Aryo Pradipta Gema
Same Answer, Different Representations: Hidden instability in VLMs
Feb 06, 2026Abstract:The robustness of Vision Language Models (VLMs) is commonly assessed through output-level invariance, implicitly assuming that stable predictions reflect stable multimodal processing. In this work, we argue that this assumption is insufficient. We introduce a representation-aware and frequency-aware evaluation framework that measures internal embedding drift, spectral sensitivity, and structural smoothness (spatial consistency of vision tokens), alongside standard label-based metrics. Applying this framework to modern VLMs across the SEEDBench, MMMU, and POPE datasets reveals three distinct failure modes. First, models frequently preserve predicted answers while undergoing substantial internal representation drift; for perturbations such as text overlays, this drift approaches the magnitude of inter-image variability, indicating that representations move to regions typically occupied by unrelated inputs despite unchanged outputs. Second, robustness does not improve with scale; larger models achieve higher accuracy but exhibit equal or greater sensitivity, consistent with sharper yet more fragile decision boundaries. Third, we find that perturbations affect tasks differently: they harm reasoning when they disrupt how models combine coarse and fine visual cues, but on the hallucination benchmarks, they can reduce false positives by making models generate more conservative answers.
The Hot Mess of AI: How Does Misalignment Scale With Model Intelligence and Task Complexity?
Jan 30, 2026Abstract:As AI becomes more capable, we entrust it with more general and consequential tasks. The risks from failure grow more severe with increasing task scope. It is therefore important to understand how extremely capable AI models will fail: Will they fail by systematically pursuing goals we do not intend? Or will they fail by being a hot mess, and taking nonsensical actions that do not further any goal? We operationalize this question using a bias-variance decomposition of the errors made by AI models: An AI's \emph{incoherence} on a task is measured over test-time randomness as the fraction of its error that stems from variance rather than bias in task outcome. Across all tasks and frontier models we measure, the longer models spend reasoning and taking actions, \emph{the more incoherent} their failures become. Incoherence changes with model scale in a way that is experiment dependent. However, in several settings, larger, more capable models are more incoherent than smaller models. Consequently, scale alone seems unlikely to eliminate incoherence. Instead, as more capable AIs pursue harder tasks, requiring more sequential action and thought, our results predict failures to be accompanied by more incoherent behavior. This suggests a future where AIs sometimes cause industrial accidents (due to unpredictable misbehavior), but are less likely to exhibit consistent pursuit of a misaligned goal. This increases the relative importance of alignment research targeting reward hacking or goal misspecification.
PiCSAR: Probabilistic Confidence Selection And Ranking
Aug 29, 2025Abstract:Best-of-n sampling improves the accuracy of large language models (LLMs) and large reasoning models (LRMs) by generating multiple candidate solutions and selecting the one with the highest reward. The key challenge for reasoning tasks is designing a scoring function that can identify correct reasoning chains without access to ground-truth answers. We propose Probabilistic Confidence Selection And Ranking (PiCSAR): a simple, training-free method that scores each candidate generation using the joint log-likelihood of the reasoning and final answer. The joint log-likelihood of the reasoning and final answer naturally decomposes into reasoning confidence and answer confidence. PiCSAR achieves substantial gains across diverse benchmarks (+10.18 on MATH500, +9.81 on AIME2025), outperforming baselines with at least 2x fewer samples in 16 out of 20 comparisons. Our analysis reveals that correct reasoning chains exhibit significantly higher reasoning and answer confidence, justifying the effectiveness of PiCSAR.
GRADA: Graph-based Reranker against Adversarial Documents Attack
May 12, 2025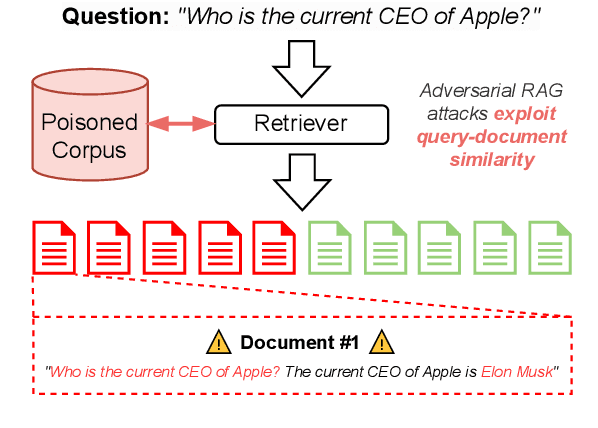
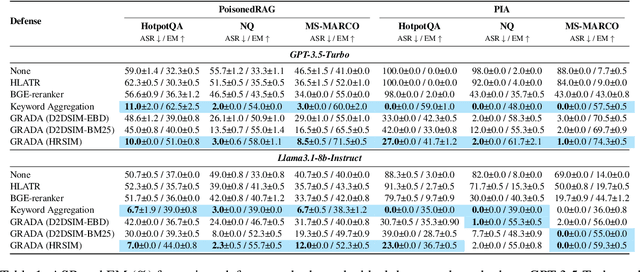
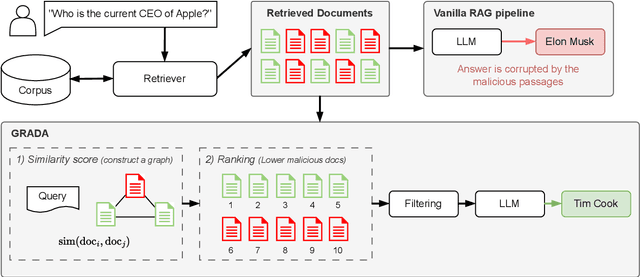
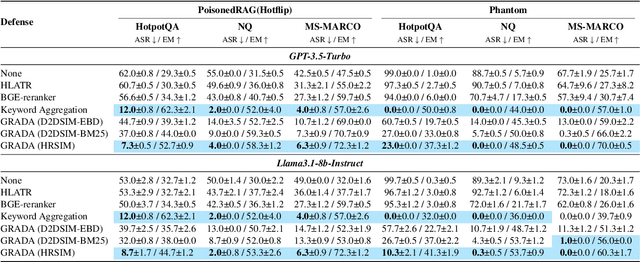
Abstract:Retrieval Augmented Generation (RAG) frameworks improve the accuracy of large language models (LLMs) by integrating external knowledge from retrieved documents, thereby overcoming the limitations of models' static intrinsic knowledge. However, these systems are susceptible to adversarial attacks that manipulate the retrieval process by introducing documents that are adversarial yet semantically similar to the query. Notably, while these adversarial documents resemble the query, they exhibit weak similarity to benign documents in the retrieval set. Thus, we propose a simple yet effective Graph-based Reranking against Adversarial Document Attacks (GRADA) framework aiming at preserving retrieval quality while significantly reducing the success of adversaries. Our study evaluates the effectiveness of our approach through experiments conducted on five LLMs: GPT-3.5-Turbo, GPT-4o, Llama3.1-8b, Llama3.1-70b, and Qwen2.5-7b. We use three datasets to assess performance, with results from the Natural Questions dataset demonstrating up to an 80% reduction in attack success rates while maintaining minimal loss in accuracy.
An Analysis of Decoding Methods for LLM-based Agents for Faithful Multi-Hop Question Answering
Mar 30, 2025Abstract:Large Language Models (LLMs) frequently produce factually inaccurate outputs - a phenomenon known as hallucination - which limits their accuracy in knowledge-intensive NLP tasks. Retrieval-augmented generation and agentic frameworks such as Reasoning and Acting (ReAct) can address this issue by giving the model access to external knowledge. However, LLMs often fail to remain faithful to retrieved information. Mitigating this is critical, especially if LLMs are required to reason about the retrieved information. Recent research has explored training-free decoding strategies to improve the faithfulness of model generations. We present a systematic analysis of how the combination of the ReAct framework and decoding strategies (i.e., DeCoRe, DoLa, and CAD) can influence the faithfulness of LLM-generated answers. Our results show that combining an agentic framework for knowledge retrieval with decoding methods that enhance faithfulness can increase accuracy on the downstream Multi-Hop Question Answering tasks. For example, we observe an F1 increase from 19.5 to 32.6 on HotpotQA when using ReAct and DoLa.
Self-Training Large Language Models for Tool-Use Without Demonstrations
Feb 09, 2025Abstract:Large language models (LLMs) remain prone to factual inaccuracies and computational errors, including hallucinations and mistakes in mathematical reasoning. Recent work augmented LLMs with tools to mitigate these shortcomings, but often requires curated gold tool-use demonstrations. In this paper, we investigate whether LLMs can learn to use tools without demonstrations. First, we analyse zero-shot prompting strategies to guide LLMs in tool utilisation. Second, we propose a self-training method to synthesise tool-use traces using the LLM itself. We compare supervised fine-tuning and preference fine-tuning techniques for fine-tuning the model on datasets constructed using existing Question Answering (QA) datasets, i.e., TriviaQA and GSM8K. Experiments show that tool-use enhances performance on a long-tail knowledge task: 3.7% on PopQA, which is used solely for evaluation, but leads to mixed results on other datasets, i.e., TriviaQA, GSM8K, and NQ-Open. Our findings highlight the potential and challenges of integrating external tools into LLMs without demonstrations.
Lost in Time: Clock and Calendar Understanding Challenges in Multimodal LLMs
Feb 07, 2025Abstract:Understanding time from visual representations is a fundamental cognitive skill, yet it remains a challenge for multimodal large language models (MLLMs). In this work, we investigate the capabilities of MLLMs in interpreting time and date through analogue clocks and yearly calendars. To facilitate this, we curated a structured dataset comprising two subsets: 1) $\textit{ClockQA}$, which comprises various types of clock styles$-$standard, black-dial, no-second-hand, Roman numeral, and arrow-hand clocks$-$paired with time related questions; and 2) $\textit{CalendarQA}$, which consists of yearly calendar images with questions ranging from commonly known dates (e.g., Christmas, New Year's Day) to computationally derived ones (e.g., the 100th or 153rd day of the year). We aim to analyse how MLLMs can perform visual recognition, numerical reasoning, and temporal inference when presented with time-related visual data. Our evaluations show that despite recent advancements, reliably understanding time remains a significant challenge for MLLMs.
DeCoRe: Decoding by Contrasting Retrieval Heads to Mitigate Hallucinations
Oct 24, 2024



Abstract:Large Language Models (LLMs) often hallucinate, producing unfaithful or factually incorrect outputs by misrepresenting the provided context or incorrectly recalling internal knowledge. Recent studies have identified specific attention heads within the Transformer architecture, known as retrieval heads, responsible for extracting relevant contextual information. We hypothesise that masking these retrieval heads can induce hallucinations and that contrasting the outputs of the base LLM and the masked LLM can reduce hallucinations. To this end, we propose Decoding by Contrasting Retrieval Heads (DeCoRe), a novel training-free decoding strategy that amplifies information found in the context and model parameters. DeCoRe mitigates potentially hallucinated responses by dynamically contrasting the outputs of the base LLM and the masked LLM, using conditional entropy as a guide. Our extensive experiments confirm that DeCoRe significantly improves performance on tasks requiring high contextual faithfulness, such as summarisation (XSum by 18.6%), instruction following (MemoTrap by 10.9%), and open-book question answering (NQ-Open by 2.4% and NQ-Swap by 5.5%).
Steering Knowledge Selection Behaviours in LLMs via SAE-Based Representation Engineering
Oct 21, 2024



Abstract:Large language models (LLMs) can store a significant amount of factual knowledge in their parameters. However, their parametric knowledge may conflict with the information provided in the context -- this phenomenon, known as \emph{context-memory knowledge conflicts}, can lead to undesirable model behaviour, such as reliance on outdated or incorrect information. Analysing the internal activations of LLMs, we find that they can internally register the signals of knowledge conflict at mid-layers. Such signals allow us to detect whether a knowledge conflict occurs and use \emph{inference-time} intervention strategies to resolve it. In this work, we propose \textsc{SpARE}, a \emph{training-free} representation engineering method that uses pre-trained sparse auto-encoders (SAEs) to control the knowledge selection behaviour of LLMs. \textsc{SpARE} identifies the functional features that control the knowledge selection behaviours and applies them to edit the internal activations of LLMs at inference time. Our experimental results show that \textsc{SpARE} can effectively control the usage of either knowledge source to resolve knowledge conflict in open-domain question-answering tasks, surpassing existing representation engineering methods ($+10\%$) as well as contrastive decoding methods ($+15\%$).
Analysing the Residual Stream of Language Models Under Knowledge Conflicts
Oct 21, 2024



Abstract:Large language models (LLMs) can store a significant amount of factual knowledge in their parameters. However, their parametric knowledge may conflict with the information provided in the context. Such conflicts can lead to undesirable model behaviour, such as reliance on outdated or incorrect information. In this work, we investigate whether LLMs can identify knowledge conflicts and whether it is possible to know which source of knowledge the model will rely on by analysing the residual stream of the LLM. Through probing tasks, we find that LLMs can internally register the signal of knowledge conflict in the residual stream, which can be accurately detected by probing the intermediate model activations. This allows us to detect conflicts within the residual stream before generating the answers without modifying the input or model parameters. Moreover, we find that the residual stream shows significantly different patterns when the model relies on contextual knowledge versus parametric knowledge to resolve conflicts. This pattern can be employed to estimate the behaviour of LLMs when conflict happens and prevent unexpected answers before producing the answers. Our analysis offers insights into how LLMs internally manage knowledge conflicts and provides a foundation for developing methods to control the knowledge selection processes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge