Christopher P. Bridge
DBT-DINO: Towards Foundation model based analysis of Digital Breast Tomosynthesis
Dec 15, 2025
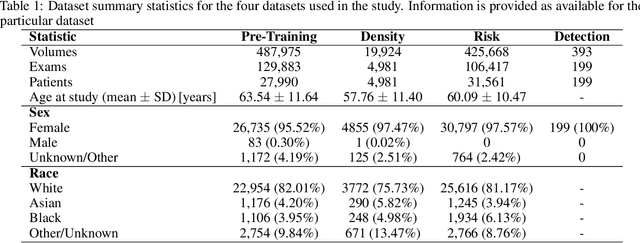
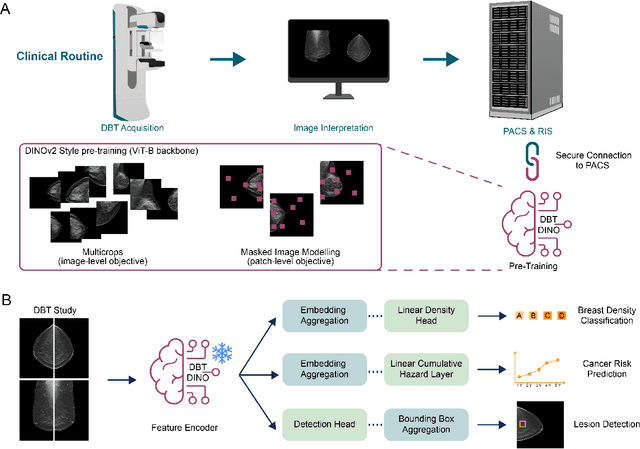
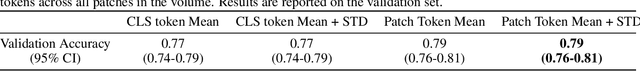
Abstract:Foundation models have shown promise in medical imaging but remain underexplored for three-dimensional imaging modalities. No foundation model currently exists for Digital Breast Tomosynthesis (DBT), despite its use for breast cancer screening. To develop and evaluate a foundation model for DBT (DBT-DINO) across multiple clinical tasks and assess the impact of domain-specific pre-training. Self-supervised pre-training was performed using the DINOv2 methodology on over 25 million 2D slices from 487,975 DBT volumes from 27,990 patients. Three downstream tasks were evaluated: (1) breast density classification using 5,000 screening exams; (2) 5-year risk of developing breast cancer using 106,417 screening exams; and (3) lesion detection using 393 annotated volumes. For breast density classification, DBT-DINO achieved an accuracy of 0.79 (95\% CI: 0.76--0.81), outperforming both the MetaAI DINOv2 baseline (0.73, 95\% CI: 0.70--0.76, p<.001) and DenseNet-121 (0.74, 95\% CI: 0.71--0.76, p<.001). For 5-year breast cancer risk prediction, DBT-DINO achieved an AUROC of 0.78 (95\% CI: 0.76--0.80) compared to DINOv2's 0.76 (95\% CI: 0.74--0.78, p=.57). For lesion detection, DINOv2 achieved a higher average sensitivity of 0.67 (95\% CI: 0.60--0.74) compared to DBT-DINO with 0.62 (95\% CI: 0.53--0.71, p=.60). DBT-DINO demonstrated better performance on cancerous lesions specifically with a detection rate of 78.8\% compared to Dinov2's 77.3\%. Using a dataset of unprecedented size, we developed DBT-DINO, the first foundation model for DBT. DBT-DINO demonstrated strong performance on breast density classification and cancer risk prediction. However, domain-specific pre-training showed variable benefits on the detection task, with ImageNet baseline outperforming DBT-DINO on general lesion detection, indicating that localized detection tasks require further methodological development.
Scalable Drift Monitoring in Medical Imaging AI
Oct 17, 2024



Abstract:The integration of artificial intelligence (AI) into medical imaging has advanced clinical diagnostics but poses challenges in managing model drift and ensuring long-term reliability. To address these challenges, we develop MMC+, an enhanced framework for scalable drift monitoring, building upon the CheXstray framework that introduced real-time drift detection for medical imaging AI models using multi-modal data concordance. This work extends the original framework's methodologies, providing a more scalable and adaptable solution for real-world healthcare settings and offers a reliable and cost-effective alternative to continuous performance monitoring addressing limitations of both continuous and periodic monitoring methods. MMC+ introduces critical improvements to the original framework, including more robust handling of diverse data streams, improved scalability with the integration of foundation models like MedImageInsight for high-dimensional image embeddings without site-specific training, and the introduction of uncertainty bounds to better capture drift in dynamic clinical environments. Validated with real-world data from Massachusetts General Hospital during the COVID-19 pandemic, MMC+ effectively detects significant data shifts and correlates them with model performance changes. While not directly predicting performance degradation, MMC+ serves as an early warning system, indicating when AI systems may deviate from acceptable performance bounds and enabling timely interventions. By emphasizing the importance of monitoring diverse data streams and evaluating data shifts alongside model performance, this work contributes to the broader adoption and integration of AI solutions in clinical settings.
Deep Learning-based Prediction of Breast Cancer Tumor and Immune Phenotypes from Histopathology
Apr 25, 2024



Abstract:The interactions between tumor cells and the tumor microenvironment (TME) dictate therapeutic efficacy of radiation and many systemic therapies in breast cancer. However, to date, there is not a widely available method to reproducibly measure tumor and immune phenotypes for each patient's tumor. Given this unmet clinical need, we applied multiple instance learning (MIL) algorithms to assess activity of ten biologically relevant pathways from the hematoxylin and eosin (H&E) slide of primary breast tumors. We employed different feature extraction approaches and state-of-the-art model architectures. Using binary classification, our models attained area under the receiver operating characteristic (AUROC) scores above 0.70 for nearly all gene expression pathways and on some cases, exceeded 0.80. Attention maps suggest that our trained models recognize biologically relevant spatial patterns of cell sub-populations from H&E. These efforts represent a first step towards developing computational H&E biomarkers that reflect facets of the TME and hold promise for augmenting precision oncology.
Automatic classification of prostate MR series type using image content and metadata
Apr 16, 2024


Abstract:With the wealth of medical image data, efficient curation is essential. Assigning the sequence type to magnetic resonance images is necessary for scientific studies and artificial intelligence-based analysis. However, incomplete or missing metadata prevents effective automation. We therefore propose a deep-learning method for classification of prostate cancer scanning sequences based on a combination of image data and DICOM metadata. We demonstrate superior results compared to metadata or image data alone, and make our code publicly available at https://github.com/deepakri201/DICOMScanClassification.
Is Open-Source There Yet? A Comparative Study on Commercial and Open-Source LLMs in Their Ability to Label Chest X-Ray Reports
Feb 19, 2024



Abstract:Introduction: With the rapid advances in large language models (LLMs), there have been numerous new open source as well as commercial models. While recent publications have explored GPT-4 in its application to extracting information of interest from radiology reports, there has not been a real-world comparison of GPT-4 to different leading open-source models. Materials and Methods: Two different and independent datasets were used. The first dataset consists of 540 chest x-ray reports that were created at the Massachusetts General Hospital between July 2019 and July 2021. The second dataset consists of 500 chest x-ray reports from the ImaGenome dataset. We then compared the commercial models GPT-3.5 Turbo and GPT-4 from OpenAI to the open-source models Mistral-7B, Mixtral-8x7B, Llama2-13B, Llama2-70B, QWEN1.5-72B and CheXbert and CheXpert-labeler in their ability to accurately label the presence of multiple findings in x-ray text reports using different prompting techniques. Results: On the ImaGenome dataset, the best performing open-source model was Llama2-70B with micro F1-scores of 0.972 and 0.970 for zero- and few-shot prompts, respectively. GPT-4 achieved micro F1-scores of 0.975 and 0.984, respectively. On the institutional dataset, the best performing open-source model was QWEN1.5-72B with micro F1-scores of 0.952 and 0.965 for zero- and few-shot prompting, respectively. GPT-4 achieved micro F1-scores of 0.975 and 0.973, respectively. Conclusion: In this paper, we show that while GPT-4 is superior to open-source models in zero-shot report labeling, the implementation of few-shot prompting can bring open-source models on par with GPT-4. This shows that open-source models could be a performant and privacy preserving alternative to GPT-4 for the task of radiology report classification.
A generalized framework to predict continuous scores from medical ordinal labels
May 30, 2023Abstract:Many variables of interest in clinical medicine, like disease severity, are recorded using discrete ordinal categories such as normal/mild/moderate/severe. These labels are used to train and evaluate disease severity prediction models. However, ordinal categories represent a simplification of an underlying continuous severity spectrum. Using continuous scores instead of ordinal categories is more sensitive to detecting small changes in disease severity over time. Here, we present a generalized framework that accurately predicts continuously valued variables using only discrete ordinal labels during model development. We found that for three clinical prediction tasks, models that take the ordinal relationship of the training labels into account outperformed conventional multi-class classification models. Particularly the continuous scores generated by ordinal classification and regression models showed a significantly higher correlation with expert rankings of disease severity and lower mean squared errors compared to the multi-class classification models. Furthermore, the use of MC dropout significantly improved the ability of all evaluated deep learning approaches to predict continuously valued scores that truthfully reflect the underlying continuous target variable. We showed that accurate continuously valued predictions can be generated even if the model development only involves discrete ordinal labels. The novel framework has been validated on three different clinical prediction tasks and has proven to bridge the gap between discrete ordinal labels and the underlying continuously valued variables.
Improving the repeatability of deep learning models with Monte Carlo dropout
Feb 15, 2022
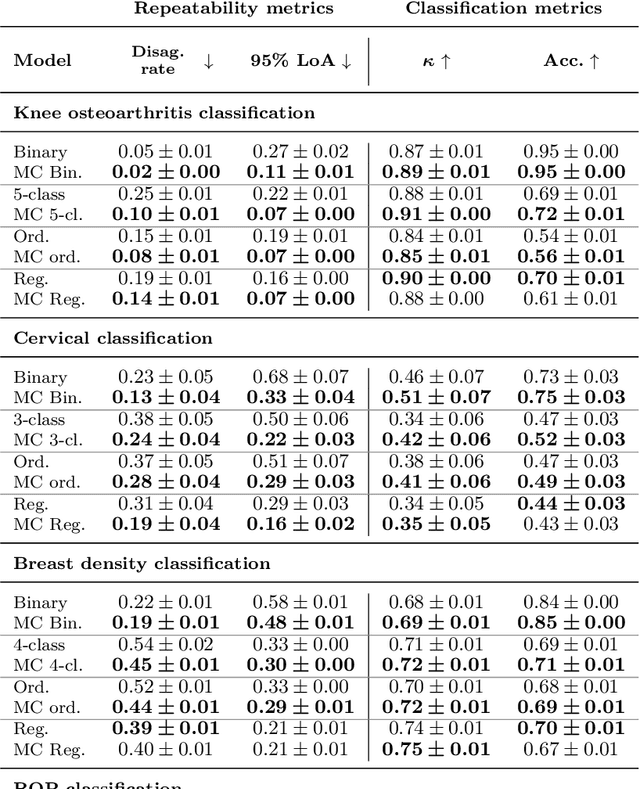
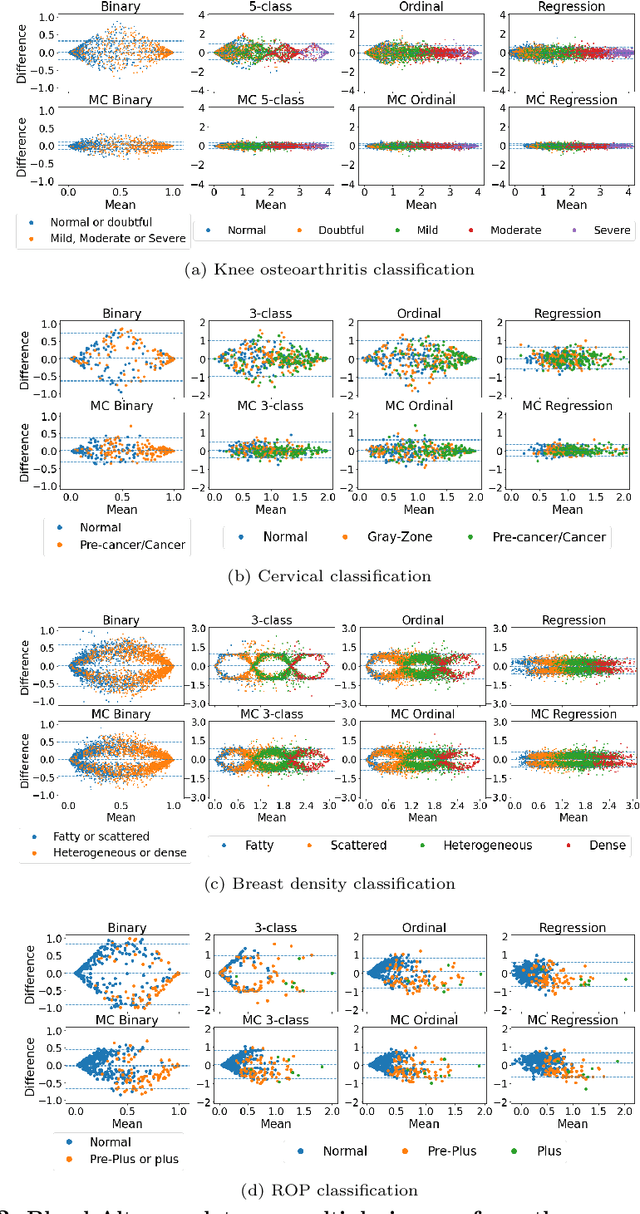
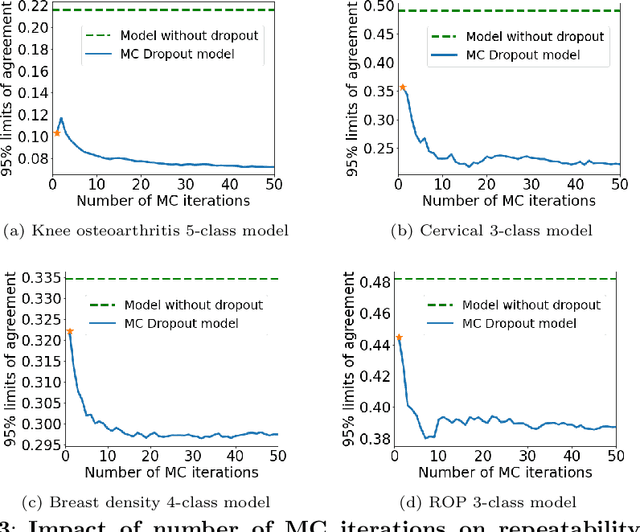
Abstract:The integration of artificial intelligence into clinical workflows requires reliable and robust models. Repeatability is a key attribute of model robustness. Repeatable models output predictions with low variation during independent tests carried out under similar conditions. During model development and evaluation, much attention is given to classification performance while model repeatability is rarely assessed, leading to the development of models that are unusable in clinical practice. In this work, we evaluate the repeatability of four model types (binary classification, multi-class classification, ordinal classification, and regression) on images that were acquired from the same patient during the same visit. We study the performance of binary, multi-class, ordinal, and regression models on four medical image classification tasks from public and private datasets: knee osteoarthritis, cervical cancer screening, breast density estimation, and retinopathy of prematurity. Repeatability is measured and compared on ResNet and DenseNet architectures. Moreover, we assess the impact of sampling Monte Carlo dropout predictions at test time on classification performance and repeatability. Leveraging Monte Carlo predictions significantly increased repeatability for all tasks on the binary, multi-class, and ordinal models leading to an average reduction of the 95\% limits of agreement by 16% points and of the disagreement rate by 7% points. The classification accuracy improved in most settings along with the repeatability. Our results suggest that beyond about 20 Monte Carlo iterations, there is no further gain in repeatability. In addition to the higher test-retest agreement, Monte Carlo predictions were better calibrated which leads to output probabilities reflecting more accurately the true likelihood of being correctly classified.
Monte Carlo dropout increases model repeatability
Nov 12, 2021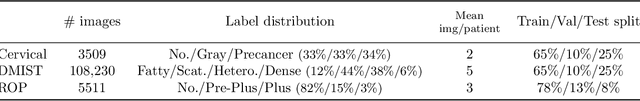
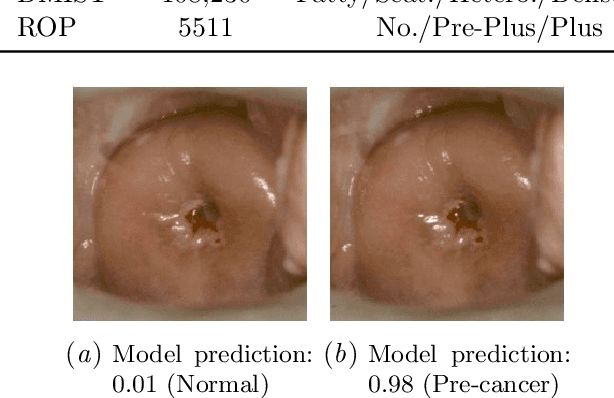
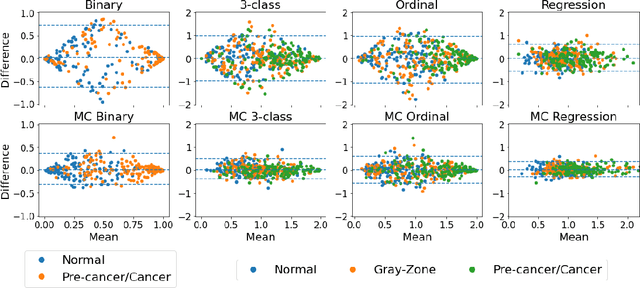
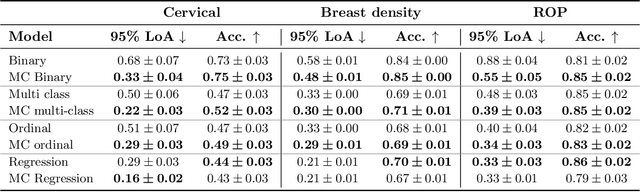
Abstract:The integration of artificial intelligence into clinical workflows requires reliable and robust models. Among the main features of robustness is repeatability. Much attention is given to classification performance without assessing the model repeatability, leading to the development of models that turn out to be unusable in practice. In this work, we evaluate the repeatability of four model types on images from the same patient that were acquired during the same visit. We study the performance of binary, multi-class, ordinal, and regression models on three medical image analysis tasks: cervical cancer screening, breast density estimation, and retinopathy of prematurity classification. Moreover, we assess the impact of sampling Monte Carlo dropout predictions at test time on classification performance and repeatability. Leveraging Monte Carlo predictions significantly increased repeatability for all tasks on the binary, multi-class, and ordinal models leading to an average reduction of the 95% limits of agreement by 17% points.
Highdicom: A Python library for standardized encoding of image annotations and machine learning model outputs in pathology and radiology
Jun 14, 2021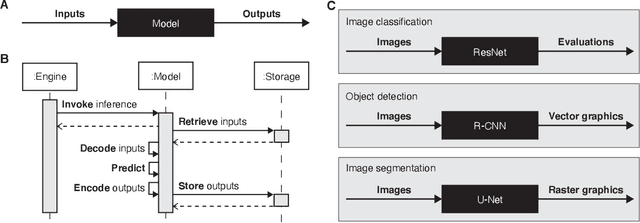
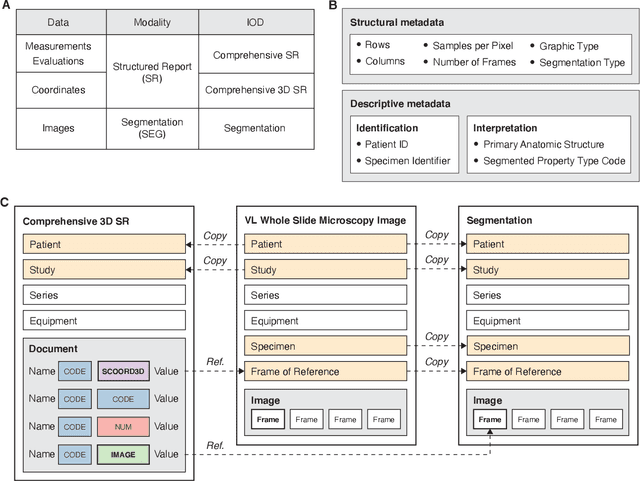
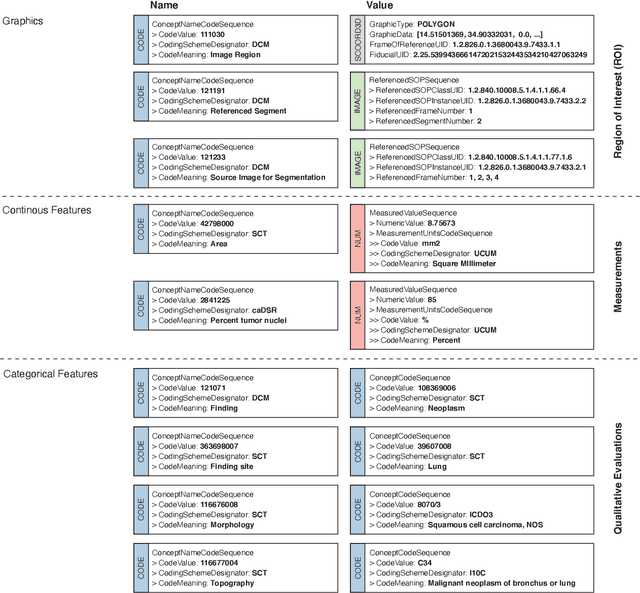
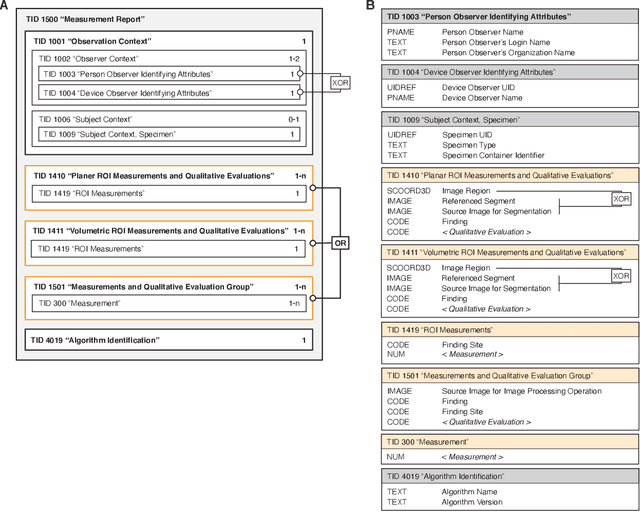
Abstract:Machine learning is revolutionizing image-based diagnostics in pathology and radiology. ML models have shown promising results in research settings, but their lack of interoperability has been a major barrier for clinical integration and evaluation. The DICOM a standard specifies Information Object Definitions and Services for the representation and communication of digital images and related information, including image-derived annotations and analysis results. However, the complexity of the standard represents an obstacle for its adoption in the ML community and creates a need for software libraries and tools that simplify working with data sets in DICOM format. Here we present the highdicom library, which provides a high-level application programming interface for the Python programming language that abstracts low-level details of the standard and enables encoding and decoding of image-derived information in DICOM format in a few lines of Python code. The highdicom library ties into the extensive Python ecosystem for image processing and machine learning. Simultaneously, by simplifying creation and parsing of DICOM-compliant files, highdicom achieves interoperability with the medical imaging systems that hold the data used to train and run ML models, and ultimately communicate and store model outputs for clinical use. We demonstrate through experiments with slide microscopy and computed tomography imaging, that, by bridging these two ecosystems, highdicom enables developers to train and evaluate state-of-the-art ML models in pathology and radiology while remaining compliant with the DICOM standard and interoperable with clinical systems at all stages. To promote standardization of ML research and streamline the ML model development and deployment process, we made the library available free and open-source.
Addressing catastrophic forgetting for medical domain expansion
Mar 24, 2021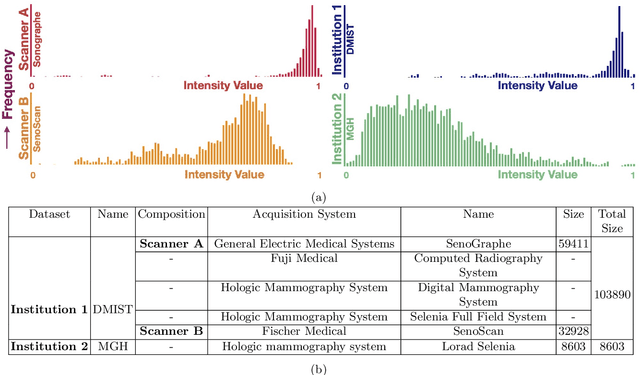
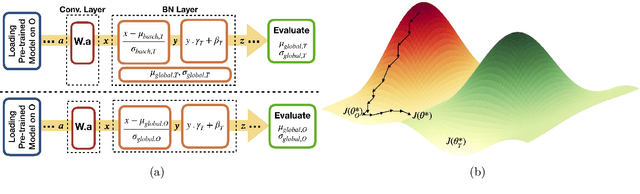
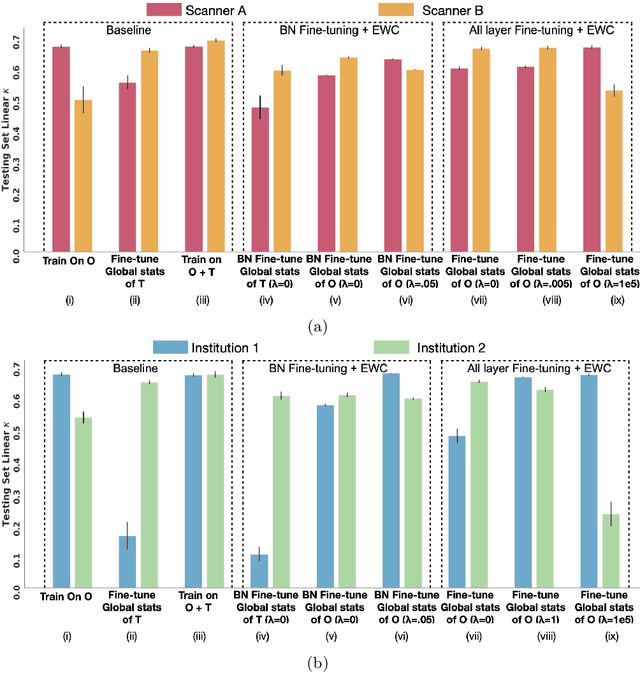
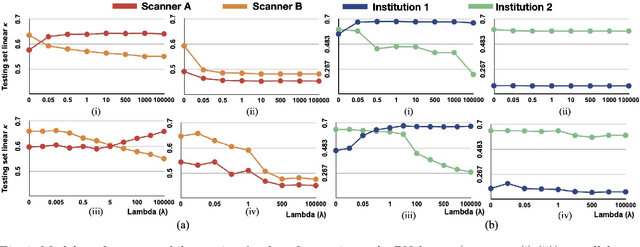
Abstract:Model brittleness is a key concern when deploying deep learning models in real-world medical settings. A model that has high performance at one institution may suffer a significant decline in performance when tested at other institutions. While pooling datasets from multiple institutions and retraining may provide a straightforward solution, it is often infeasible and may compromise patient privacy. An alternative approach is to fine-tune the model on subsequent institutions after training on the original institution. Notably, this approach degrades model performance at the original institution, a phenomenon known as catastrophic forgetting. In this paper, we develop an approach to address catastrophic forget-ting based on elastic weight consolidation combined with modulation of batch normalization statistics under two scenarios: first, for expanding the domain from one imaging system's data to another imaging system's, and second, for expanding the domain from a large multi-institutional dataset to another single institution dataset. We show that our approach outperforms several other state-of-the-art approaches and provide theoretical justification for the efficacy of batch normalization modulation. The results of this study are generally applicable to the deployment of any clinical deep learning model which requires domain expansion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge