Daoqiang Zhang
JCCS-PFGM: A Novel Circle-Supervision based Poisson Flow Generative Model for Multiphase CECT Progressive Low-Dose Reconstruction with Joint Condition
Jun 13, 2023Abstract:Multiphase contrast-enhanced computed tomography (CECT) scan is clinically significant to demonstrate the anatomy at different phases. In practice, such a multiphase CECT scan inherently takes longer time and deposits much more radiation dose into a patient body than a regular CT scan, and reduction of the radiation dose typically compromise the CECT image quality and its diagnostic value. With Joint Condition and Circle-Supervision, here we propose a novel Poisson Flow Generative Model (JCCS-PFGM) to promote the progressive low-dose reconstruction for multiphase CECT. JCCS-PFGM is characterized by the following three aspects: a progressive low-dose reconstruction scheme, a circle-supervision strategy, and a joint condition mechanism. Our extensive experiments are performed on a clinical dataset consisting of 11436 images. The results show that our JCCS-PFGM achieves promising PSNR up to 46.3dB, SSIM up to 98.5%, and MAE down to 9.67 HU averagely on phases I, II and III, in quantitative evaluations, as well as gains high-quality readable visualizations in qualitative assessments. All of these findings reveal our method a great potential to be adapted for clinical CECT scans at a much-reduced radiation dose.
Uncertainty-inspired Open Set Learning for Retinal Anomaly Identification
Apr 08, 2023



Abstract:Failure to recognize samples from the classes unseen during training is a major limit of artificial intelligence (AI) in real-world implementation of retinal anomaly classification. To resolve this obstacle, we propose an uncertainty-inspired open-set (UIOS) model which was trained with fundus images of 9 common retinal conditions. Besides the probability of each category, UIOS also calculates an uncertainty score to express its confidence. Our UIOS model with thresholding strategy achieved an F1 score of 99.55%, 97.01% and 91.91% for the internal testing set, external testing set and non-typical testing set, respectively, compared to the F1 score of 92.20%, 80.69% and 64.74% by the standard AI model. Furthermore, UIOS correctly predicted high uncertainty scores, which prompted the need for a manual check, in the datasets of rare retinal diseases, low-quality fundus images, and non-fundus images. This work provides a robust method for real-world screening of retinal anomalies.
Model Barrier: A Compact Un-Transferable Isolation Domain for Model Intellectual Property Protection
Mar 20, 2023Abstract:As scientific and technological advancements result from human intellectual labor and computational costs, protecting model intellectual property (IP) has become increasingly important to encourage model creators and owners. Model IP protection involves preventing the use of well-trained models on unauthorized domains. To address this issue, we propose a novel approach called Compact Un-Transferable Isolation Domain (CUTI-domain), which acts as a barrier to block illegal transfers from authorized to unauthorized domains. Specifically, CUTI-domain blocks cross-domain transfers by highlighting the private style features of the authorized domain, leading to recognition failure on unauthorized domains with irrelevant private style features. Moreover, we provide two solutions for using CUTI-domain depending on whether the unauthorized domain is known or not: target-specified CUTI-domain and target-free CUTI-domain. Our comprehensive experimental results on four digit datasets, CIFAR10 & STL10, and VisDA-2017 dataset demonstrate that CUTI-domain can be easily implemented as a plug-and-play module with different backbones, providing an efficient solution for model IP protection.
sMRI-PatchNet: A novel explainable patch-based deep learning network for Alzheimer's disease diagnosis and discriminative atrophy localisation with Structural MRI
Feb 20, 2023



Abstract:Structural magnetic resonance imaging (sMRI) can identify subtle brain changes due to its high contrast for soft tissues and high spatial resolution. It has been widely used in diagnosing neurological brain diseases, such as Alzheimer disease (AD). However, the size of 3D high-resolution data poses a significant challenge for data analysis and processing. Since only a few areas of the brain show structural changes highly associated with AD, the patch-based methods dividing the whole image data into several small regular patches have shown promising for more efficient sMRI-based image analysis. The major challenges of the patch-based methods on sMRI include identifying the discriminative patches, combining features from the discrete discriminative patches, and designing appropriate classifiers. This work proposes a novel patch-based deep learning network (sMRI-PatchNet) with explainable patch localisation and selection for AD diagnosis using sMRI. Specifically, it consists of two primary components: 1) A fast and efficient explainable patch selection mechanism for determining the most discriminative patches based on computing the SHapley Additive exPlanations (SHAP) contribution to a transfer learning model for AD diagnosis on massive medical data; and 2) A novel patch-based network for extracting deep features and AD classfication from the selected patches with position embeddings to retain position information, capable of capturing the global and local information of inter- and intra-patches. This method has been applied for the AD classification and the prediction of the transitional state moderate cognitive impairment (MCI) conversion with real datasets.
Complementary Labels Learning with Augmented Classes
Nov 19, 2022



Abstract:Complementary Labels Learning (CLL) arises in many real-world tasks such as private questions classification and online learning, which aims to alleviate the annotation cost compared with standard supervised learning. Unfortunately, most previous CLL algorithms were in a stable environment rather than an open and dynamic scenarios, where data collected from unseen augmented classes in the training process might emerge in the testing phase. In this paper, we propose a novel problem setting called Complementary Labels Learning with Augmented Classes (CLLAC), which brings the challenge that classifiers trained by complementary labels should not only be able to classify the instances from observed classes accurately, but also recognize the instance from the Augmented Classes in the testing phase. Specifically, by using unlabeled data, we propose an unbiased estimator of classification risk for CLLAC, which is guaranteed to be provably consistent. Moreover, we provide generalization error bound for proposed method which shows that the optimal parametric convergence rate is achieved for estimation error. Finally, the experimental results on several benchmark datasets verify the effectiveness of the proposed method.
Unsupervised Domain Adaptation via Style-Aware Self-intermediate Domain
Sep 05, 2022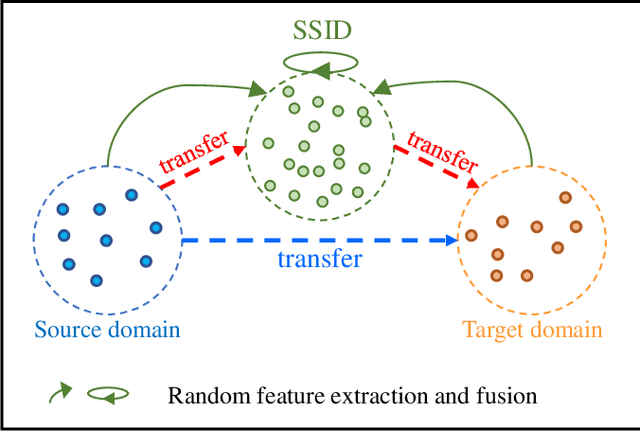
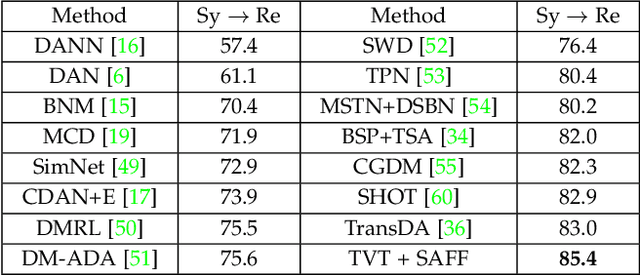
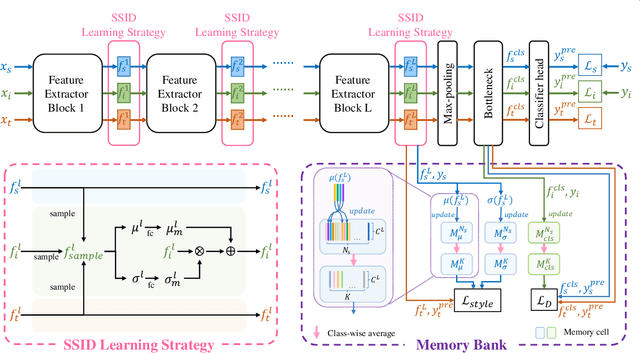
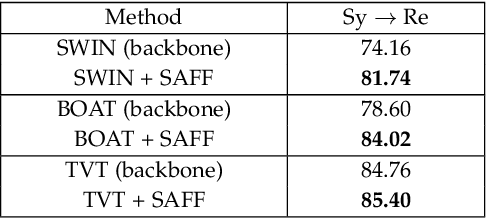
Abstract:Unsupervised domain adaptation (UDA) has attracted considerable attention, which transfers knowledge from a label-rich source domain to a related but unlabeled target domain. Reducing inter-domain differences has always been a crucial factor to improve performance in UDA, especially for tasks where there is a large gap between source and target domains. To this end, we propose a novel style-aware feature fusion method (SAFF) to bridge the large domain gap and transfer knowledge while alleviating the loss of class-discriminative information. Inspired by the human transitive inference and learning ability, a novel style-aware self-intermediate domain (SSID) is investigated to link two seemingly unrelated concepts through a series of intermediate auxiliary synthesized concepts. Specifically, we propose a novel learning strategy of SSID, which selects samples from both source and target domains as anchors, and then randomly fuses the object and style features of these anchors to generate labeled and style-rich intermediate auxiliary features for knowledge transfer. Moreover, we design an external memory bank to store and update specified labeled features to obtain stable class features and class-wise style features. Based on the proposed memory bank, the intra- and inter-domain loss functions are designed to improve the class recognition ability and feature compatibility, respectively. Meanwhile, we simulate the rich latent feature space of SSID by infinite sampling and the convergence of the loss function by mathematical theory. Finally, we conduct comprehensive experiments on commonly used domain adaptive benchmarks to evaluate the proposed SAFF, and the experimental results show that the proposed SAFF can be easily combined with different backbone networks and obtain better performance as a plug-in-plug-out module.
Learning from Positive and Unlabeled Data with Augmented Classes
Jul 27, 2022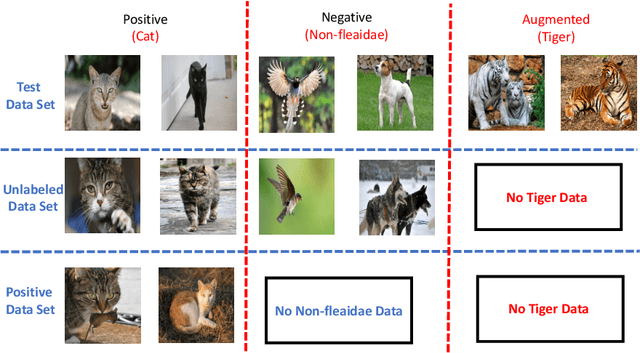
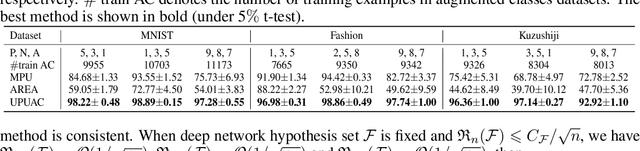
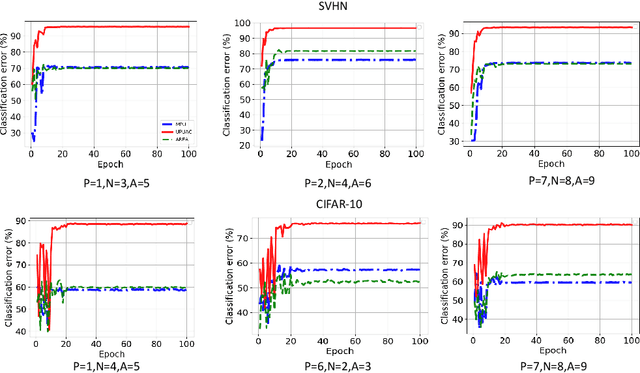
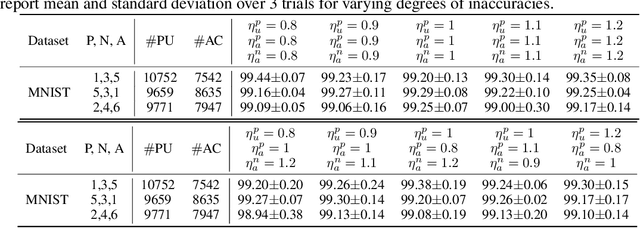
Abstract:Positive Unlabeled (PU) learning aims to learn a binary classifier from only positive and unlabeled data, which is utilized in many real-world scenarios. However, existing PU learning algorithms cannot deal with the real-world challenge in an open and changing scenario, where examples from unobserved augmented classes may emerge in the testing phase. In this paper, we propose an unbiased risk estimator for PU learning with Augmented Classes (PUAC) by utilizing unlabeled data from the augmented classes distribution, which can be easily collected in many real-world scenarios. Besides, we derive the estimation error bound for the proposed estimator, which provides a theoretical guarantee for its convergence to the optimal solution. Experiments on multiple realistic datasets demonstrate the effectiveness of proposed approach.
Layer-Wise Partitioning and Merging for Efficient and Scalable Deep Learning
Jul 22, 2022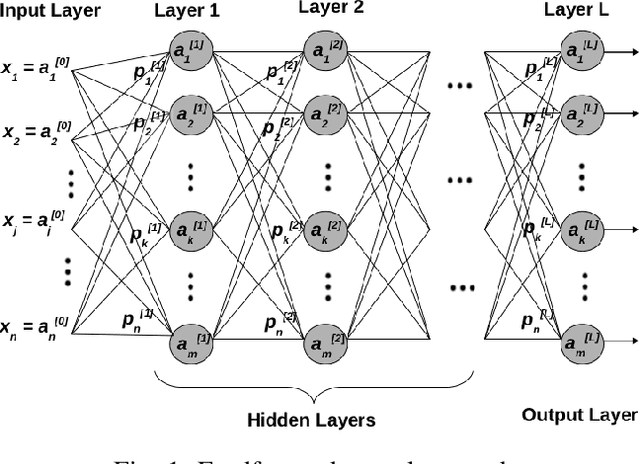
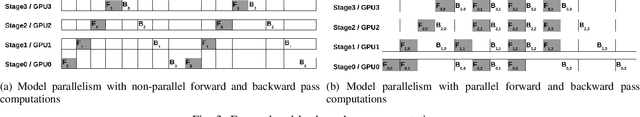
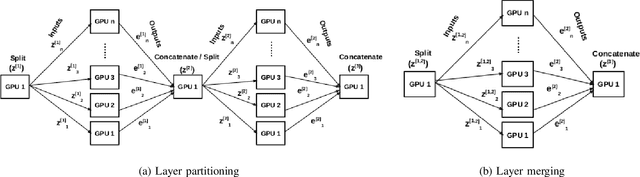

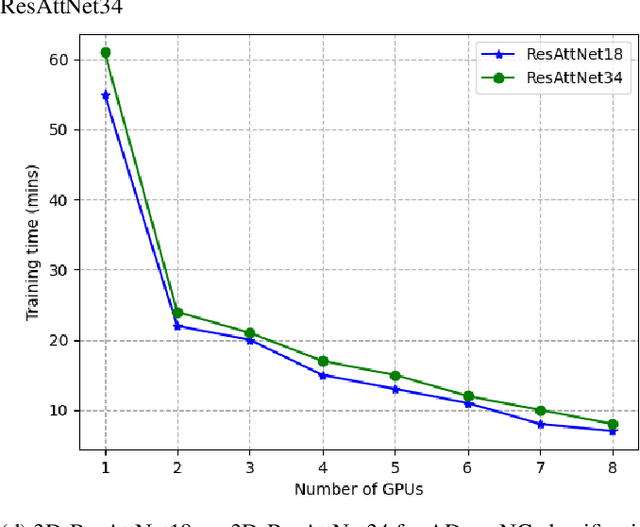
Abstract:Deep Neural Network (DNN) models are usually trained sequentially from one layer to another, which causes forward, backward and update locking's problems, leading to poor performance in terms of training time. The existing parallel strategies to mitigate these problems provide suboptimal runtime performance. In this work, we have proposed a novel layer-wise partitioning and merging, forward and backward pass parallel framework to provide better training performance. The novelty of the proposed work consists of 1) a layer-wise partition and merging model which can minimise communication overhead between devices without the memory cost of existing strategies during the training process; 2) a forward pass and backward pass parallelisation and optimisation to address the update locking problem and minimise the total training cost. The experimental evaluation on real use cases shows that the proposed method outperforms the state-of-the-art approaches in terms of training speed; and achieves almost linear speedup without compromising the accuracy performance of the non-parallel approach.
InfoAT: Improving Adversarial Training Using the Information Bottleneck Principle
Jun 23, 2022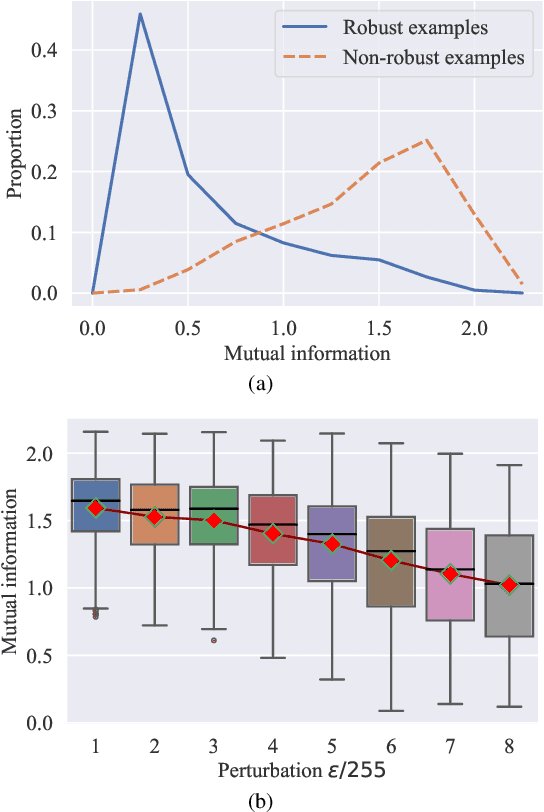
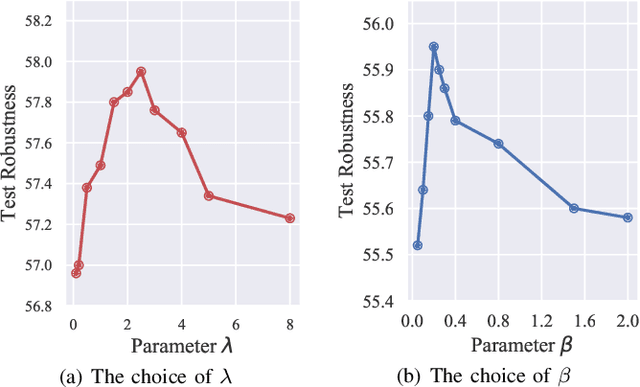
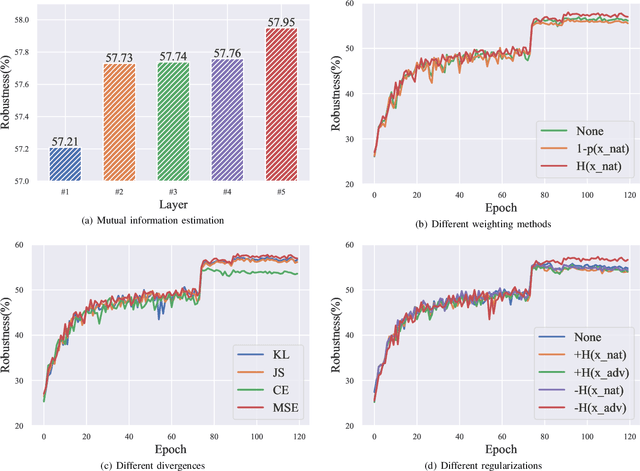
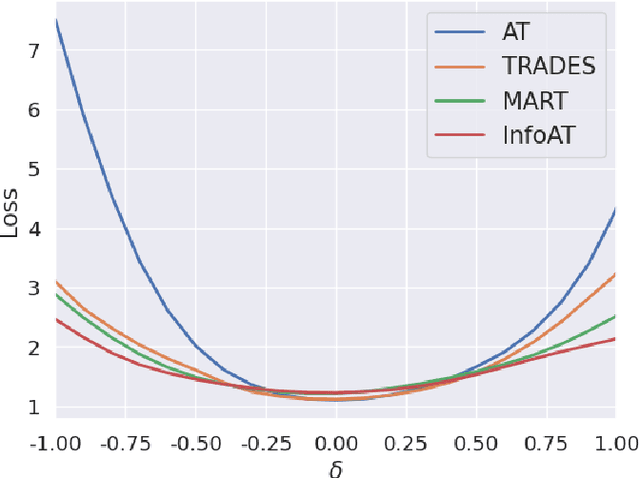
Abstract:Adversarial training (AT) has shown excellent high performance in defending against adversarial examples. Recent studies demonstrate that examples are not equally important to the final robustness of models during AT, that is, the so-called hard examples that can be attacked easily exhibit more influence than robust examples on the final robustness. Therefore, guaranteeing the robustness of hard examples is crucial for improving the final robustness of the model. However, defining effective heuristics to search for hard examples is still difficult. In this article, inspired by the information bottleneck (IB) principle, we uncover that an example with high mutual information of the input and its associated latent representation is more likely to be attacked. Based on this observation, we propose a novel and effective adversarial training method (InfoAT). InfoAT is encouraged to find examples with high mutual information and exploit them efficiently to improve the final robustness of models. Experimental results show that InfoAT achieves the best robustness among different datasets and models in comparison with several state-of-the-art methods.
Low-Dose CT Denoising via Sinogram Inner-Structure Transformer
Apr 18, 2022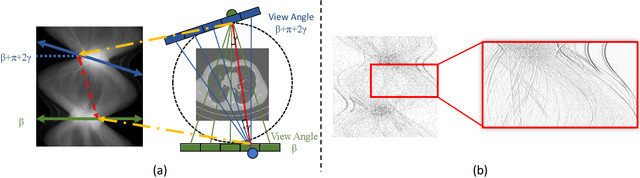
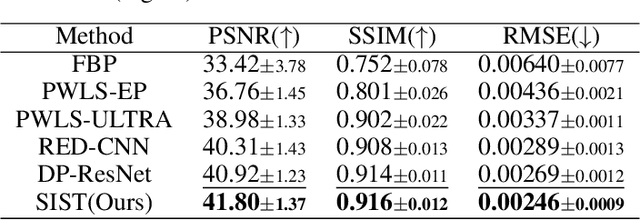
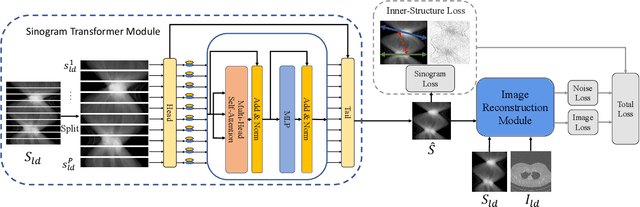
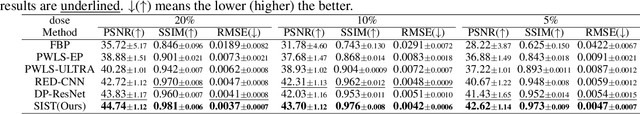
Abstract:Low-Dose Computed Tomography (LDCT) technique, which reduces the radiation harm to human bodies, is now attracting increasing interest in the medical imaging field. As the image quality is degraded by low dose radiation, LDCT exams require specialized reconstruction methods or denoising algorithms. However, most of the recent effective methods overlook the inner-structure of the original projection data (sinogram) which limits their denoising ability. The inner-structure of the sinogram represents special characteristics of the data in the sinogram domain. By maintaining this structure while denoising, the noise can be obviously restrained. Therefore, we propose an LDCT denoising network namely Sinogram Inner-Structure Transformer (SIST) to reduce the noise by utilizing the inner-structure in the sinogram domain. Specifically, we study the CT imaging mechanism and statistical characteristics of sinogram to design the sinogram inner-structure loss including the global and local inner-structure for restoring high-quality CT images. Besides, we propose a sinogram transformer module to better extract sinogram features. The transformer architecture using a self-attention mechanism can exploit interrelations between projections of different view angles, which achieves an outstanding performance in sinogram denoising. Furthermore, in order to improve the performance in the image domain, we propose the image reconstruction module to complementarily denoise both in the sinogram and image domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge