Paulo E. Arratia
Data-driven particle dynamics: Structure-preserving coarse-graining for emergent behavior in non-equilibrium systems
Aug 18, 2025Abstract:Multiscale systems are ubiquitous in science and technology, but are notoriously challenging to simulate as short spatiotemporal scales must be appropriately linked to emergent bulk physics. When expensive high-dimensional dynamical systems are coarse-grained into low-dimensional models, the entropic loss of information leads to emergent physics which are dissipative, history-dependent, and stochastic. To machine learn coarse-grained dynamics from time-series observations of particle trajectories, we propose a framework using the metriplectic bracket formalism that preserves these properties by construction; most notably, the framework guarantees discrete notions of the first and second laws of thermodynamics, conservation of momentum, and a discrete fluctuation-dissipation balance crucial for capturing non-equilibrium statistics. We introduce the mathematical framework abstractly before specializing to a particle discretization. As labels are generally unavailable for entropic state variables, we introduce a novel self-supervised learning strategy to identify emergent structural variables. We validate the method on benchmark systems and demonstrate its utility on two challenging examples: (1) coarse-graining star polymers at challenging levels of coarse-graining while preserving non-equilibrium statistics, and (2) learning models from high-speed video of colloidal suspensions that capture coupling between local rearrangement events and emergent stochastic dynamics. We provide open-source implementations in both PyTorch and LAMMPS, enabling large-scale inference and extensibility to diverse particle-based systems.
Multi-environment model estimation for motility analysis of Caenorhabditis Elegans
Jul 08, 2010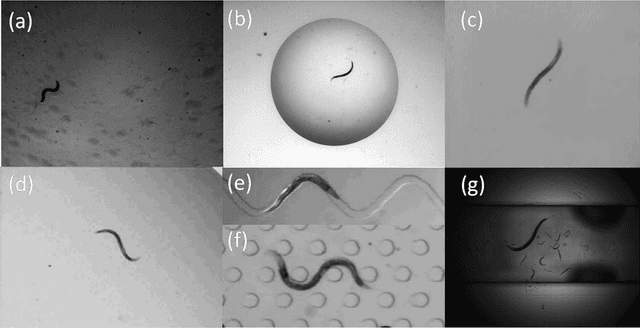
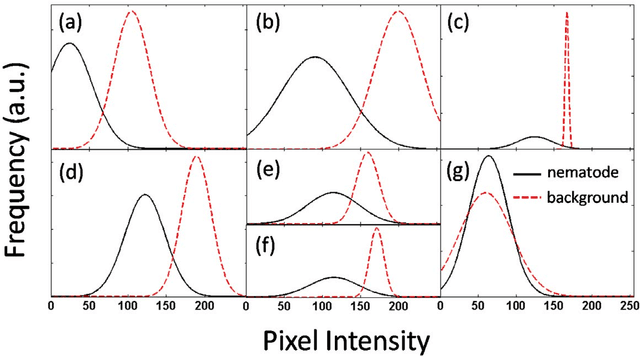
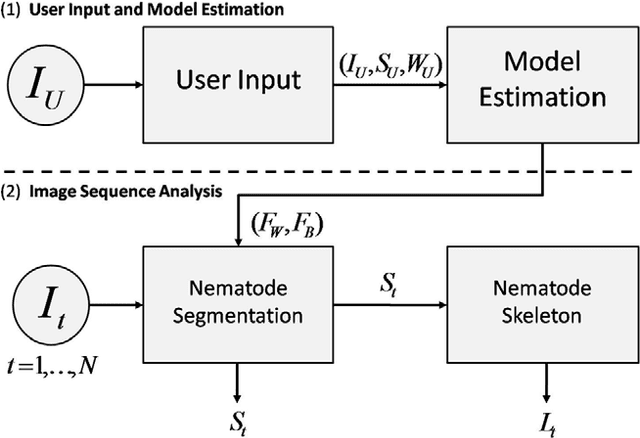
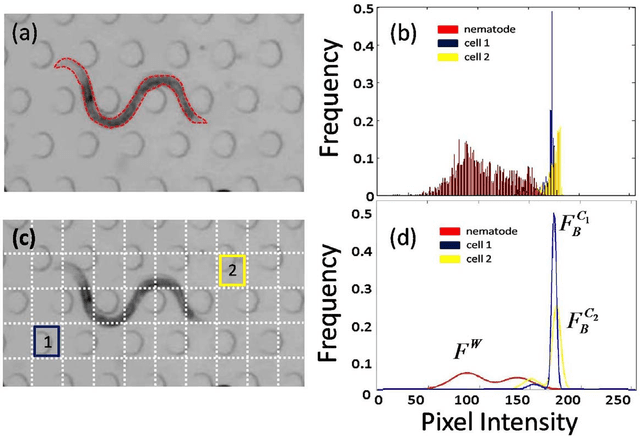
Abstract:The nematode Caenorhabditis elegans is a well-known model organism used to investigate fundamental questions in biology. Motility assays of this small roundworm are designed to study the relationships between genes and behavior. Commonly, motility analysis is used to classify nematode movements and characterize them quantitatively. Over the past years, C. elegans' motility has been studied across a wide range of environments, including crawling on substrates, swimming in fluids, and locomoting through microfluidic substrates. However, each environment often requires customized image processing tools relying on heuristic parameter tuning. In the present study, we propose a novel Multi-Environment Model Estimation (MEME) framework for automated image segmentation that is versatile across various environments. The MEME platform is constructed around the concept of Mixture of Gaussian (MOG) models, where statistical models for both the background environment and the nematode appearance are explicitly learned and used to accurately segment a target nematode. Our method is designed to simplify the burden often imposed on users; here, only a single image which includes a nematode in its environment must be provided for model learning. In addition, our platform enables the extraction of nematode `skeletons' for straightforward motility quantification. We test our algorithm on various locomotive environments and compare performances with an intensity-based thresholding method. Overall, MEME outperforms the threshold-based approach for the overwhelming majority of cases examined. Ultimately, MEME provides researchers with an attractive platform for C. elegans' segmentation and `skeletonizing' across a wide range of motility assays.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge