Mannudeep Kalra
Scalable Drift Monitoring in Medical Imaging AI
Oct 17, 2024



Abstract:The integration of artificial intelligence (AI) into medical imaging has advanced clinical diagnostics but poses challenges in managing model drift and ensuring long-term reliability. To address these challenges, we develop MMC+, an enhanced framework for scalable drift monitoring, building upon the CheXstray framework that introduced real-time drift detection for medical imaging AI models using multi-modal data concordance. This work extends the original framework's methodologies, providing a more scalable and adaptable solution for real-world healthcare settings and offers a reliable and cost-effective alternative to continuous performance monitoring addressing limitations of both continuous and periodic monitoring methods. MMC+ introduces critical improvements to the original framework, including more robust handling of diverse data streams, improved scalability with the integration of foundation models like MedImageInsight for high-dimensional image embeddings without site-specific training, and the introduction of uncertainty bounds to better capture drift in dynamic clinical environments. Validated with real-world data from Massachusetts General Hospital during the COVID-19 pandemic, MMC+ effectively detects significant data shifts and correlates them with model performance changes. While not directly predicting performance degradation, MMC+ serves as an early warning system, indicating when AI systems may deviate from acceptable performance bounds and enabling timely interventions. By emphasizing the importance of monitoring diverse data streams and evaluating data shifts alongside model performance, this work contributes to the broader adoption and integration of AI solutions in clinical settings.
Fact-Checking of AI-Generated Reports
Jul 27, 2023Abstract:With advances in generative artificial intelligence (AI), it is now possible to produce realistic-looking automated reports for preliminary reads of radiology images. This can expedite clinical workflows, improve accuracy and reduce overall costs. However, it is also well-known that such models often hallucinate, leading to false findings in the generated reports. In this paper, we propose a new method of fact-checking of AI-generated reports using their associated images. Specifically, the developed examiner differentiates real and fake sentences in reports by learning the association between an image and sentences describing real or potentially fake findings. To train such an examiner, we first created a new dataset of fake reports by perturbing the findings in the original ground truth radiology reports associated with images. Text encodings of real and fake sentences drawn from these reports are then paired with image encodings to learn the mapping to real/fake labels. The utility of such an examiner is demonstrated for verifying automatically generated reports by detecting and removing fake sentences. Future generative AI approaches can use the resulting tool to validate their reports leading to a more responsible use of AI in expediting clinical workflows.
Soft Activation Mapping of Lung Nodules in Low-Dose CT images
Oct 30, 2018
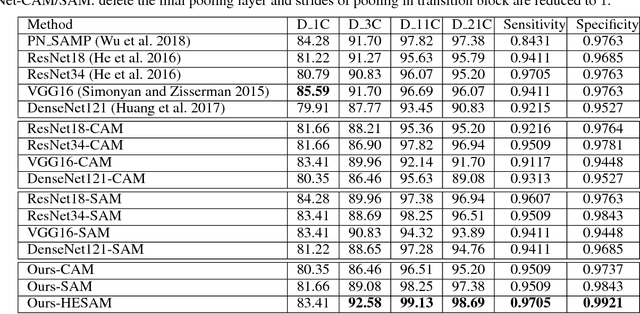
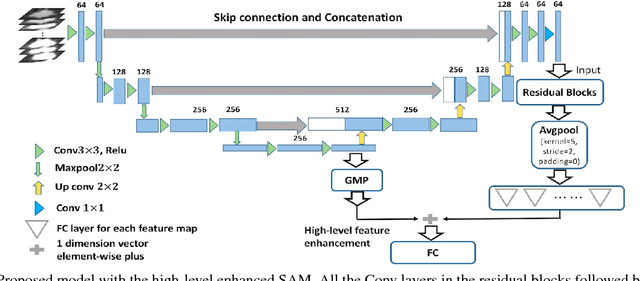
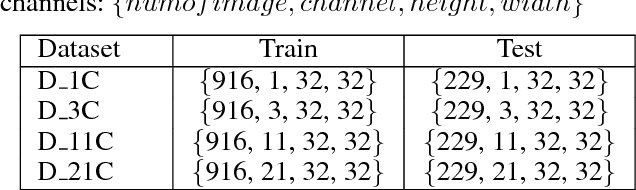
Abstract:As a popular deep learning model, the convolutional neural network (CNN) has produced promising results in analyzing lung nodules and tumors in low-dose CT images. However, this approach still suffers from the lack of labeled data, which is a major challenge for further improvement in the screening and diagnostic performance of CNN. Accurate localization and characterization of nodules provides crucial pathological clues, especially relevant size, attenuation, shape, margins, and growth or stability of lesions, with which the sensitivity and specificity of detection and classification can be increased. To address this challenge, in this paper we develop a soft activation mapping (SAM) to enable fine-grained lesion analysis with a CNN so that it can access rich radiomics features. By combining high-level convolutional features with SAM, we further propose a high-level feature enhancement scheme to localize lesions precisely from multiple CT slices, which helps alleviate overfitting without any additional data augmentation. Experiments on the LIDC-IDRI benchmark dataset indicate that our proposed approach achieves a state-of-the-art predictive performance, reducing the false positive rate. Moreover, the SAM method focuses on irregular margins which are often linked to malignancy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge