Chaoli Wang
Leveraging Multimodal LLMs for Built Environment and Housing Attribute Assessment from Street-View Imagery
Apr 22, 2026Abstract:We present a novel framework for automatically evaluating building conditions nationwide in the United States by leveraging large language models (LLMs) and Google Street View (GSV) imagery. By fine-tuning Gemma 3 27B on a modest human-labeled dataset, our approach achieves strong alignment with human mean opinion scores (MOS), outperforming even individual raters on SRCC and PLCC relative to the MOS benchmark. To enhance efficiency, we apply knowledge distillation, transferring the capabilities of Gemma 3 27B to a smaller Gemma 3 4B model that achieves comparable performance with a 3x speedup. Further, we distill the knowledge into a CNN-based model (EfficientNetV2-M) and a transformer (SwinV2-B), delivering close performance while achieving a 30x speed gain. Furthermore, we investigate LLMs' capabilities for assessing an extensive list of built environment and housing attributes through a human-AI alignment study and develop a visualization dashboard that integrates LLM assessment outcomes for downstream analysis by homeowners. Our framework offers a flexible and efficient solution for large-scale building condition assessment, enabling high accuracy with minimal human labeling effort.
CODE-GEN: A Human-in-the-Loop RAG-Based Agentic AI System for Multiple-Choice Question Generation
Apr 05, 2026Abstract:We present CODE-GEN, a human-in-the-Loop, retrieval-augmented generation (RAG)-based agentic AI system for generating context-aligned multiple-choice questions to develop student code reasoning and comprehension abilities. CODE-GEN employs an agentic AI architecture in which a Generator agent produces multiple-choice coding comprehension questions aligned with course-specific learning objectives, while a Validator agent independently assesses content quality across seven pedagogical dimensions. Both agents are augmented with specialized tools that enhance computational accuracy and verify code outputs. To evaluate the effectiveness of CODE-GEN, we conducted an evaluation study involving six human subject-matter experts (SMEs) who judged 288 AI-generated questions. The SMEs produced a total of 2,016 human-AI rating pairs, indicating agreement or disagreement with the assessments of Validator, along with 131 instances of qualitative feedback. Analyses of SME judgments show strong system performance, with human-validated success rates ranging from 79.9% to 98.6% across the seven pedagogical dimensions. The analysis of qualitative feedback reveals that CODE-GEN achieves high reliability on dimensions well suited to computational verification and explicit criteria matching, including question clarity, code validity, concept alignment, and correct answer validity. In contrast, human expertise remains essential for dimensions requiring deeper instructional judgment, such as designing pedagogically meaningful distractors and providing high-quality feedback that reinforces understanding. These findings inform the strategic allocation of human and AI effort in AI-assisted educational content generation.
SciVisAgentBench: A Benchmark for Evaluating Scientific Data Analysis and Visualization Agents
Mar 31, 2026Abstract:Recent advances in large language models (LLMs) have enabled agentic systems that translate natural language intent into executable scientific visualization (SciVis) tasks. Despite rapid progress, the community lacks a principled and reproducible benchmark for evaluating these emerging SciVis agents in realistic, multi-step analysis settings. We present SciVisAgentBench, a comprehensive and extensible benchmark for evaluating scientific data analysis and visualization agents. Our benchmark is grounded in a structured taxonomy spanning four dimensions: application domain, data type, complexity level, and visualization operation. It currently comprises 108 expert-crafted cases covering diverse SciVis scenarios. To enable reliable assessment, we introduce a multimodal outcome-centric evaluation pipeline that combines LLM-based judging with deterministic evaluators, including image-based metrics, code checkers, rule-based verifiers, and case-specific evaluators. We also conduct a validity study with 12 SciVis experts to examine the agreement between human and LLM judges. Using this framework, we evaluate representative SciVis agents and general-purpose coding agents to establish initial baselines and reveal capability gaps. SciVisAgentBench is designed as a living benchmark to support systematic comparison, diagnose failure modes, and drive progress in agentic SciVis. The benchmark is available at https://scivisagentbench.github.io/.
Sketch2CT: Multimodal Diffusion for Structure-Aware 3D Medical Volume Generation
Mar 23, 2026Abstract:Diffusion probabilistic models have demonstrated significant potential in generating high-quality, realistic medical images, providing a promising solution to the persistent challenge of data scarcity in the medical field. Nevertheless, producing 3D medical volumes with anatomically consistent structures under multimodal conditions remains a complex and unresolved problem. We introduce Sketch2CT, a multimodal diffusion framework for structure-aware 3D medical volume generation, jointly guided by a user-provided 2D sketch and a textual description that captures 3D geometric semantics. The framework initially generates 3D segmentation masks of the target organ from random noise, conditioned on both modalities. To effectively align and fuse these inputs, we propose two key modules that refine sketch features with localized textual cues and integrate global sketch-text representations. Built upon a capsule-attention backbone, these modules leverage the complementary strengths of sketches and text to produce anatomically accurate organ shapes. The synthesized segmentation masks subsequently guide a latent diffusion model for 3D CT volume synthesis, enabling realistic reconstruction of organ appearances that are consistent with user-defined sketches and descriptions. Extensive experiments on public CT datasets demonstrate that Sketch2CT achieves superior performance in generating multimodal medical volumes. Its controllable, low-cost generation pipeline enables principled, efficient augmentation of medical datasets. Code is available at https://github.com/adlsn/Sketch2CT.
SGI: Structured 2D Gaussians for Efficient and Compact Large Image Representation
Mar 10, 2026Abstract:2D Gaussian Splatting has emerged as a novel image representation technique that can support efficient rendering on low-end devices. However, scaling to high-resolution images requires optimizing and storing millions of unstructured Gaussian primitives independently, leading to slow convergence and redundant parameters. To address this, we propose Structured Gaussian Image (SGI), a compact and efficient framework for representing high-resolution images. SGI decomposes a complex image into multi-scale local spaces defined by a set of seeds. Each seed corresponds to a spatially coherent region and, together with lightweight multi-layer perceptrons (MLPs), generates structured implicit 2D neural Gaussians. This seed-based formulation imposes structural regularity on otherwise unstructured Gaussian primitives, which facilitates entropy-based compression at the seed level to reduce the total storage. However, optimizing seed parameters directly on high-resolution images is a challenging and non-trivial task. Therefore, we designed a multi-scale fitting strategy that refines the seed representation in a coarse-to-fine manner, substantially accelerating convergence. Quantitative and qualitative evaluations demonstrate that SGI achieves up to 7.5x compression over prior non-quantized 2D Gaussian methods and 1.6x over quantized ones, while also delivering 1.6x and 6.5x faster optimization, respectively, without degrading, and often improving, image fidelity. Code is available at https://github.com/zx-pan/SGI.
An Evaluation-Centric Paradigm for Scientific Visualization Agents
Sep 18, 2025
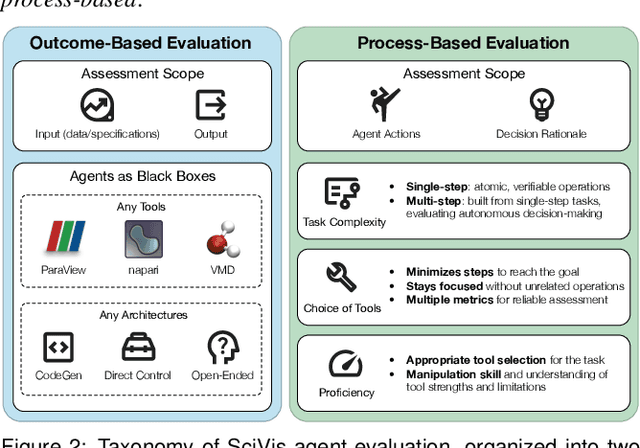
Abstract:Recent advances in multi-modal large language models (MLLMs) have enabled increasingly sophisticated autonomous visualization agents capable of translating user intentions into data visualizations. However, measuring progress and comparing different agents remains challenging, particularly in scientific visualization (SciVis), due to the absence of comprehensive, large-scale benchmarks for evaluating real-world capabilities. This position paper examines the various types of evaluation required for SciVis agents, outlines the associated challenges, provides a simple proof-of-concept evaluation example, and discusses how evaluation benchmarks can facilitate agent self-improvement. We advocate for a broader collaboration to develop a SciVis agentic evaluation benchmark that would not only assess existing capabilities but also drive innovation and stimulate future development in the field.
TexGS-VolVis: Expressive Scene Editing for Volume Visualization via Textured Gaussian Splatting
Jul 18, 2025



Abstract:Advancements in volume visualization (VolVis) focus on extracting insights from 3D volumetric data by generating visually compelling renderings that reveal complex internal structures. Existing VolVis approaches have explored non-photorealistic rendering techniques to enhance the clarity, expressiveness, and informativeness of visual communication. While effective, these methods often rely on complex predefined rules and are limited to transferring a single style, restricting their flexibility. To overcome these limitations, we advocate the representation of VolVis scenes using differentiable Gaussian primitives combined with pretrained large models to enable arbitrary style transfer and real-time rendering. However, conventional 3D Gaussian primitives tightly couple geometry and appearance, leading to suboptimal stylization results. To address this, we introduce TexGS-VolVis, a textured Gaussian splatting framework for VolVis. TexGS-VolVis employs 2D Gaussian primitives, extending each Gaussian with additional texture and shading attributes, resulting in higher-quality, geometry-consistent stylization and enhanced lighting control during inference. Despite these improvements, achieving flexible and controllable scene editing remains challenging. To further enhance stylization, we develop image- and text-driven non-photorealistic scene editing tailored for TexGS-VolVis and 2D-lift-3D segmentation to enable partial editing with fine-grained control. We evaluate TexGS-VolVis both qualitatively and quantitatively across various volume rendering scenes, demonstrating its superiority over existing methods in terms of efficiency, visual quality, and editing flexibility.
MC-INR: Efficient Encoding of Multivariate Scientific Simulation Data using Meta-Learning and Clustered Implicit Neural Representations
Jul 03, 2025



Abstract:Implicit Neural Representations (INRs) are widely used to encode data as continuous functions, enabling the visualization of large-scale multivariate scientific simulation data with reduced memory usage. However, existing INR-based methods face three main limitations: (1) inflexible representation of complex structures, (2) primarily focusing on single-variable data, and (3) dependence on structured grids. Thus, their performance degrades when applied to complex real-world datasets. To address these limitations, we propose a novel neural network-based framework, MC-INR, which handles multivariate data on unstructured grids. It combines meta-learning and clustering to enable flexible encoding of complex structures. To further improve performance, we introduce a residual-based dynamic re-clustering mechanism that adaptively partitions clusters based on local error. We also propose a branched layer to leverage multivariate data through independent branches simultaneously. Experimental results demonstrate that MC-INR outperforms existing methods on scientific data encoding tasks.
iVR-GS: Inverse Volume Rendering for Explorable Visualization via Editable 3D Gaussian Splatting
Apr 24, 2025



Abstract:In volume visualization, users can interactively explore the three-dimensional data by specifying color and opacity mappings in the transfer function (TF) or adjusting lighting parameters, facilitating meaningful interpretation of the underlying structure. However, rendering large-scale volumes demands powerful GPUs and high-speed memory access for real-time performance. While existing novel view synthesis (NVS) methods offer faster rendering speeds with lower hardware requirements, the visible parts of a reconstructed scene are fixed and constrained by preset TF settings, significantly limiting user exploration. This paper introduces inverse volume rendering via Gaussian splatting (iVR-GS), an innovative NVS method that reduces the rendering cost while enabling scene editing for interactive volume exploration. Specifically, we compose multiple iVR-GS models associated with basic TFs covering disjoint visible parts to make the entire volumetric scene visible. Each basic model contains a collection of 3D editable Gaussians, where each Gaussian is a 3D spatial point that supports real-time scene rendering and editing. We demonstrate the superior reconstruction quality and composability of iVR-GS against other NVS solutions (Plenoxels, CCNeRF, and base 3DGS) on various volume datasets. The code is available at https://github.com/TouKaienn/iVR-GS.
AI-Powered Automated Model Construction for Patient-Specific CFD Simulations of Aortic Flows
Mar 16, 2025



Abstract:Image-based modeling is essential for understanding cardiovascular hemodynamics and advancing the diagnosis and treatment of cardiovascular diseases. Constructing patient-specific vascular models remains labor-intensive, error-prone, and time-consuming, limiting their clinical applications. This study introduces a deep-learning framework that automates the creation of simulation-ready vascular models from medical images. The framework integrates a segmentation module for accurate voxel-based vessel delineation with a surface deformation module that performs anatomically consistent and unsupervised surface refinements guided by medical image data. By unifying voxel segmentation and surface deformation into a single cohesive pipeline, the framework addresses key limitations of existing methods, enhancing geometric accuracy and computational efficiency. Evaluated on publicly available datasets, the proposed approach demonstrates state-of-the-art performance in segmentation and mesh quality while significantly reducing manual effort and processing time. This work advances the scalability and reliability of image-based computational modeling, facilitating broader applications in clinical and research settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge