Christoph Lippert
Digital Health & Machine Learning, Hasso Plattner Institute, University of Potsdam, Germany, Hasso Plattner Institute for Digital Health at Mount Sinai, Icahn School of Medicine at Mount Sinai, NYC, USA
Towards Visually Explaining Statistical Tests with Applications in Biomedical Imaging
Jan 20, 2026Abstract:Deep neural two-sample tests have recently shown strong power for detecting distributional differences between groups, yet their black-box nature limits interpretability and practical adoption in biomedical analysis. Moreover, most existing post-hoc explainability methods rely on class labels, making them unsuitable for label-free statistical testing settings. We propose an explainable deep statistical testing framework that augments deep two-sample tests with sample-level and feature-level explanations, revealing which individual samples and which input features drive statistically significant group differences. Our method highlights which image regions and which individual samples contribute most to the detected group difference, providing spatial and instance-wise insight into the test's decision. Applied to biomedical imaging data, the proposed framework identifies influential samples and highlights anatomically meaningful regions associated with disease-related variation. This work bridges statistical inference and explainable AI, enabling interpretable, label-free population analysis in medical imaging.
AnatomiX, an Anatomy-Aware Grounded Multimodal Large Language Model for Chest X-Ray Interpretation
Jan 06, 2026Abstract:Multimodal medical large language models have shown impressive progress in chest X-ray interpretation but continue to face challenges in spatial reasoning and anatomical understanding. Although existing grounding techniques improve overall performance, they often fail to establish a true anatomical correspondence, resulting in incorrect anatomical understanding in the medical domain. To address this gap, we introduce AnatomiX, a multitask multimodal large language model explicitly designed for anatomically grounded chest X-ray interpretation. Inspired by the radiological workflow, AnatomiX adopts a two stage approach: first, it identifies anatomical structures and extracts their features, and then leverages a large language model to perform diverse downstream tasks such as phrase grounding, report generation, visual question answering, and image understanding. Extensive experiments across multiple benchmarks demonstrate that AnatomiX achieves superior anatomical reasoning and delivers over 25% improvement in performance on anatomy grounding, phrase grounding, grounded diagnosis and grounded captioning tasks compared to existing approaches. Code and pretrained model are available at https://github.com/aneesurhashmi/anatomix
JAPAN: Joint Adaptive Prediction Areas with Normalising-Flows
May 29, 2025
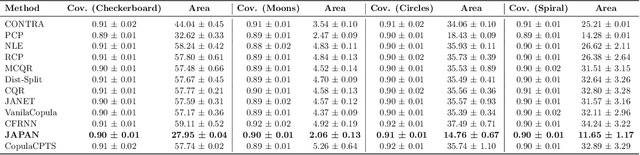


Abstract:Conformal prediction provides a model-agnostic framework for uncertainty quantification with finite-sample validity guarantees, making it an attractive tool for constructing reliable prediction sets. However, existing approaches commonly rely on residual-based conformity scores, which impose geometric constraints and struggle when the underlying distribution is multimodal. In particular, they tend to produce overly conservative prediction areas centred around the mean, often failing to capture the true shape of complex predictive distributions. In this work, we introduce JAPAN (Joint Adaptive Prediction Areas with Normalising-Flows), a conformal prediction framework that uses density-based conformity scores. By leveraging flow-based models, JAPAN estimates the (predictive) density and constructs prediction areas by thresholding on the estimated density scores, enabling compact, potentially disjoint, and context-adaptive regions that retain finite-sample coverage guarantees. We theoretically motivate the efficiency of JAPAN and empirically validate it across multivariate regression and forecasting tasks, demonstrating good calibration and tighter prediction areas compared to existing baselines. We also provide several \emph{extensions} adding flexibility to our proposed framework.
Token Cropr: Faster ViTs for Quite a Few Tasks
Dec 01, 2024



Abstract:The adoption of Vision Transformers (ViTs) in resource-constrained applications necessitates improvements in inference throughput. To this end several token pruning and merging approaches have been proposed that improve efficiency by successively reducing the number of tokens. However, it remains an open problem to design a token reduction method that is fast, maintains high performance, and is applicable to various vision tasks. In this work, we present a token pruner that uses auxiliary prediction heads that learn to select tokens end-to-end based on task relevance. These auxiliary heads can be removed after training, leading to throughput close to that of a random pruner. We evaluate our method on image classification, semantic segmentation, object detection, and instance segmentation, and show speedups of 1.5 to 4x with small drops in performance. As a best case, on the ADE20k semantic segmentation benchmark, we observe a 2x speedup relative to the no-pruning baseline, with a negligible performance penalty of 0.1 median mIoU across 5 seeds.
Conformalised Conditional Normalising Flows for Joint Prediction Regions in time series
Nov 26, 2024
Abstract:Conformal Prediction offers a powerful framework for quantifying uncertainty in machine learning models, enabling the construction of prediction sets with finite-sample validity guarantees. While easily adaptable to non-probabilistic models, applying conformal prediction to probabilistic generative models, such as Normalising Flows is not straightforward. This work proposes a novel method to conformalise conditional normalising flows, specifically addressing the problem of obtaining prediction regions for multi-step time series forecasting. Our approach leverages the flexibility of normalising flows to generate potentially disjoint prediction regions, leading to improved predictive efficiency in the presence of potential multimodal predictive distributions.
Deep Nonparametric Conditional Independence Tests for Images
Nov 09, 2024Abstract:Conditional independence tests (CITs) test for conditional dependence between random variables. As existing CITs are limited in their applicability to complex, high-dimensional variables such as images, we introduce deep nonparametric CITs (DNCITs). The DNCITs combine embedding maps, which extract feature representations of high-dimensional variables, with nonparametric CITs applicable to these feature representations. For the embedding maps, we derive general properties on their parameter estimators to obtain valid DNCITs and show that these properties include embedding maps learned through (conditional) unsupervised or transfer learning. For the nonparametric CITs, appropriate tests are selected and adapted to be applicable to feature representations. Through simulations, we investigate the performance of the DNCITs for different embedding maps and nonparametric CITs under varying confounder dimensions and confounder relationships. We apply the DNCITs to brain MRI scans and behavioral traits, given confounders, of healthy individuals from the UK Biobank (UKB), confirming null results from a number of ambiguous personality neuroscience studies with a larger data set and with our more powerful tests. In addition, in a confounder control study, we apply the DNCITs to brain MRI scans and a confounder set to test for sufficient confounder control, leading to a potential reduction in the confounder dimension under improved confounder control compared to existing state-of-the-art confounder control studies for the UKB. Finally, we provide an R package implementing the DNCITs.
JANET: Joint Adaptive predictioN-region Estimation for Time-series
Jul 08, 2024Abstract:Conformal prediction provides machine learning models with prediction sets that offer theoretical guarantees, but the underlying assumption of exchangeability limits its applicability to time series data. Furthermore, existing approaches struggle to handle multi-step ahead prediction tasks, where uncertainty estimates across multiple future time points are crucial. We propose JANET (Joint Adaptive predictioN-region Estimation for Time-series), a novel framework for constructing conformal prediction regions that are valid for both univariate and multivariate time series. JANET generalises the inductive conformal framework and efficiently produces joint prediction regions with controlled K-familywise error rates, enabling flexible adaptation to specific application needs. Our empirical evaluation demonstrates JANET's superior performance in multi-step prediction tasks across diverse time series datasets, highlighting its potential for reliable and interpretable uncertainty quantification in sequential data.
On the Challenges and Opportunities in Generative AI
Feb 28, 2024Abstract:The field of deep generative modeling has grown rapidly and consistently over the years. With the availability of massive amounts of training data coupled with advances in scalable unsupervised learning paradigms, recent large-scale generative models show tremendous promise in synthesizing high-resolution images and text, as well as structured data such as videos and molecules. However, we argue that current large-scale generative AI models do not sufficiently address several fundamental issues that hinder their widespread adoption across domains. In this work, we aim to identify key unresolved challenges in modern generative AI paradigms that should be tackled to further enhance their capabilities, versatility, and reliability. By identifying these challenges, we aim to provide researchers with valuable insights for exploring fruitful research directions, thereby fostering the development of more robust and accessible generative AI solutions.
MixerFlow for Image Modelling
Oct 25, 2023



Abstract:Normalising flows are statistical models that transform a complex density into a simpler density through the use of bijective transformations enabling both density estimation and data generation from a single model. In the context of image modelling, the predominant choice has been the Glow-based architecture, whereas alternative architectures remain largely unexplored in the research community. In this work, we propose a novel architecture called MixerFlow, based on the MLP-Mixer architecture, further unifying the generative and discriminative modelling architectures. MixerFlow offers an effective mechanism for weight sharing for flow-based models. Our results demonstrate better density estimation on image datasets under a fixed computational budget and scales well as the image resolution increases, making MixeFlow a powerful yet simple alternative to the Glow-based architectures. We also show that MixerFlow provides more informative embeddings than Glow-based architectures.
A Probabilistic Approach to Self-Supervised Learning using Cyclical Stochastic Gradient MCMC
Aug 02, 2023Abstract:In this paper we present a practical Bayesian self-supervised learning method with Cyclical Stochastic Gradient Hamiltonian Monte Carlo (cSGHMC). Within this framework, we place a prior over the parameters of a self-supervised learning model and use cSGHMC to approximate the high dimensional and multimodal posterior distribution over the embeddings. By exploring an expressive posterior over the embeddings, Bayesian self-supervised learning produces interpretable and diverse representations. Marginalizing over these representations yields a significant gain in performance, calibration and out-of-distribution detection on a variety of downstream classification tasks. We provide experimental results on multiple classification tasks on four challenging datasets. Moreover, we demonstrate the effectiveness of the proposed method in out-of-distribution detection using the SVHN and CIFAR-10 datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge