Davood Karimi
Fetal-BET: Brain Extraction Tool for Fetal MRI
Oct 02, 2023



Abstract:Fetal brain extraction is a necessary first step in most computational fetal brain MRI pipelines. However, it has been a very challenging task due to non-standard fetal head pose, fetal movements during examination, and vastly heterogeneous appearance of the developing fetal brain and the neighboring fetal and maternal anatomy across various sequences and scanning conditions. Development of a machine learning method to effectively address this task requires a large and rich labeled dataset that has not been previously available. As a result, there is currently no method for accurate fetal brain extraction on various fetal MRI sequences. In this work, we first built a large annotated dataset of approximately 72,000 2D fetal brain MRI images. Our dataset covers the three common MRI sequences including T2-weighted, diffusion-weighted, and functional MRI acquired with different scanners. Moreover, it includes normal and pathological brains. Using this dataset, we developed and validated deep learning methods, by exploiting the power of the U-Net style architectures, the attention mechanism, multi-contrast feature learning, and data augmentation for fast, accurate, and generalizable automatic fetal brain extraction. Our approach leverages the rich information from multi-contrast (multi-sequence) fetal MRI data, enabling precise delineation of the fetal brain structures. Evaluations on independent test data show that our method achieves accurate brain extraction on heterogeneous test data acquired with different scanners, on pathological brains, and at various gestational stages. This robustness underscores the potential utility of our deep learning model for fetal brain imaging and image analysis.
Characterizing normal perinatal development of the human brain structural connectivity
Aug 22, 2023Abstract:Early brain development is characterized by the formation of a highly organized structural connectome. The interconnected nature of this connectome underlies the brain's cognitive abilities and influences its response to diseases and environmental factors. Hence, quantitative assessment of structural connectivity in the perinatal stage is useful for studying normal and abnormal neurodevelopment. However, estimation of the connectome from diffusion MRI data involves complex computations. For the perinatal period, these computations are further challenged by the rapid brain development and imaging difficulties. Combined with high inter-subject variability, these factors make it difficult to chart the normal development of the structural connectome. As a result, there is a lack of reliable normative baselines of structural connectivity metrics at this critical stage in brain development. In this study, we developed a computational framework, based on spatio-temporal averaging, for determining such baselines. We used this framework to analyze the structural connectivity between 33 and 44 postmenstrual weeks using data from 166 subjects. Our results unveiled clear and strong trends in the development of structural connectivity in perinatal stage. Connection weighting based on fractional anisotropy and neurite density produced the most consistent results. We observed increases in global and local efficiency, a decrease in characteristic path length, and widespread strengthening of the connections within and across brain lobes and hemispheres. We also observed asymmetry patterns that were consistent between different connection weighting approaches. The new computational method and results are useful for assessing normal and abnormal development of the structural connectome early in life.
TBSS++: A novel computational method for Tract-Based Spatial Statistics
Jul 07, 2023Abstract:Diffusion-weighted magnetic resonance imaging (dMRI) is widely used to assess the brain white matter. One of the most common computations in dMRI involves cross-subject tract-specific analysis, whereby dMRI-derived biomarkers are compared between cohorts of subjects. The accuracy and reliability of these studies hinges on the ability to compare precisely the same white matter tracts across subjects. This is an intricate and error-prone computation. Existing computational methods such as Tract-Based Spatial Statistics (TBSS) suffer from a host of shortcomings and limitations that can seriously undermine the validity of the results. We present a new computational framework that overcomes the limitations of existing methods via (i) accurate segmentation of the tracts, and (ii) precise registration of data from different subjects/scans. The registration is based on fiber orientation distributions. To further improve the alignment of cross-subject data, we create detailed atlases of white matter tracts. These atlases serve as an unbiased reference space where the data from all subjects is registered for comparison. Extensive evaluations show that, compared with TBSS, our proposed framework offers significantly higher reproducibility and robustness to data perturbations. Our method promises a drastic improvement in accuracy and reproducibility of cross-subject dMRI studies that are routinely used in neuroscience and medical research.
Direct segmentation of brain white matter tracts in diffusion MRI
Jul 05, 2023



Abstract:The brain white matter consists of a set of tracts that connect distinct regions of the brain. Segmentation of these tracts is often needed for clinical and research studies. Diffusion-weighted MRI offers unique contrast to delineate these tracts. However, existing segmentation methods rely on intermediate computations such as tractography or estimation of fiber orientation density. These intermediate computations, in turn, entail complex computations that can result in unnecessary errors. Moreover, these intermediate computations often require dense multi-shell measurements that are unavailable in many clinical and research applications. As a result, current methods suffer from low accuracy and poor generalizability. Here, we propose a new deep learning method that segments these tracts directly from the diffusion MRI data, thereby sidestepping the intermediate computation errors. Our experiments show that this method can achieve segmentation accuracy that is on par with the state of the art methods (mean Dice Similarity Coefficient of 0.826). Compared with the state of the art, our method offers far superior generalizability to undersampled data that are typical of clinical studies and to data obtained with different acquisition protocols. Moreover, we propose a new method for detecting inaccurate segmentations and show that it is more accurate than standard methods that are based on estimation uncertainty quantification. The new methods can serve many critically important clinical and scientific applications that require accurate and reliable non-invasive segmentation of white matter tracts.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Atlas-powered deep learning (ADL) -- application to diffusion weighted MRI
May 05, 2022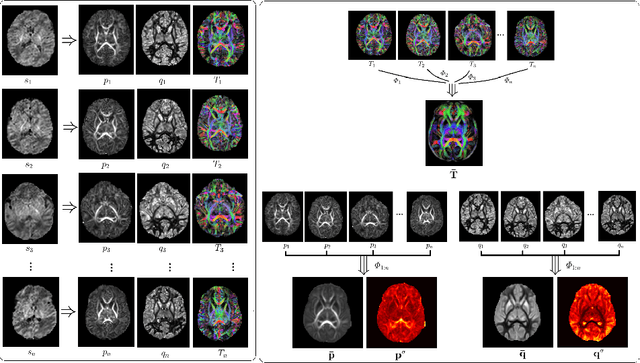

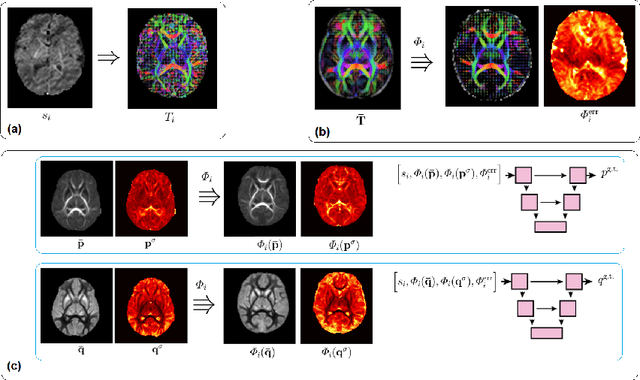

Abstract:Deep learning has a great potential for estimating biomarkers in diffusion weighted magnetic resonance imaging (dMRI). Atlases, on the other hand, are a unique tool for modeling the spatio-temporal variability of biomarkers. In this paper, we propose the first framework to exploit both deep learning and atlases for biomarker estimation in dMRI. Our framework relies on non-linear diffusion tensor registration to compute biomarker atlases and to estimate atlas reliability maps. We also use nonlinear tensor registration to align the atlas to a subject and to estimate the error of this alignment. We use the biomarker atlas, atlas reliability map, and alignment error map, in addition to the dMRI signal, as inputs to a deep learning model for biomarker estimation. We use our framework to estimate fractional anisotropy and neurite orientation dispersion from down-sampled dMRI data on a test cohort of 70 newborn subjects. Results show that our method significantly outperforms standard estimation methods as well as recent deep learning techniques. Our method is also more robust to stronger measurement down-sampling factors. Our study shows that the advantages of deep learning and atlases can be synergistically combined to achieve unprecedented accuracy in biomarker estimation from dMRI data.
Deep Learning Framework for Real-time Fetal Brain Segmentation in MRI
May 02, 2022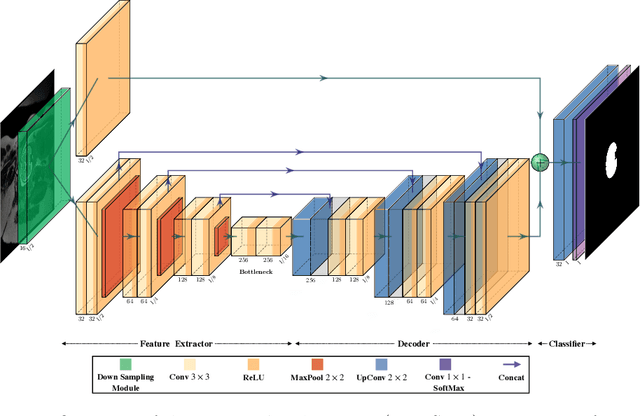
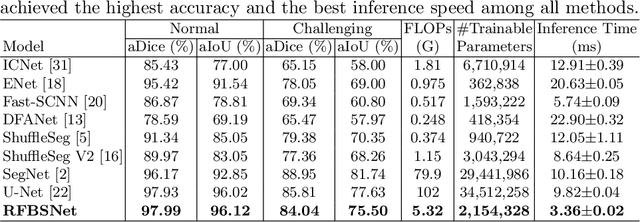
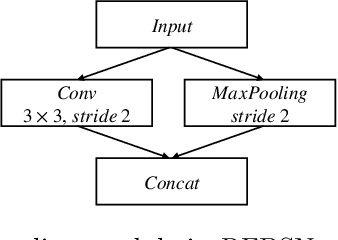
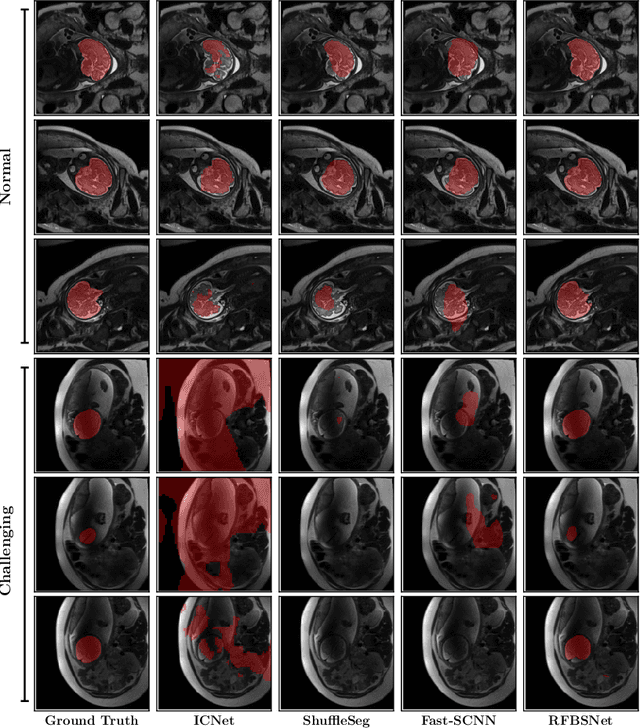
Abstract:Fetal brain segmentation is an important first step for slice-level motion correction and slice-to-volume reconstruction in fetal MRI. Fast and accurate segmentation of the fetal brain on fetal MRI is required to achieve real-time fetal head pose estimation and motion tracking for slice re-acquisition and steering. To address this critical unmet need, in this work we analyzed the speed-accuracy performance of a variety of deep neural network models, and devised a symbolically small convolutional neural network that combines spatial details at high resolution with context features extracted at lower resolutions. We used multiple branches with skip connections to maintain high accuracy while devising a parallel combination of convolution and pooling operations as an input downsampling module to further reduce inference time. We trained our model as well as eight alternative, state-of-the-art networks with manually-labeled fetal brain MRI slices and tested on two sets of normal and challenging test cases. Experimental results show that our network achieved the highest accuracy and lowest inference time among all of the compared state-of-the-art real-time segmentation methods. We achieved average Dice scores of 97.99\% and 84.04\% on the normal and challenging test sets, respectively, with an inference time of 3.36 milliseconds per image on an NVIDIA GeForce RTX 2080 Ti. Code, data, and the trained models are available at https://github.com/bchimagine/real_time_fetal_brain_segmentation.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022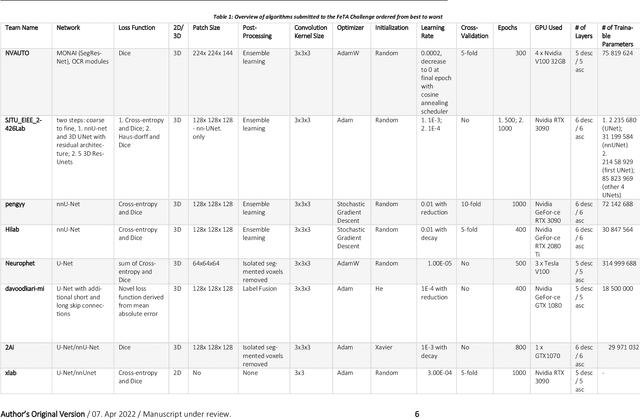
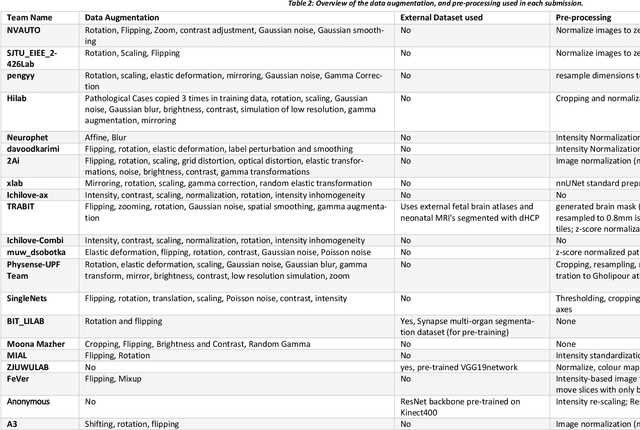
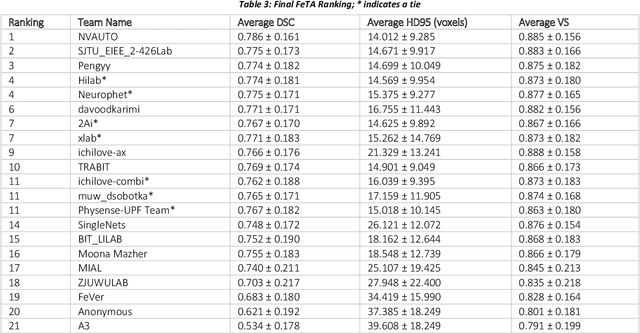
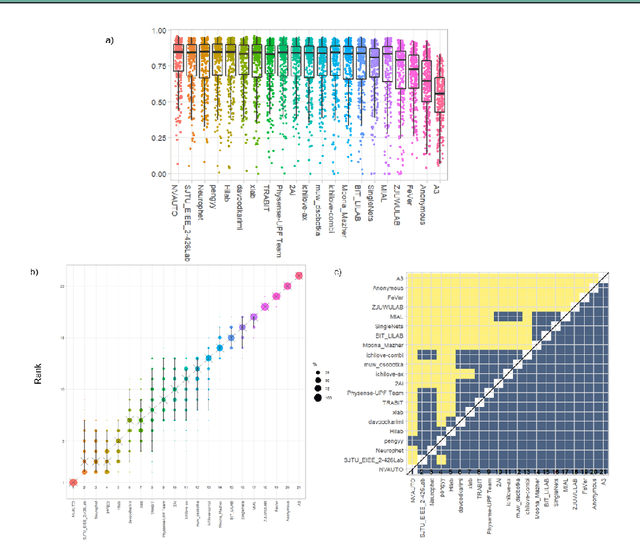
Abstract:In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
Learning to segment fetal brain tissue from noisy annotations
Mar 25, 2022
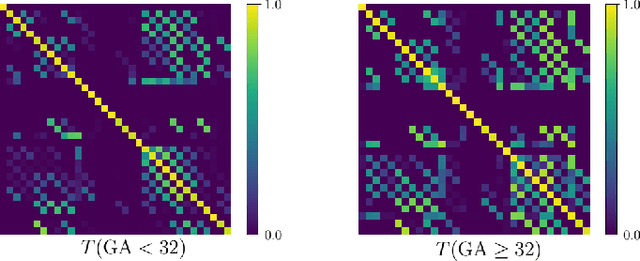
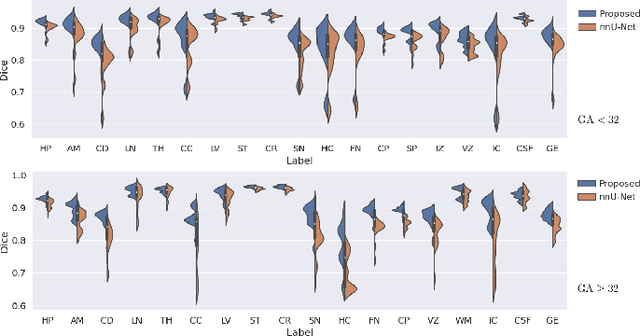
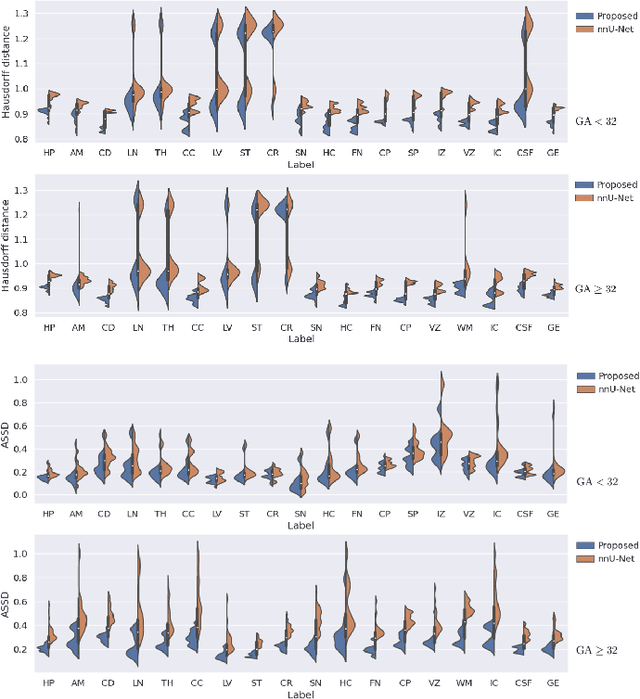
Abstract:Automatic fetal brain tissue segmentation can enhance the quantitative assessment of brain development at this critical stage. Deep learning methods represent the state of the art in medical image segmentation and have also achieved impressive results in brain segmentation. However, effective training of a deep learning model to perform this task requires a large number of training images to represent the rapid development of the transient fetal brain structures. On the other hand, manual multi-label segmentation of a large number of 3D images is prohibitive. To address this challenge, we segmented 272 training images, covering 19-39 gestational weeks, using an automatic multi-atlas segmentation strategy based on deformable registration and probabilistic atlas fusion, and manually corrected large errors in those segmentations. Since this process generated a large training dataset with noisy segmentations, we developed a novel label smoothing procedure and a loss function to train a deep learning model with smoothed noisy segmentations. Our proposed methods properly account for the uncertainty in tissue boundaries. We evaluated our method on 23 manually-segmented test images of a separate set of fetuses. Results show that our method achieves an average Dice similarity coefficient of 0.893 and 0.916 for the transient structures of younger and older fetuses, respectively. Our method generated results that were significantly more accurate than several state-of-the-art methods including nnU-Net that achieved the closest results to our method. Our trained model can serve as a valuable tool to enhance the accuracy and reproducibility of fetal brain analysis in MRI.
Diffusion Tensor Estimation with Transformer Neural Networks
Jan 14, 2022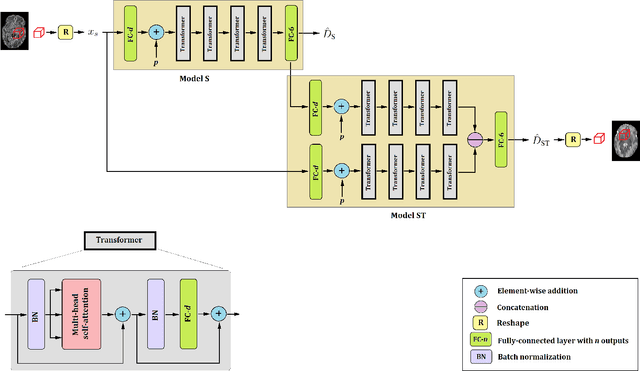
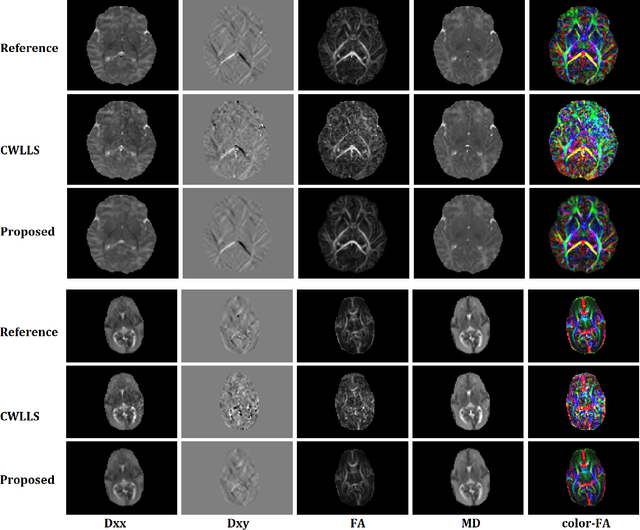
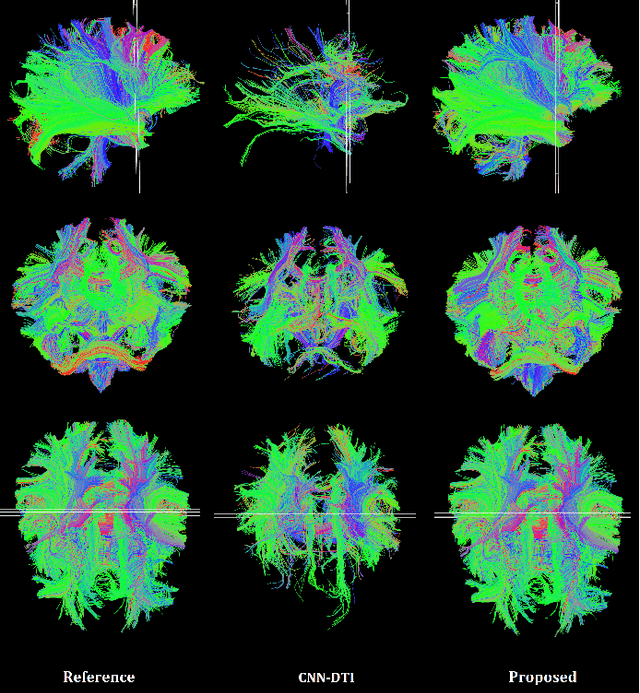
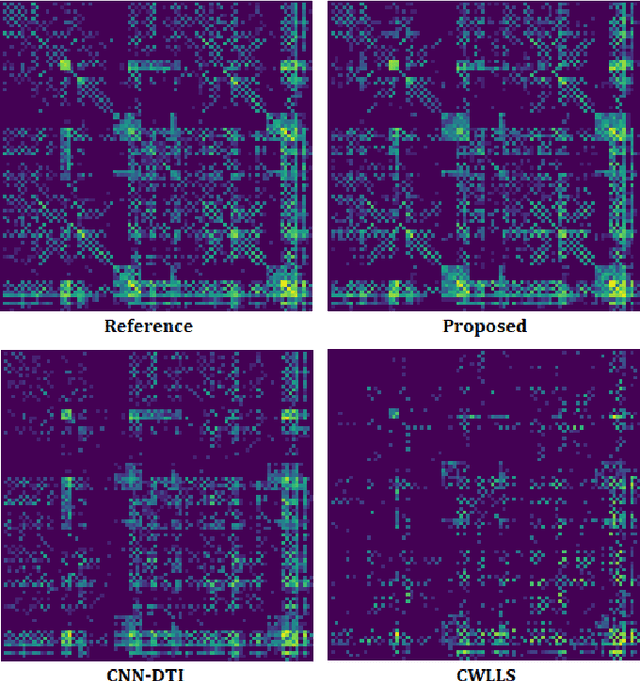
Abstract:Diffusion tensor imaging (DTI) is the most widely used tool for studying brain white matter development and degeneration. However, standard DTI estimation methods depend on a large number of high-quality measurements. This would require long scan times and can be particularly difficult to achieve with certain patient populations such as neonates. Here, we propose a method that can accurately estimate the diffusion tensor from only six diffusion-weighted measurements. Our method achieves this by learning to exploit the relationships between the diffusion signals and tensors in neighboring voxels. Our model is based on transformer networks, which represent the state of the art in modeling the relationship between signals in a sequence. In particular, our model consists of two such networks. The first network estimates the diffusion tensor based on the diffusion signals in a neighborhood of voxels. The second network provides more accurate tensor estimations by learning the relationships between the diffusion signals as well as the tensors estimated by the first network in neighboring voxels. Our experiments with three datasets show that our proposed method achieves highly accurate estimations of the diffusion tensor and is significantly superior to three competing methods. Estimations produced by our method with six measurements are comparable with those of standard estimation methods with 30-88 measurements. Hence, our method promises shorter scan times and more reliable assessment of brain white matter, particularly in non-cooperative patients such as neonates and infants.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge