Christos Davatzikos
from the iSTAGING consortium, for the ADNI
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023



Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
Adapting Machine Learning Diagnostic Models to New Populations Using a Small Amount of Data: Results from Clinical Neuroscience
Aug 06, 2023



Abstract:Machine learning (ML) has shown great promise for revolutionizing a number of areas, including healthcare. However, it is also facing a reproducibility crisis, especially in medicine. ML models that are carefully constructed from and evaluated on a training set might not generalize well on data from different patient populations or acquisition instrument settings and protocols. We tackle this problem in the context of neuroimaging of Alzheimer's disease (AD), schizophrenia (SZ) and brain aging. We develop a weighted empirical risk minimization approach that optimally combines data from a source group, e.g., subjects are stratified by attributes such as sex, age group, race and clinical cohort to make predictions on a target group, e.g., other sex, age group, etc. using a small fraction (10%) of data from the target group. We apply this method to multi-source data of 15,363 individuals from 20 neuroimaging studies to build ML models for diagnosis of AD and SZ, and estimation of brain age. We found that this approach achieves substantially better accuracy than existing domain adaptation techniques: it obtains area under curve greater than 0.95 for AD classification, area under curve greater than 0.7 for SZ classification and mean absolute error less than 5 years for brain age prediction on all target groups, achieving robustness to variations of scanners, protocols, and demographic or clinical characteristics. In some cases, it is even better than training on all data from the target group, because it leverages the diversity and size of a larger training set. We also demonstrate the utility of our models for prognostic tasks such as predicting disease progression in individuals with mild cognitive impairment. Critically, our brain age prediction models lead to new clinical insights regarding correlations with neurophysiological tests.
The Brain Tumor Segmentation Challenge 2023: Brain MR Image Synthesis for Tumor Segmentation
May 20, 2023
Abstract:Automated brain tumor segmentation methods are well established, reaching performance levels with clear clinical utility. Most algorithms require four input magnetic resonance imaging (MRI) modalities, typically T1-weighted images with and without contrast enhancement, T2-weighted images, and FLAIR images. However, some of these sequences are often missing in clinical practice, e.g., because of time constraints and/or image artifacts (such as patient motion). Therefore, substituting missing modalities to recover segmentation performance in these scenarios is highly desirable and necessary for the more widespread adoption of such algorithms in clinical routine. In this work, we report the set-up of the Brain MR Image Synthesis Benchmark (BraSyn), organized in conjunction with the Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2023. The objective of the challenge is to benchmark image synthesis methods that realistically synthesize missing MRI modalities given multiple available images to facilitate automated brain tumor segmentation pipelines. The image dataset is multi-modal and diverse, created in collaboration with various hospitals and research institutions.
The Brain Tumor Segmentation Challenge 2023: Local Synthesis of Healthy Brain Tissue via Inpainting
May 15, 2023



Abstract:A myriad of algorithms for the automatic analysis of brain MR images is available to support clinicians in their decision-making. For brain tumor patients, the image acquisition time series typically starts with a scan that is already pathological. This poses problems, as many algorithms are designed to analyze healthy brains and provide no guarantees for images featuring lesions. Examples include but are not limited to algorithms for brain anatomy parcellation, tissue segmentation, and brain extraction. To solve this dilemma, we introduce the BraTS 2023 inpainting challenge. Here, the participants' task is to explore inpainting techniques to synthesize healthy brain scans from lesioned ones. The following manuscript contains the task formulation, dataset, and submission procedure. Later it will be updated to summarize the findings of the challenge. The challenge is organized as part of the BraTS 2023 challenge hosted at the MICCAI 2023 conference in Vancouver, Canada.
Gene-SGAN: a method for discovering disease subtypes with imaging and genetic signatures via multi-view weakly-supervised deep clustering
Jan 25, 2023


Abstract:Disease heterogeneity has been a critical challenge for precision diagnosis and treatment, especially in neurologic and neuropsychiatric diseases. Many diseases can display multiple distinct brain phenotypes across individuals, potentially reflecting disease subtypes that can be captured using MRI and machine learning methods. However, biological interpretability and treatment relevance are limited if the derived subtypes are not associated with genetic drivers or susceptibility factors. Herein, we describe Gene-SGAN - a multi-view, weakly-supervised deep clustering method - which dissects disease heterogeneity by jointly considering phenotypic and genetic data, thereby conferring genetic correlations to the disease subtypes and associated endophenotypic signatures. We first validate the generalizability, interpretability, and robustness of Gene-SGAN in semi-synthetic experiments. We then demonstrate its application to real multi-site datasets from 28,858 individuals, deriving subtypes of Alzheimer's disease and brain endophenotypes associated with hypertension, from MRI and SNP data. Derived brain phenotypes displayed significant differences in neuroanatomical patterns, genetic determinants, biological and clinical biomarkers, indicating potentially distinct underlying neuropathologic processes, genetic drivers, and susceptibility factors. Overall, Gene-SGAN is broadly applicable to disease subtyping and endophenotype discovery, and is herein tested on disease-related, genetically-driven neuroimaging phenotypes.
Deep Learning Based Detection of Enlarged Perivascular Spaces on Brain MRI
Sep 27, 2022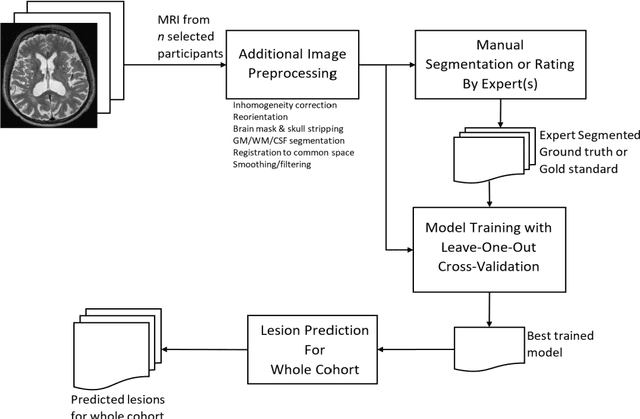
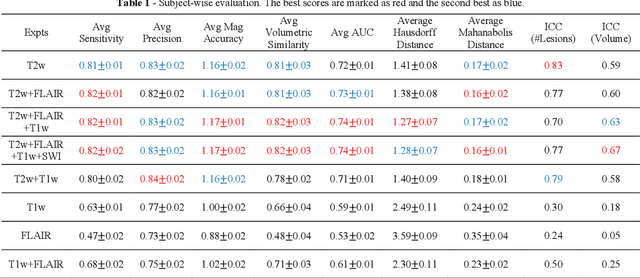
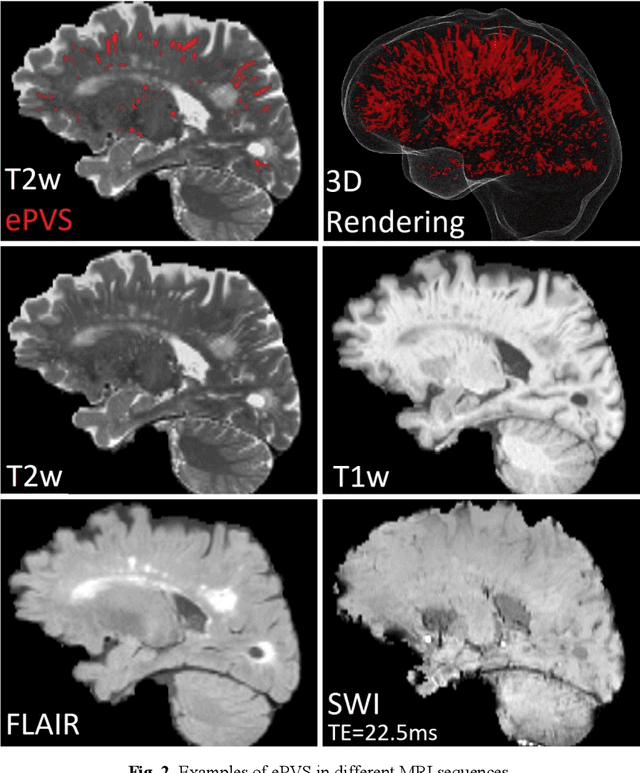
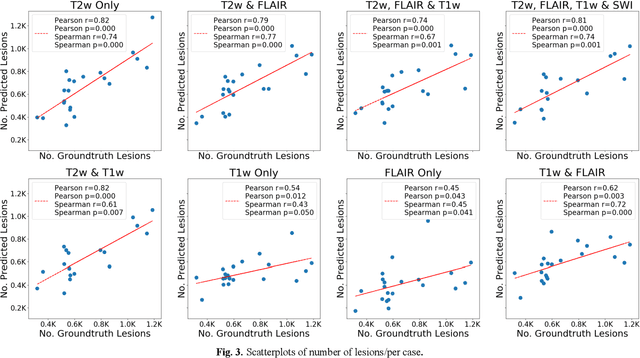
Abstract:Deep learning has been demonstrated effective in many neuroimaging applications. However, in many scenarios the number of imaging sequences capturing information related to small vessel disease lesions is insufficient to support data-driven techniques. Additionally, cohort-based studies may not always have the optimal or essential imaging sequences for accurate lesion detection. Therefore, it is necessary to determine which of these imaging sequences are essential for accurate detection. In this study we aimed to find the optimal combination of magnetic resonance imaging (MRI) sequences for deep learning-based detection of enlarged perivascular spaces (ePVS). To this end, we implemented an effective light-weight U-Net adapted for ePVS detection and comprehensively investigated different combinations of information from susceptibility weighted imaging (SWI), fluid-attenuated inversion recovery (FLAIR), T1-weighted (T1w) and T2-weighted (T2w) MRI sequences. We conclude that T2w MRI is the most important for accurate ePVS detection, and the incorporation of SWI, FLAIR and T1w MRI in the deep neural network could make insignificant improvements in accuracy.
Applications of Generative Adversarial Networks in Neuroimaging and Clinical Neuroscience
Jun 14, 2022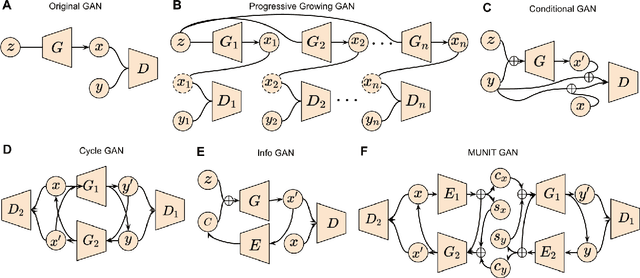
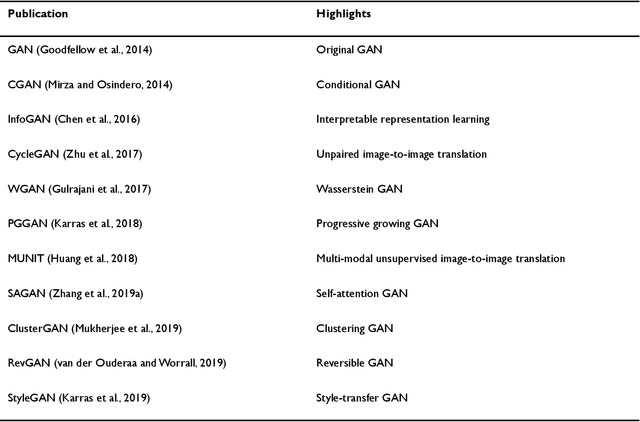
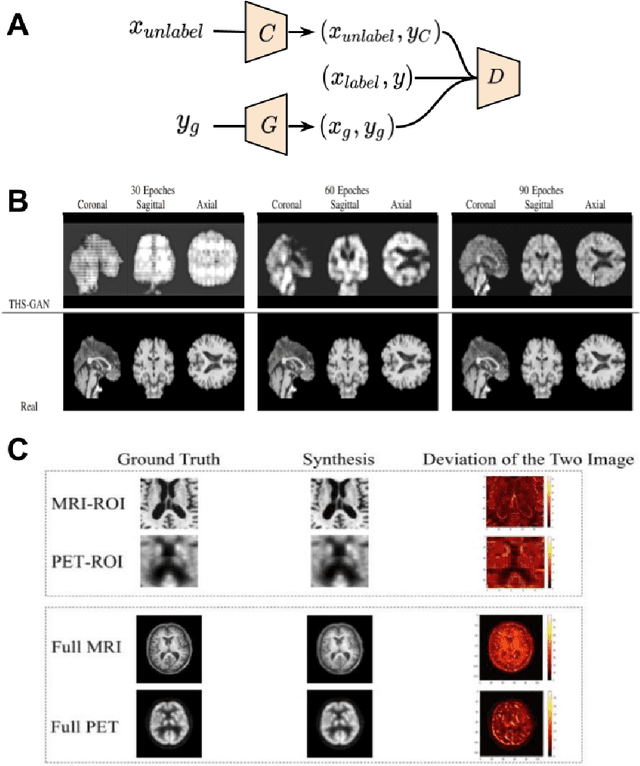
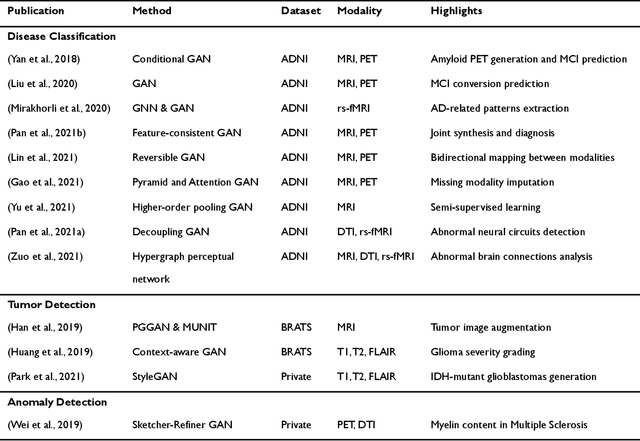
Abstract:Generative adversarial networks (GANs) are one powerful type of deep learning models that have been successfully utilized in numerous fields. They belong to a broader family called generative methods, which generate new data with a probabilistic model by learning sample distribution from real examples. In the clinical context, GANs have shown enhanced capabilities in capturing spatially complex, nonlinear, and potentially subtle disease effects compared to traditional generative methods. This review appraises the existing literature on the applications of GANs in imaging studies of various neurological conditions, including Alzheimer's disease, brain tumors, brain aging, and multiple sclerosis. We provide an intuitive explanation of various GAN methods for each application and further discuss the main challenges, open questions, and promising future directions of leveraging GANs in neuroimaging. We aim to bridge the gap between advanced deep learning methods and neurology research by highlighting how GANs can be leveraged to support clinical decision making and contribute to a better understanding of the structural and functional patterns of brain diseases.
Machine Learning Models Are Not Necessarily Biased When Constructed Properly: Evidence from Neuroimaging Studies
May 26, 2022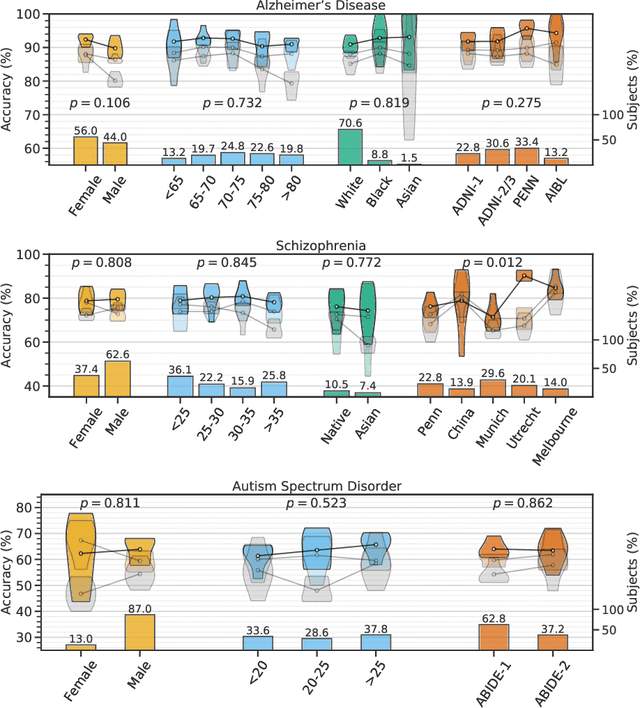
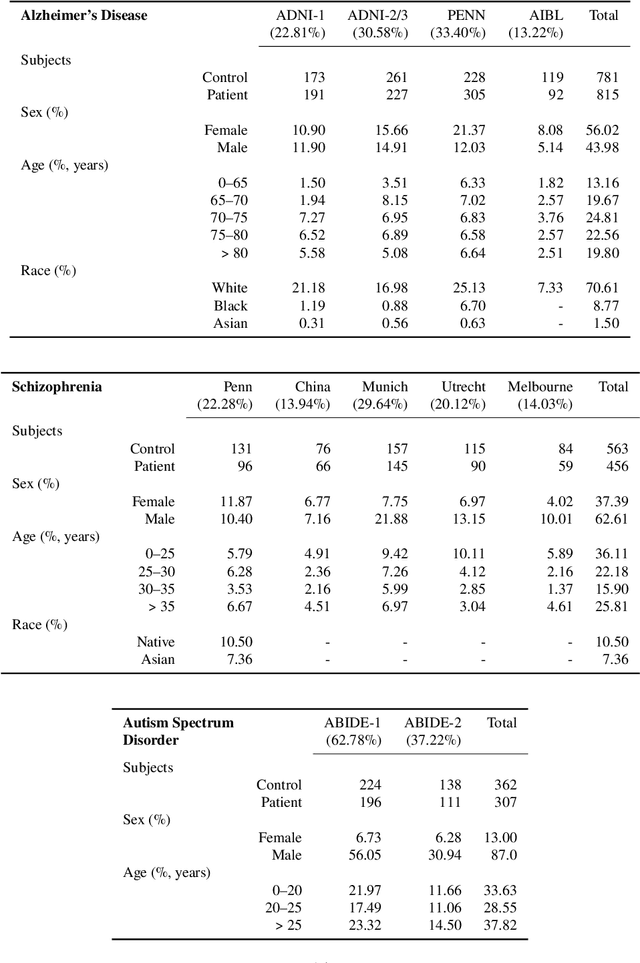
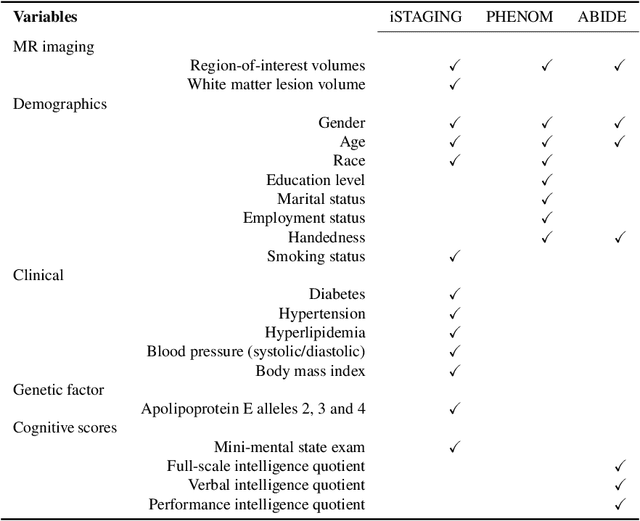
Abstract:Despite the great promise that machine learning has offered in many fields of medicine, it has also raised concerns about potential biases and poor generalization across genders, age distributions, races and ethnicities, hospitals, and data acquisition equipment and protocols. In the current study, and in the context of three brain diseases, we provide experimental data which support that when properly trained, machine learning models can generalize well across diverse conditions and do not suffer from biases. Specifically, by using multi-study magnetic resonance imaging consortia for diagnosing Alzheimer's disease, schizophrenia, and autism spectrum disorder, we find that, the accuracy of well-trained models is consistent across different subgroups pertaining to attributes such as gender, age, and racial groups, as also different clinical studies. We find that models that incorporate multi-source data from demographic, clinical, genetic factors and cognitive scores are also unbiased. These models have better predictive accuracy across subgroups than those trained only with structural measures in some cases but there are also situations when these additional features do not help.
Surreal-GAN:Semi-Supervised Representation Learning via GAN for uncovering heterogeneous disease-related imaging patterns
May 09, 2022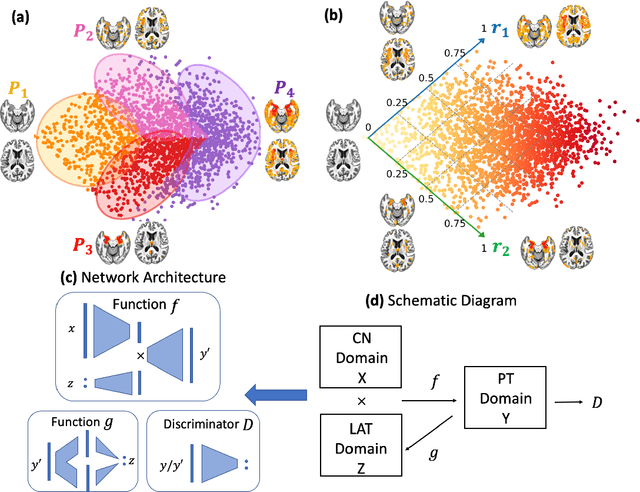
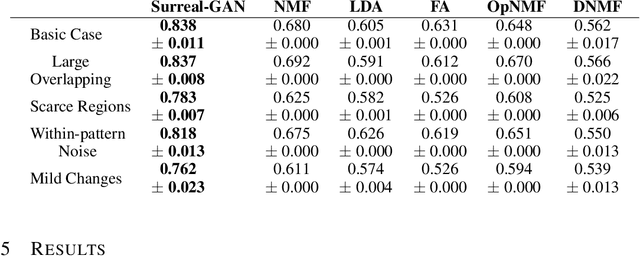
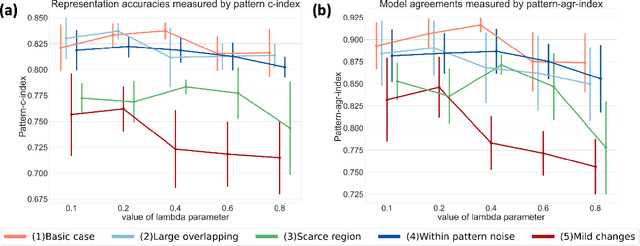
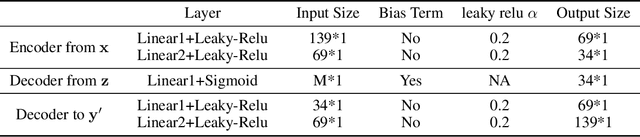
Abstract:A plethora of machine learning methods have been applied to imaging data, enabling the construction of clinically relevant imaging signatures of neurological and neuropsychiatric diseases. Oftentimes, such methods don't explicitly model the heterogeneity of disease effects, or approach it via nonlinear models that are not interpretable. Moreover, unsupervised methods may parse heterogeneity that is driven by nuisance confounding factors that affect brain structure or function, rather than heterogeneity relevant to a pathology of interest. On the other hand, semi-supervised clustering methods seek to derive a dichotomous subtype membership, ignoring the truth that disease heterogeneity spatially and temporally extends along a continuum. To address the aforementioned limitations, herein, we propose a novel method, termed Surreal-GAN (Semi-SUpeRvised ReprEsentAtion Learning via GAN). Using cross-sectional imaging data, Surreal-GAN dissects underlying disease-related heterogeneity under the principle of semi-supervised clustering (cluster mappings from normal control to patient), proposes a continuously dimensional representation, and infers the disease severity of patients at individual level along each dimension. The model first learns a transformation function from normal control (CN) domain to the patient (PT) domain with latent variables controlling transformation directions. An inverse mapping function together with regularization on function continuity, pattern orthogonality and monotonicity was also imposed to make sure that the transformation function captures necessarily meaningful imaging patterns with clinical significance. We first validated the model through extensive semi-synthetic experiments, and then demonstrate its potential in capturing biologically plausible imaging patterns in Alzheimer's disease (AD).
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge