Cao Xiao
Fast Graph Attention Networks Using Effective Resistance Based Graph Sparsification
Jun 17, 2020
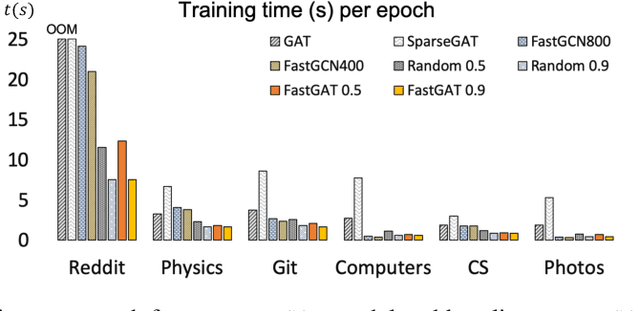
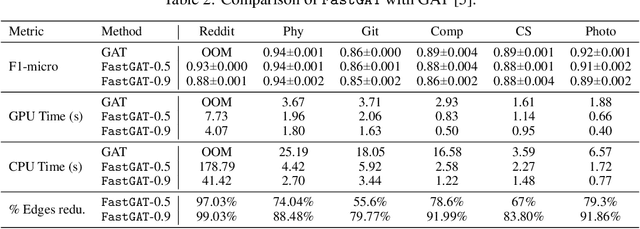
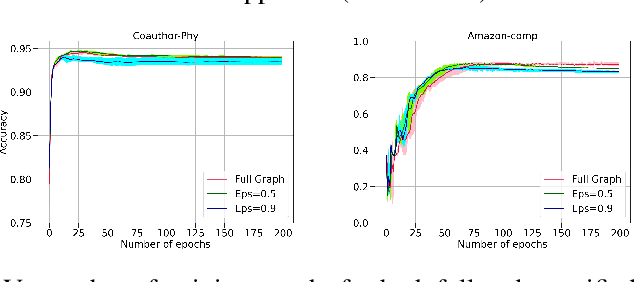
Abstract:The attention mechanism has demonstrated superior performance for inference over nodes in graph neural networks (GNNs), however, they result in a high computational burden during both training and inference. We propose FastGAT, a method to make attention based GNNs lightweight by using spectral sparsification to generate an optimal pruning of the input graph. This results in a per-epoch time that is almost linear in the number of graph nodes as opposed to quadratic. Further, we provide a re-formulation of a specific attention based GNN, Graph Attention Network (GAT) that interprets it as a graph convolution method using the random walk normalized graph Laplacian. Using this framework, we theoretically prove that spectral sparsification preserves the features computed by the GAT model, thereby justifying our FastGAT algorithm. We experimentally evaluate FastGAT on several large real world graph datasets for node classification tasks, FastGAT can dramatically reduce (up to 10x) the computational time and memory requirements, allowing the usage of attention based GNNs on large graphs.
COMPOSE: Cross-Modal Pseudo-Siamese Network for Patient Trial Matching
Jun 15, 2020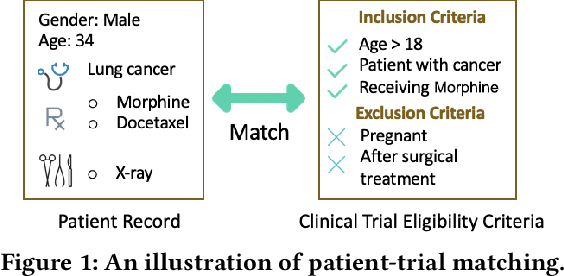
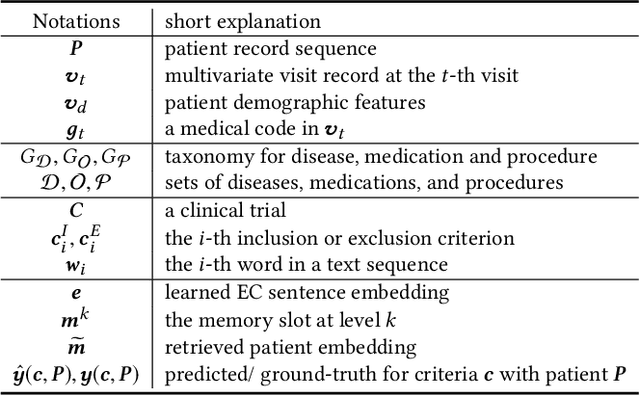
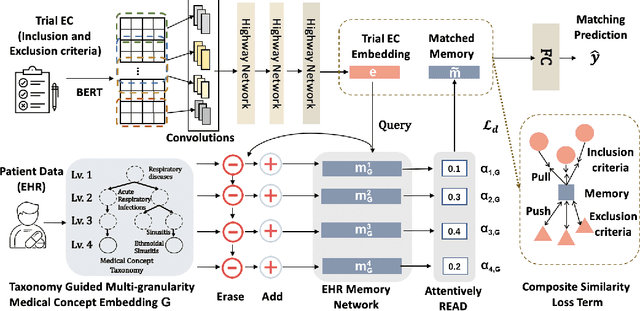
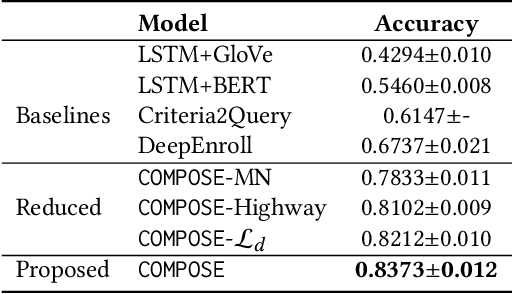
Abstract:Clinical trials play important roles in drug development but often suffer from expensive, inaccurate and insufficient patient recruitment. The availability of massive electronic health records (EHR) data and trial eligibility criteria (EC) bring a new opportunity to data driven patient recruitment. One key task named patient-trial matching is to find qualified patients for clinical trials given structured EHR and unstructured EC text (both inclusion and exclusion criteria). How to match complex EC text with longitudinal patient EHRs? How to embed many-to-many relationships between patients and trials? How to explicitly handle the difference between inclusion and exclusion criteria? In this paper, we proposed CrOss-Modal PseudO-SiamEse network (COMPOSE) to address these challenges for patient-trial matching. One path of the network encodes EC using convolutional highway network. The other path processes EHR with multi-granularity memory network that encodes structured patient records into multiple levels based on medical ontology. Using the EC embedding as query, COMPOSE performs attentional record alignment and thus enables dynamic patient-trial matching. COMPOSE also introduces a composite loss term to maximize the similarity between patient records and inclusion criteria while minimize the similarity to the exclusion criteria. Experiment results show COMPOSE can reach 98.0% AUC on patient-criteria matching and 83.7% accuracy on patient-trial matching, which leads 24.3% improvement over the best baseline on real-world patient-trial matching tasks.
CHEER: Rich Model Helps Poor Model via Knowledge Infusion
May 21, 2020
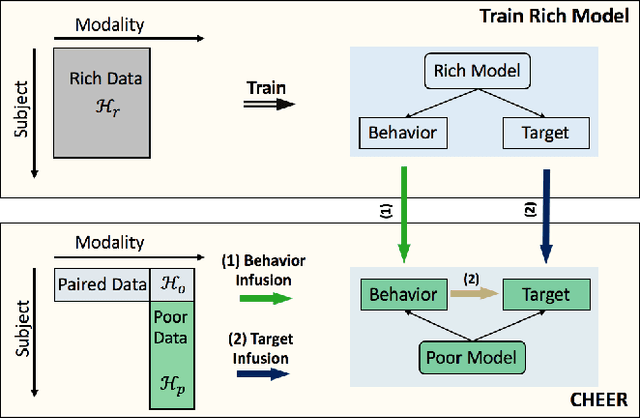
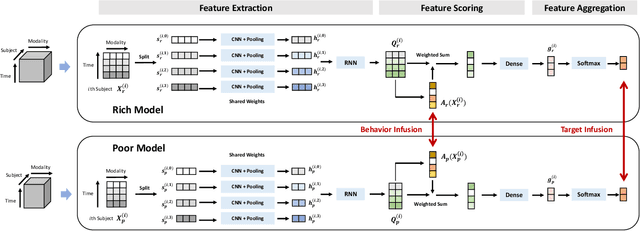
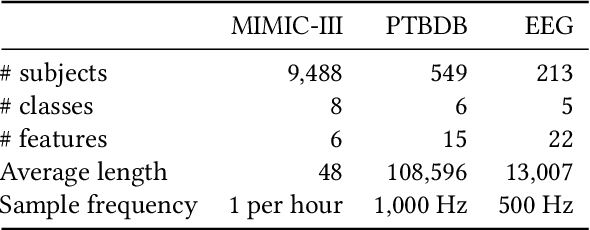
Abstract:There is a growing interest in applying deep learning (DL) to healthcare, driven by the availability of data with multiple feature channels in rich-data environments (e.g., intensive care units). However, in many other practical situations, we can only access data with much fewer feature channels in a poor-data environments (e.g., at home), which often results in predictive models with poor performance. How can we boost the performance of models learned from such poor-data environment by leveraging knowledge extracted from existing models trained using rich data in a related environment? To address this question, we develop a knowledge infusion framework named CHEER that can succinctly summarize such rich model into transferable representations, which can be incorporated into the poor model to improve its performance. The infused model is analyzed theoretically and evaluated empirically on several datasets. Our empirical results showed that CHEER outperformed baselines by 5.60% to 46.80% in terms of the macro-F1 score on multiple physiological datasets.
SkipGNN: Predicting Molecular Interactions with Skip-Graph Networks
Apr 30, 2020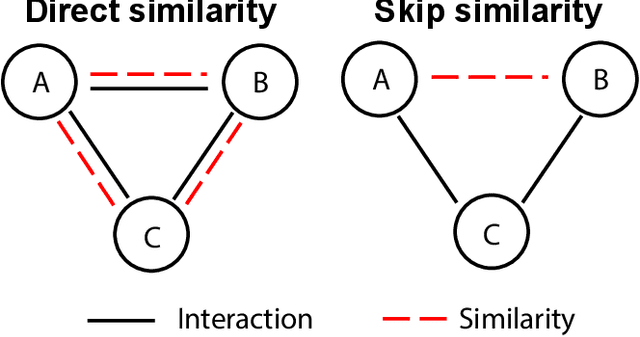
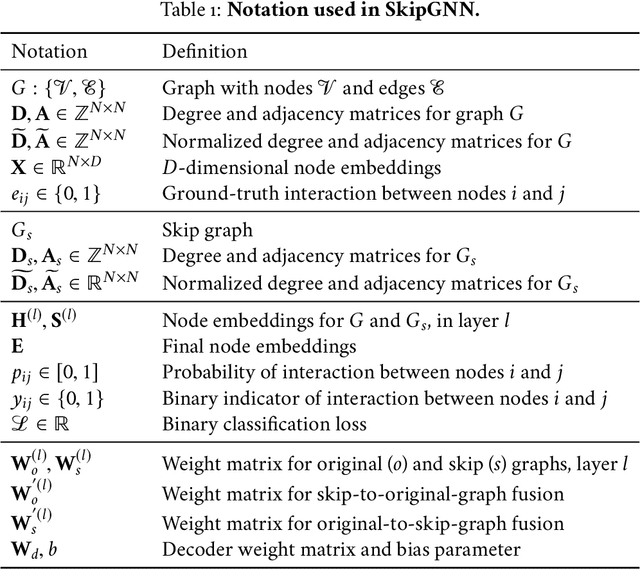
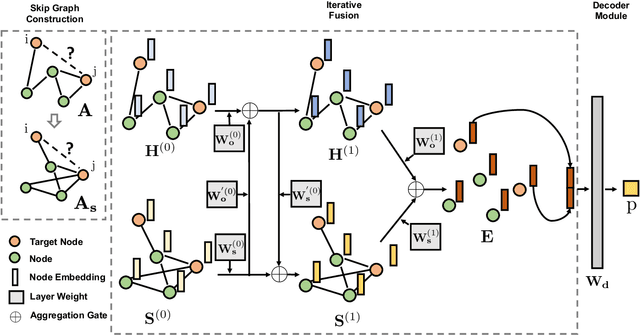
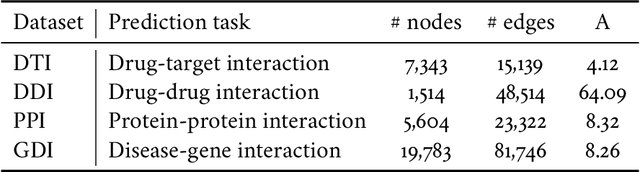
Abstract:Molecular interaction networks are powerful resources for the discovery. They are increasingly used with machine learning methods to predict biologically meaningful interactions. While deep learning on graphs has dramatically advanced the prediction prowess, current graph neural network (GNN) methods are optimized for prediction on the basis of direct similarity between interacting nodes. In biological networks, however, similarity between nodes that do not directly interact has proved incredibly useful in the last decade across a variety of interaction networks. Here, we present SkipGNN, a graph neural network approach for the prediction of molecular interactions. SkipGNN predicts molecular interactions by not only aggregating information from direct interactions but also from second-order interactions, which we call skip similarity. In contrast to existing GNNs, SkipGNN receives neural messages from two-hop neighbors as well as immediate neighbors in the interaction network and non-linearly transforms the messages to obtain useful information for prediction. To inject skip similarity into a GNN, we construct a modified version of the original network, called the skip graph. We then develop an iterative fusion scheme that optimizes a GNN using both the skip graph and the original graph. Experiments on four interaction networks, including drug-drug, drug-target, protein-protein, and gene-disease interactions, show that SkipGNN achieves superior and robust performance, outperforming existing methods by up to 28.8\% of area under the precision recall curve (PR-AUC). Furthermore, we show that unlike popular GNNs, SkipGNN learns biologically meaningful embeddings and performs especially well on noisy, incomplete interaction networks.
MolTrans: Molecular Interaction Transformer for Drug Target Interaction Prediction
Apr 23, 2020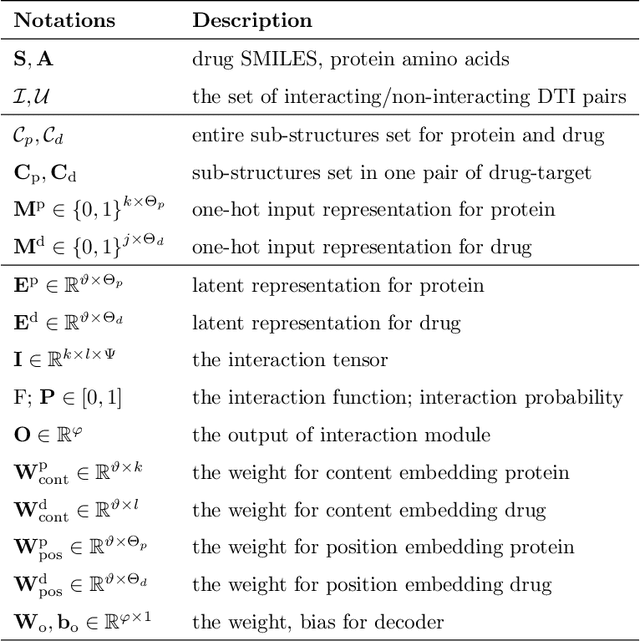
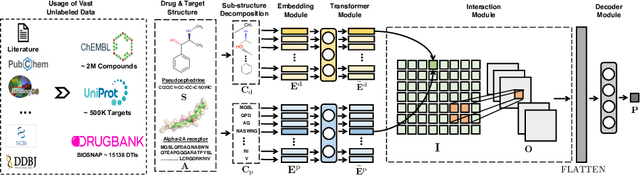
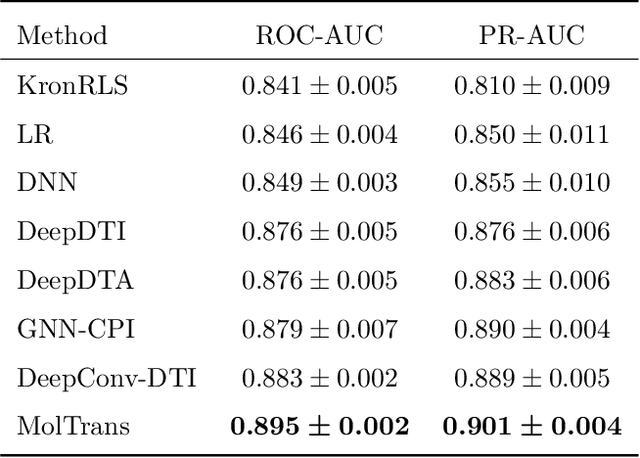
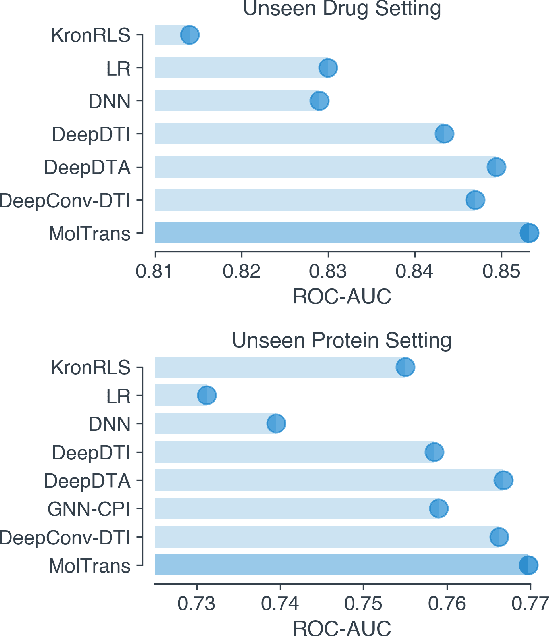
Abstract:Drug target interaction (DTI) prediction is a foundational task for in silico drug discovery, which is costly and time-consuming due to the need of experimental search over large drug compound space. Recent years have witnessed promising progress for deep learning in DTI predictions. However, the following challenges are still open: (1) the sole data-driven molecular representation learning approaches ignore the sub-structural nature of DTI, thus produce results that are less accurate and difficult to explain; (2) existing methods focus on limited labeled data while ignoring the value of massive unlabelled molecular data. We propose a Molecular Interaction Transformer (MolTrans) to address these limitations via: (1) knowledge inspired sub-structural pattern mining algorithm and interaction modeling module for more accurate and interpretable DTI prediction; (2) an augmented transformer encoder to better extract and capture the semantic relations among substructures extracted from massive unlabeled biomedical data. We evaluate MolTrans on real world data and show it improved DTI prediction performance compared to state-of-the-art baselines.
DeepPurpose: a Deep Learning Based Drug Repurposing Toolkit
Apr 19, 2020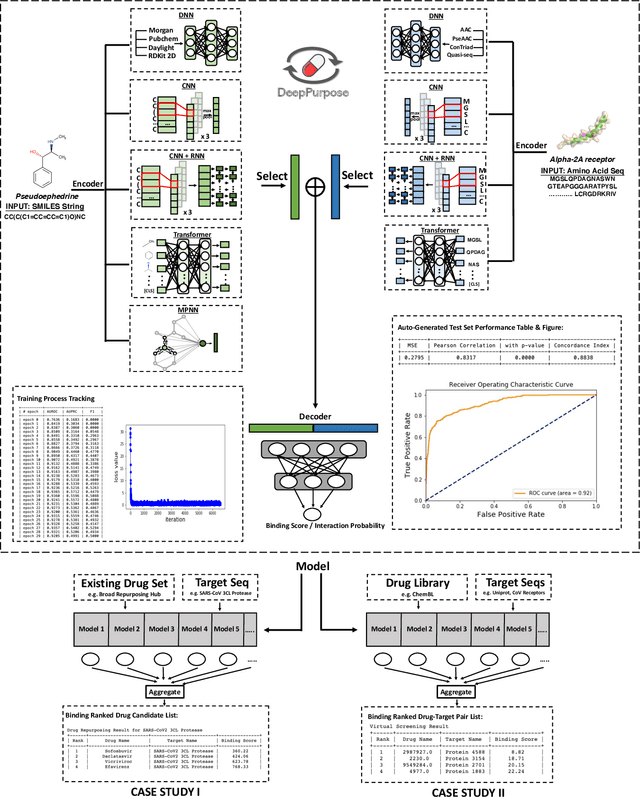
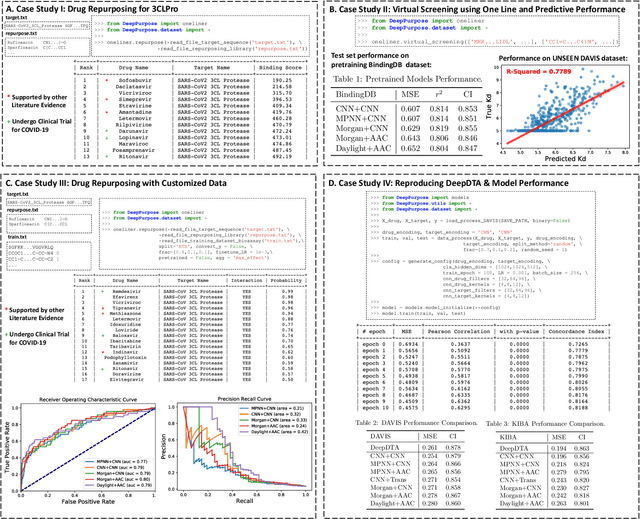
Abstract:We present DeepPurpose, a deep learning toolkit for simple and efficient drug repurposing. With a few lines of code, DeepPurpose generates drug candidates based on aggregating five pretrained state-of-the-art models while offering flexibility for users to train their own models with 15 drug/target encodings and $50+$ novel architectures. We demonstrated DeepPurpose using case studies, including repurposing for COVID-19 where promising candidates under trials are ranked high in our results.
SUOD: A Scalable Unsupervised Outlier Detection Framework
Mar 11, 2020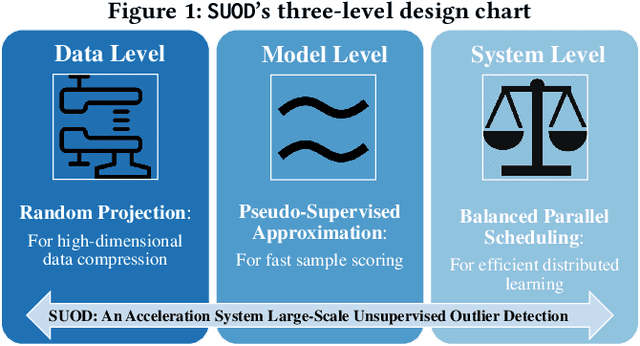
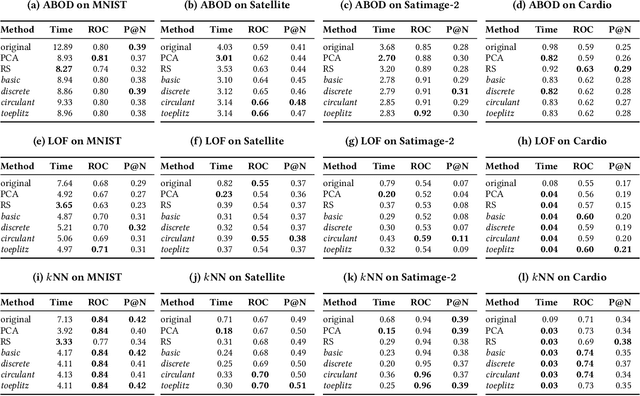
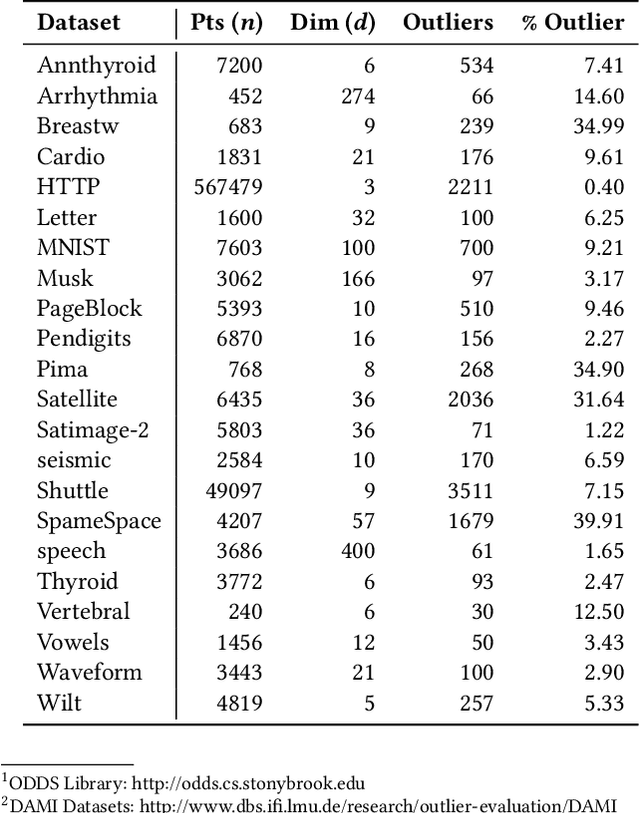
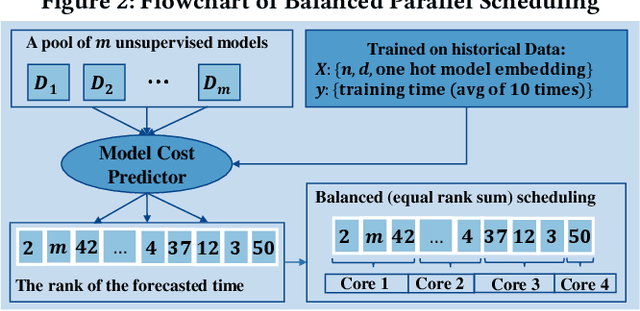
Abstract:Outlier detection is a key data mining task for identifying abnormal objects from massive data. Due to the high expense of acquiring ground truth, practitioners lean towards building a large number of unsupervised models for further combination and analysis, rather than relying on a single model out of reliability consideration. However, this poses scalability challenge to high-dimensional, large datasets. In this study, we propose a three-module framework called SUOD to address the challenge. It can accelerate outlier model building and scoring when a large number of base models are used. It focuses on three complementary levels to speed up the process while controlling prediction performance degradation at the same time. At the data level, its Random Projection module projects high-dimensional data onto diversified low-dimensional subspaces while preserving the pairwise distance relationship. At the model level, SUOD's Pseudo-supervised Approximation module can approximate and replace fitted unsupervised models by low-cost supervised regressors, leading to fast offline scoring on new-coming samples with better interpretability. At the system level, Balanced Parallel Scheduling module mitigates the workload imbalance within distributed systems, which is helpful for heterogeneous outlier ensembles. As the three modules are independent with different focuses, they have great flexibility to "mix and match". The framework is also designed with extensibility in mind. One may customize each module based on specific use cases, and the framework may be generalized to other learning tasks as well. Extensive experiments on more than 20 benchmark datasets demonstrate SUOD's effectiveness. In addition, a real-world deployment system on fraudulent claim analysis by IQVIA is also discussed. The full framework, documentation, and examples are openly shared at https://github.com/yzhao062/SUOD.
CLARA: Clinical Report Auto-completion
Mar 04, 2020
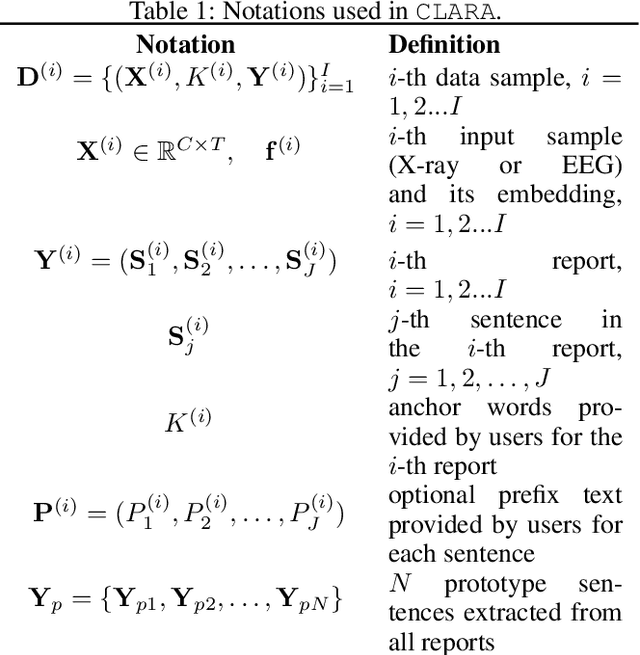

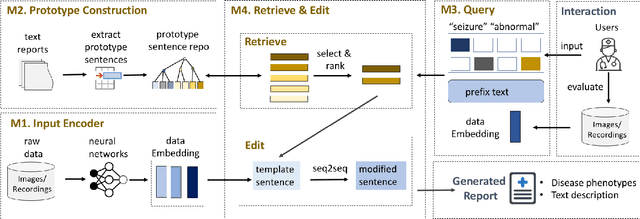
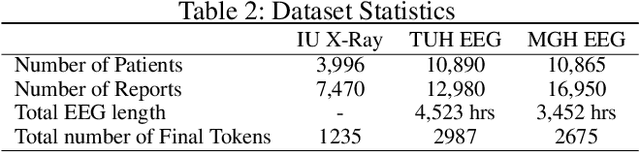
Abstract:Generating clinical reports from raw recordings such as X-rays and electroencephalogram (EEG) is an essential and routine task for doctors. However, it is often time-consuming to write accurate and detailed reports. Most existing methods try to generate the whole reports from the raw input with limited success because 1) generated reports often contain errors that need manual review and correction, 2) it does not save time when doctors want to write additional information into the report, and 3) the generated reports are not customized based on individual doctors' preference. We propose {\it CL}inic{\it A}l {\it R}eport {\it A}uto-completion (CLARA), an interactive method that generates reports in a sentence by sentence fashion based on doctors' anchor words and partially completed sentences. CLARA searches for most relevant sentences from existing reports as the template for the current report. The retrieved sentences are sequentially modified by combining with the input feature representations to create the final report. In our experimental evaluation, CLARA achieved 0.393 CIDEr and 0.248 BLEU-4 on X-ray reports and 0.482 CIDEr and 0.491 BLEU-4 for EEG reports for sentence-level generation, which is up to 35% improvement over the best baseline. Also via our qualitative evaluation, CLARA is shown to produce reports which have a significantly higher level of approval by doctors in a user study (3.74 out of 5 for CLARA vs 2.52 out of 5 for the baseline).
REST: Robust and Efficient Neural Networks for Sleep Monitoring in the Wild
Jan 29, 2020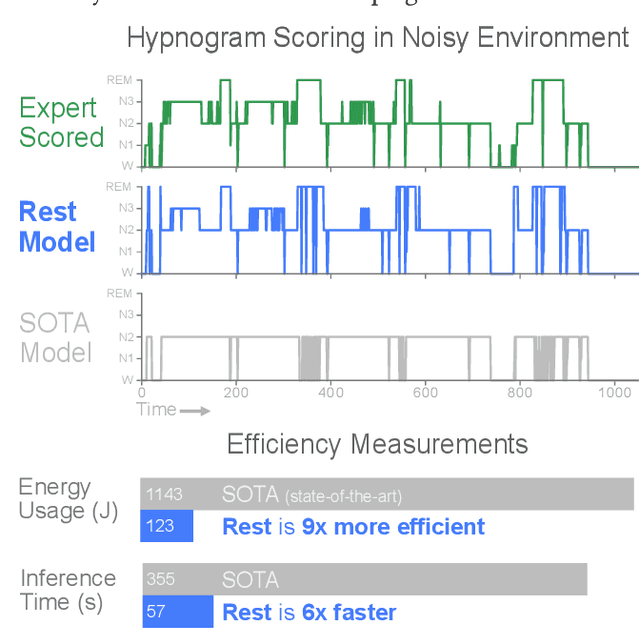
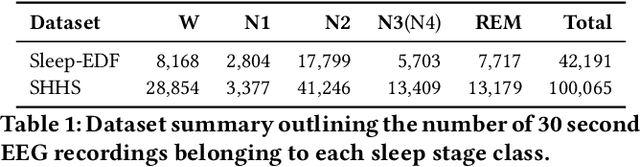
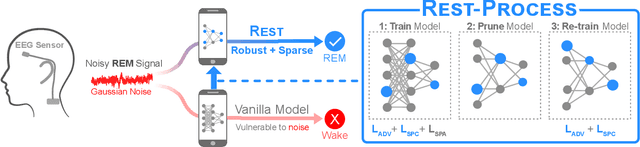
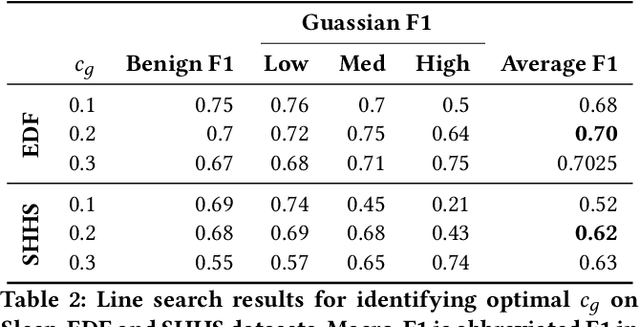
Abstract:In recent years, significant attention has been devoted towards integrating deep learning technologies in the healthcare domain. However, to safely and practically deploy deep learning models for home health monitoring, two significant challenges must be addressed: the models should be (1) robust against noise; and (2) compact and energy-efficient. We propose REST, a new method that simultaneously tackles both issues via 1) adversarial training and controlling the Lipschitz constant of the neural network through spectral regularization while 2) enabling neural network compression through sparsity regularization. We demonstrate that REST produces highly-robust and efficient models that substantially outperform the original full-sized models in the presence of noise. For the sleep staging task over single-channel electroencephalogram (EEG), the REST model achieves a macro-F1 score of 0.67 vs. 0.39 achieved by a state-of-the-art model in the presence of Gaussian noise while obtaining 19x parameter reduction and 15x MFLOPS reduction on two large, real-world EEG datasets. By deploying these models to an Android application on a smartphone, we quantitatively observe that REST allows models to achieve up to 17x energy reduction and 9x faster inference. We open-source the code repository with this paper: https://github.com/duggalrahul/REST.
StageNet: Stage-Aware Neural Networks for Health Risk Prediction
Jan 24, 2020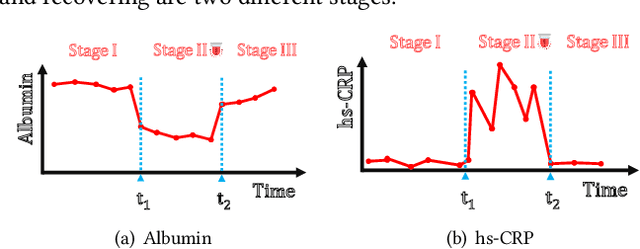
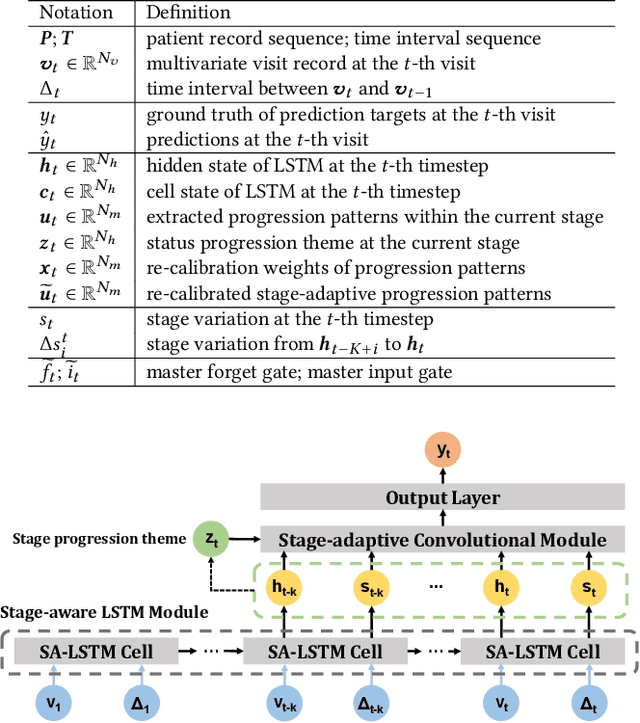
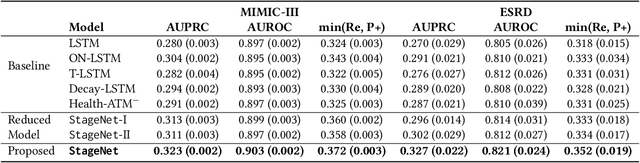
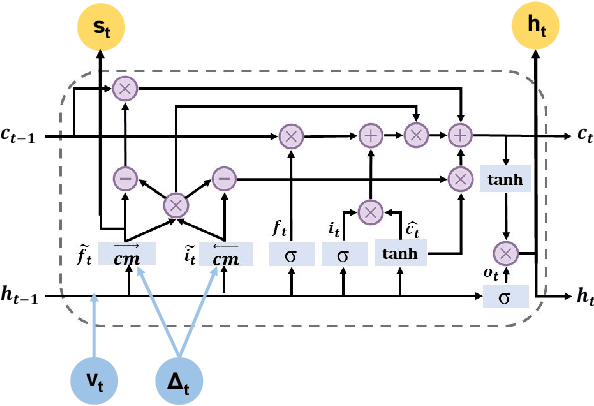
Abstract:Deep learning has demonstrated success in health risk prediction especially for patients with chronic and progressing conditions. Most existing works focus on learning disease Network (StageNet) model to extract disease stage information from patient data and integrate it into risk prediction. StageNet is enabled by (1) a stage-aware long short-term memory (LSTM) module that extracts health stage variations unsupervisedly; (2) a stage-adaptive convolutional module that incorporates stage-related progression patterns into risk prediction. We evaluate StageNet on two real-world datasets and show that StageNet outperforms state-of-the-art models in risk prediction task and patient subtyping task. Compared to the best baseline model, StageNet achieves up to 12% higher AUPRC for risk prediction task on two real-world patient datasets. StageNet also achieves over 58% higher Calinski-Harabasz score (a cluster quality metric) for a patient subtyping task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge