Efstratios Gavves
Dynamic Prototype Adaptation with Distillation for Few-shot Point Cloud Segmentation
Jan 29, 2024Abstract:Few-shot point cloud segmentation seeks to generate per-point masks for previously unseen categories, using only a minimal set of annotated point clouds as reference. Existing prototype-based methods rely on support prototypes to guide the segmentation of query point clouds, but they encounter challenges when significant object variations exist between the support prototypes and query features. In this work, we present dynamic prototype adaptation (DPA), which explicitly learns task-specific prototypes for each query point cloud to tackle the object variation problem. DPA achieves the adaptation through prototype rectification, aligning vanilla prototypes from support with the query feature distribution, and prototype-to-query attention, extracting task-specific context from query point clouds. Furthermore, we introduce a prototype distillation regularization term, enabling knowledge transfer between early-stage prototypes and their deeper counterparts during adaption. By iteratively applying these adaptations, we generate task-specific prototypes for accurate mask predictions on query point clouds. Extensive experiments on two popular benchmarks show that DPA surpasses state-of-the-art methods by a significant margin, e.g., 7.43\% and 6.39\% under the 2-way 1-shot setting on S3DIS and ScanNet, respectively. Code is available at https://github.com/jliu4ai/DPA.
How to Train Neural Field Representations: A Comprehensive Study and Benchmark
Dec 16, 2023



Abstract:Neural fields (NeFs) have recently emerged as a versatile method for modeling signals of various modalities, including images, shapes, and scenes. Subsequently, a number of works have explored the use of NeFs as representations for downstream tasks, e.g. classifying an image based on the parameters of a NeF that has been fit to it. However, the impact of the NeF hyperparameters on their quality as downstream representation is scarcely understood and remains largely unexplored. This is in part caused by the large amount of time required to fit datasets of neural fields. In this work, we propose $\verb|fit-a-nef|$, a JAX-based library that leverages parallelization to enable fast optimization of large-scale NeF datasets, resulting in a significant speed-up. With this library, we perform a comprehensive study that investigates the effects of different hyperparameters -- including initialization, network architecture, and optimization strategies -- on fitting NeFs for downstream tasks. Our study provides valuable insights on how to train NeFs and offers guidance for optimizing their effectiveness in downstream applications. Finally, based on the proposed library and our analysis, we propose Neural Field Arena, a benchmark consisting of neural field variants of popular vision datasets, including MNIST, CIFAR, variants of ImageNet, and ShapeNetv2. Our library and the Neural Field Arena will be open-sourced to introduce standardized benchmarking and promote further research on neural fields.
Motion Flow Matching for Human Motion Synthesis and Editing
Dec 14, 2023Abstract:Human motion synthesis is a fundamental task in computer animation. Recent methods based on diffusion models or GPT structure demonstrate commendable performance but exhibit drawbacks in terms of slow sampling speeds and error accumulation. In this paper, we propose \emph{Motion Flow Matching}, a novel generative model designed for human motion generation featuring efficient sampling and effectiveness in motion editing applications. Our method reduces the sampling complexity from thousand steps in previous diffusion models to just ten steps, while achieving comparable performance in text-to-motion and action-to-motion generation benchmarks. Noticeably, our approach establishes a new state-of-the-art Fr\'echet Inception Distance on the KIT-ML dataset. What is more, we tailor a straightforward motion editing paradigm named \emph{sampling trajectory rewriting} leveraging the ODE-style generative models and apply it to various editing scenarios including motion prediction, motion in-between prediction, motion interpolation, and upper-body editing. Our code will be released.
Data Augmentations in Deep Weight Spaces
Nov 15, 2023Abstract:Learning in weight spaces, where neural networks process the weights of other deep neural networks, has emerged as a promising research direction with applications in various fields, from analyzing and editing neural fields and implicit neural representations, to network pruning and quantization. Recent works designed architectures for effective learning in that space, which takes into account its unique, permutation-equivariant, structure. Unfortunately, so far these architectures suffer from severe overfitting and were shown to benefit from large datasets. This poses a significant challenge because generating data for this learning setup is laborious and time-consuming since each data sample is a full set of network weights that has to be trained. In this paper, we address this difficulty by investigating data augmentations for weight spaces, a set of techniques that enable generating new data examples on the fly without having to train additional input weight space elements. We first review several recently proposed data augmentation schemes %that were proposed recently and divide them into categories. We then introduce a novel augmentation scheme based on the Mixup method. We evaluate the performance of these techniques on existing benchmarks as well as new benchmarks we generate, which can be valuable for future studies.
Latent Field Discovery In Interacting Dynamical Systems With Neural Fields
Oct 31, 2023



Abstract:Systems of interacting objects often evolve under the influence of field effects that govern their dynamics, yet previous works have abstracted away from such effects, and assume that systems evolve in a vacuum. In this work, we focus on discovering these fields, and infer them from the observed dynamics alone, without directly observing them. We theorize the presence of latent force fields, and propose neural fields to learn them. Since the observed dynamics constitute the net effect of local object interactions and global field effects, recently popularized equivariant networks are inapplicable, as they fail to capture global information. To address this, we propose to disentangle local object interactions -- which are $\mathrm{SE}(n)$ equivariant and depend on relative states -- from external global field effects -- which depend on absolute states. We model interactions with equivariant graph networks, and combine them with neural fields in a novel graph network that integrates field forces. Our experiments show that we can accurately discover the underlying fields in charged particles settings, traffic scenes, and gravitational n-body problems, and effectively use them to learn the system and forecast future trajectories.
Time Does Tell: Self-Supervised Time-Tuning of Dense Image Representations
Aug 22, 2023Abstract:Spatially dense self-supervised learning is a rapidly growing problem domain with promising applications for unsupervised segmentation and pretraining for dense downstream tasks. Despite the abundance of temporal data in the form of videos, this information-rich source has been largely overlooked. Our paper aims to address this gap by proposing a novel approach that incorporates temporal consistency in dense self-supervised learning. While methods designed solely for images face difficulties in achieving even the same performance on videos, our method improves not only the representation quality for videos-but also images. Our approach, which we call time-tuning, starts from image-pretrained models and fine-tunes them with a novel self-supervised temporal-alignment clustering loss on unlabeled videos. This effectively facilitates the transfer of high-level information from videos to image representations. Time-tuning improves the state-of-the-art by 8-10% for unsupervised semantic segmentation on videos and matches it for images. We believe this method paves the way for further self-supervised scaling by leveraging the abundant availability of videos. The implementation can be found here : https://github.com/SMSD75/Timetuning
Neural Modulation Fields for Conditional Cone Beam Neural Tomography
Jul 17, 2023



Abstract:Conventional Computed Tomography (CT) methods require large numbers of noise-free projections for accurate density reconstructions, limiting their applicability to the more complex class of Cone Beam Geometry CT (CBCT) reconstruction. Recently, deep learning methods have been proposed to overcome these limitations, with methods based on neural fields (NF) showing strong performance, by approximating the reconstructed density through a continuous-in-space coordinate based neural network. Our focus is on improving such methods, however, unlike previous work, which requires training an NF from scratch for each new set of projections, we instead propose to leverage anatomical consistencies over different scans by training a single conditional NF on a dataset of projections. We propose a novel conditioning method where local modulations are modeled per patient as a field over the input domain through a Neural Modulation Field (NMF). The resulting Conditional Cone Beam Neural Tomography (CondCBNT) shows improved performance for both high and low numbers of available projections on noise-free and noisy data.
Learning Lie Group Symmetry Transformations with Neural Networks
Jul 04, 2023Abstract:The problem of detecting and quantifying the presence of symmetries in datasets is useful for model selection, generative modeling, and data analysis, amongst others. While existing methods for hard-coding transformations in neural networks require prior knowledge of the symmetries of the task at hand, this work focuses on discovering and characterizing unknown symmetries present in the dataset, namely, Lie group symmetry transformations beyond the traditional ones usually considered in the field (rotation, scaling, and translation). Specifically, we consider a scenario in which a dataset has been transformed by a one-parameter subgroup of transformations with different parameter values for each data point. Our goal is to characterize the transformation group and the distribution of the parameter values. The results showcase the effectiveness of the approach in both these settings.
BISCUIT: Causal Representation Learning from Binary Interactions
Jun 16, 2023



Abstract:Identifying the causal variables of an environment and how to intervene on them is of core value in applications such as robotics and embodied AI. While an agent can commonly interact with the environment and may implicitly perturb the behavior of some of these causal variables, often the targets it affects remain unknown. In this paper, we show that causal variables can still be identified for many common setups, e.g., additive Gaussian noise models, if the agent's interactions with a causal variable can be described by an unknown binary variable. This happens when each causal variable has two different mechanisms, e.g., an observational and an interventional one. Using this identifiability result, we propose BISCUIT, a method for simultaneously learning causal variables and their corresponding binary interaction variables. On three robotic-inspired datasets, BISCUIT accurately identifies causal variables and can even be scaled to complex, realistic environments for embodied AI.
Graph Switching Dynamical Systems
Jun 01, 2023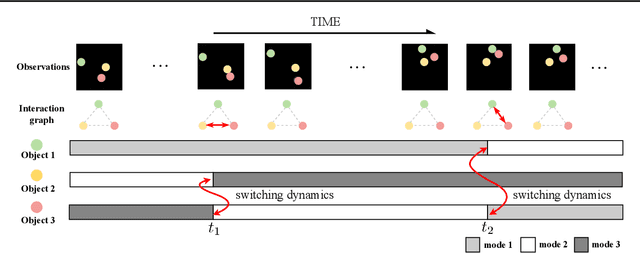
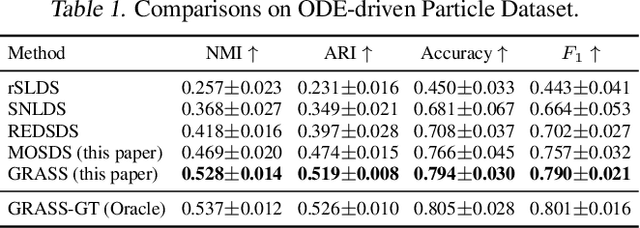
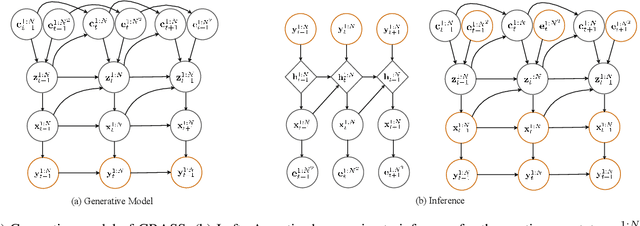
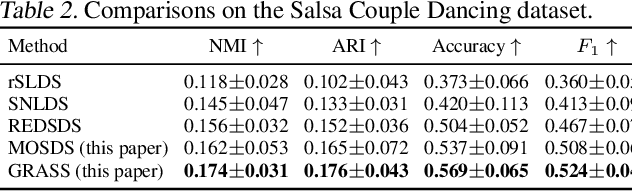
Abstract:Dynamical systems with complex behaviours, e.g. immune system cells interacting with a pathogen, are commonly modelled by splitting the behaviour into different regimes, or modes, each with simpler dynamics, and then learning the switching behaviour from one mode to another. Switching Dynamical Systems (SDS) are a powerful tool that automatically discovers these modes and mode-switching behaviour from time series data. While effective, these methods focus on independent objects, where the modes of one object are independent of the modes of the other objects. In this paper, we focus on the more general interacting object setting for switching dynamical systems, where the per-object dynamics also depends on an unknown and dynamically changing subset of other objects and their modes. To this end, we propose a novel graph-based approach for switching dynamical systems, GRAph Switching dynamical Systems (GRASS), in which we use a dynamic graph to characterize interactions between objects and learn both intra-object and inter-object mode-switching behaviour. We introduce two new datasets for this setting, a synthesized ODE-driven particles dataset and a real-world Salsa Couple Dancing dataset. Experiments show that GRASS can consistently outperforms previous state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge