Andreas Maier
Pattern Recognition Lab, FAU Erlangen-Nürnberg, Germany
Inter-Species Cell Detection: Datasets on pulmonary hemosiderophages in equine, human and feline specimens
Aug 19, 2021
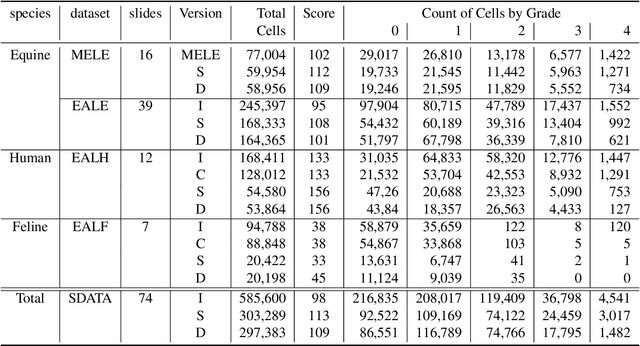
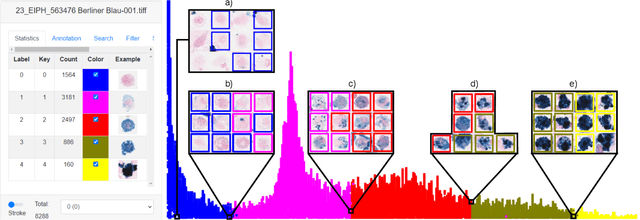
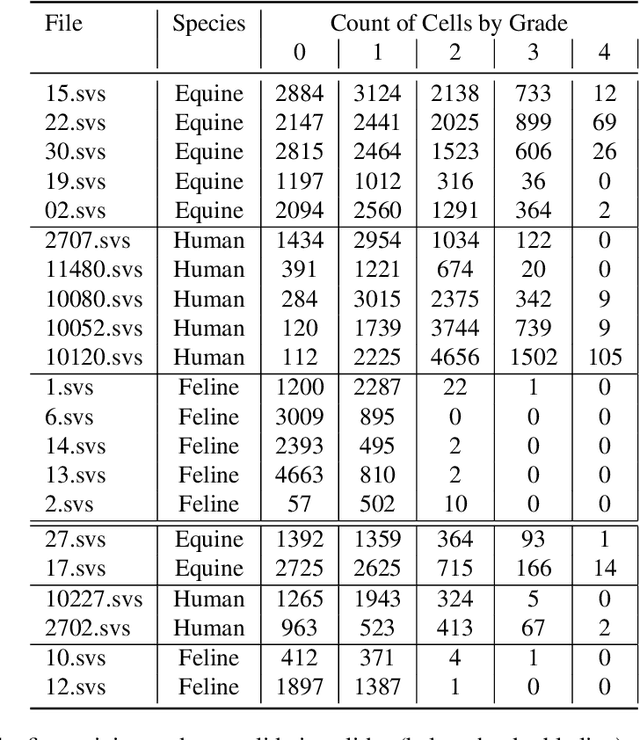
Abstract:Pulmonary hemorrhage (P-Hem) occurs among multiple species and can have various causes. Cytology of bronchoalveolarlavage fluid (BALF) using a 5-tier scoring system of alveolar macrophages based on their hemosiderin content is considered the most sensitive diagnostic method. We introduce a novel, fully annotated multi-species P-Hem dataset which consists of 74 cytology whole slide images (WSIs) with equine, feline and human samples. To create this high-quality and high-quantity dataset, we developed an annotation pipeline combining human expertise with deep learning and data visualisation techniques. We applied a deep learning-based object detection approach trained on 17 expertly annotated equine WSIs, to the remaining 39 equine, 12 human and 7 feline WSIs. The resulting annotations were semi-automatically screened for errors on multiple types of specialised annotation maps and finally reviewed by a trained pathologists. Our dataset contains a total of 297,383 hemosiderophages classified into five grades. It is one of the largest publicly availableWSIs datasets with respect to the number of annotations, the scanned area and the number of species covered.
Known Operator Learning and Hybrid Machine Learning in Medical Imaging --- A Review of the Past, the Present, and the Future
Aug 10, 2021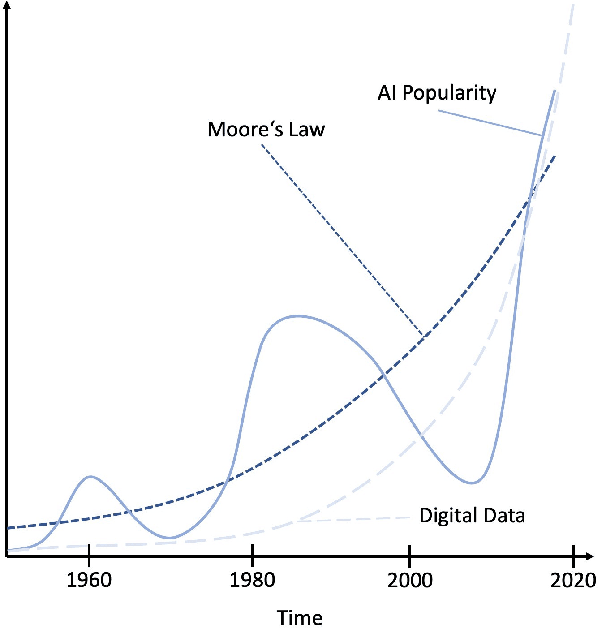
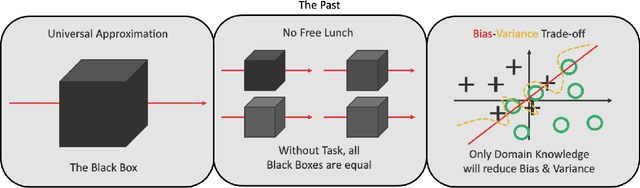
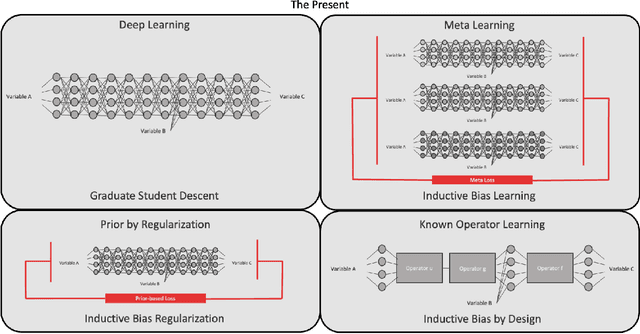
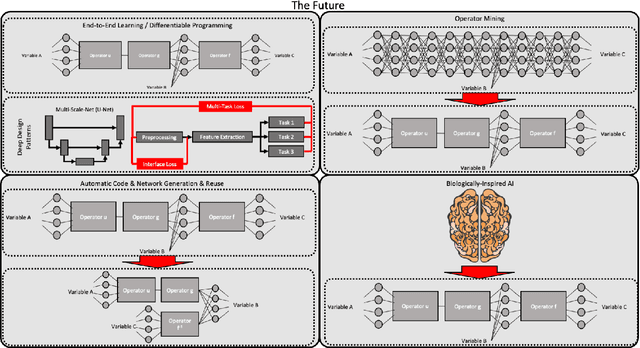
Abstract:In this article, we perform a review of the state-of-the-art of hybrid machine learning in medical imaging. We start with a short summary of the general developments of the past in machine learning and how general and specialized approaches have been in competition in the past decades. A particular focus will be the theoretical and experimental evidence pro and contra hybrid modelling. Next, we inspect several new developments regarding hybrid machine learning with a particular focus on so-called known operator learning and how hybrid approaches gain more and more momentum across essentially all applications in medical imaging and medical image analysis. As we will point out by numerous examples, hybrid models are taking over in image reconstruction and analysis. Even domains such as physical simulation and scanner and acquisition design are being addressed using machine learning grey box modelling approaches. Towards the end of the article, we will investigate a few future directions and point out relevant areas in which hybrid modelling, meta learning, and other domains will likely be able to drive the state-of-the-art ahead.
Deep Iterative 2D/3D Registration
Jul 21, 2021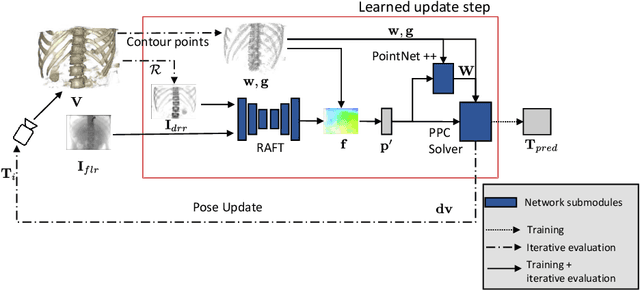
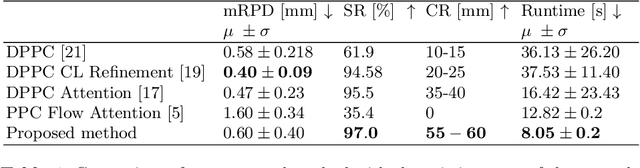
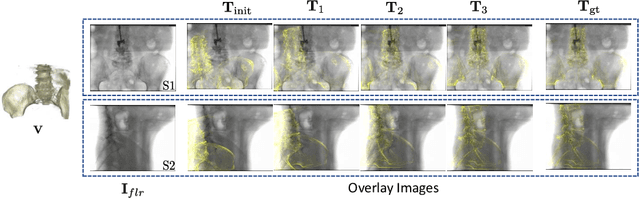
Abstract:Deep Learning-based 2D/3D registration methods are highly robust but often lack the necessary registration accuracy for clinical application. A refinement step using the classical optimization-based 2D/3D registration method applied in combination with Deep Learning-based techniques can provide the required accuracy. However, it also increases the runtime. In this work, we propose a novel Deep Learning driven 2D/3D registration framework that can be used end-to-end for iterative registration tasks without relying on any further refinement step. We accomplish this by learning the update step of the 2D/3D registration framework using Point-to-Plane Correspondences. The update step is learned using iterative residual refinement-based optical flow estimation, in combination with the Point-to-Plane correspondence solver embedded as a known operator. Our proposed method achieves an average runtime of around 8s, a mean re-projection distance error of 0.60 $\pm$ 0.40 mm with a success ratio of 97 percent and a capture range of 60 mm. The combination of high registration accuracy, high robustness, and fast runtime makes our solution ideal for clinical applications.
Automatic and explainable grading of meningiomas from histopathology images
Jul 19, 2021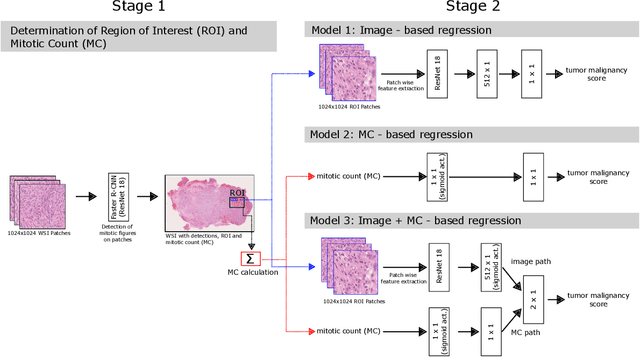
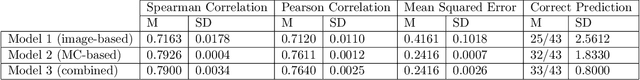
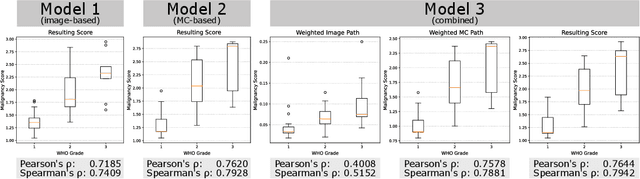
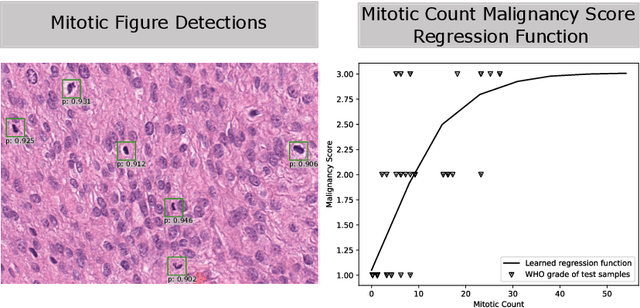
Abstract:Meningioma is one of the most prevalent brain tumors in adults. To determine its malignancy, it is graded by a pathologist into three grades according to WHO standards. This grade plays a decisive role in treatment, and yet may be subject to inter-rater discordance. In this work, we present and compare three approaches towards fully automatic meningioma grading from histology whole slide images. All approaches are following a two-stage paradigm, where we first identify a region of interest based on the detection of mitotic figures in the slide using a state-of-the-art object detection deep learning network. This region of highest mitotic rate is considered characteristic for biological tumor behavior. In the second stage, we calculate a score corresponding to tumor malignancy based on information contained in this region using three different settings. In a first approach, image patches are sampled from this region and regression is based on morphological features encoded by a ResNet-based network. We compare this to learning a logistic regression from the determined mitotic count, an approach which is easily traceable and explainable. Lastly, we combine both approaches in a single network. We trained the pipeline on 951 slides from 341 patients and evaluated them on a separate set of 141 slides from 43 patients. All approaches yield a high correlation to the WHO grade. The logistic regression and the combined approach had the best results in our experiments, yielding correct predictions in 32 and 33 of all cases, respectively, with the image-based approach only predicting 25 cases correctly. Spearman's correlation was 0.716, 0.792 and 0.790 respectively. It may seem counterintuitive at first that morphological features provided by image patches do not improve model performance. Yet, this mirrors the criteria of the grading scheme, where mitotic count is the only unequivocal parameter.
FlexParser -- the adaptive log file parser for continuous results in a changing world
Jun 06, 2021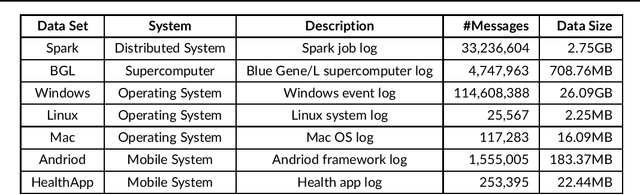
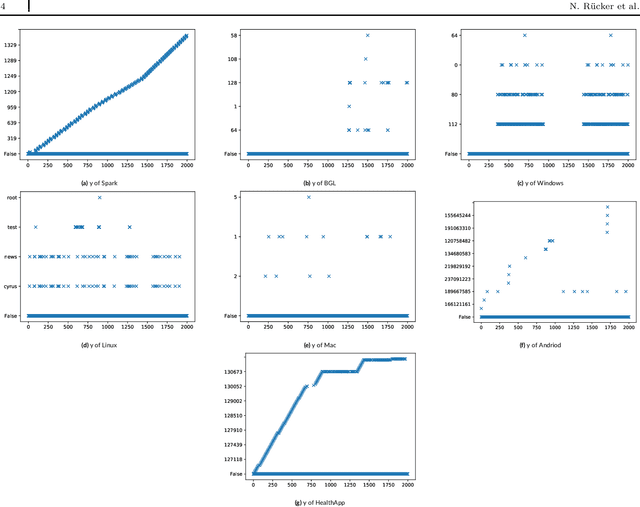
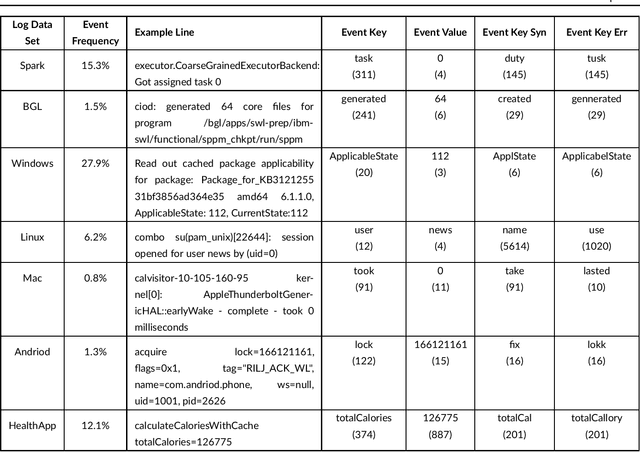
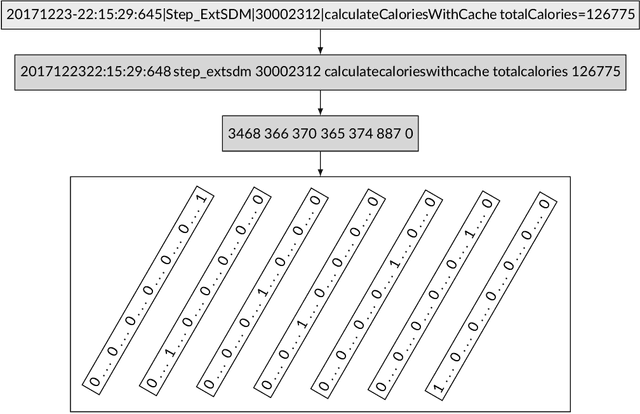
Abstract:Any modern system writes events into files, called log files. Those contain crucial information which are subject to various analyses. Examples range from cybersecurity, intrusion detection over usage analyses to trouble shooting. Before data analysis is possible, desired information needs to be extracted first out of the semi-structured log messages. State of the art event parsing often assumes static log events. However, any modern system is updated consistently and with updates also log file structures can change. We call those changes 'mutations' and study parsing performance for different mutation cases. Latest research discovers mutations using anomaly detection post mortem, however, does not cover actual continuous parsing. Thus, we propose a novel, flexible parser, called FlexParser which can extract desired values despite gradual changes in the log messages. It implies basic text preprocessing followed by a supervised Deep Learning method. We train a stateful LSTM on parsing one event per data set. Statefulness enforces the model to learn log message structures across several messages. Our model was tested on seven different, publicly available log file data sets and various kinds of mutations. Exhibiting an average F1-Score of 0.98, it outperforms other Deep Learning methods as well as state-of-the-art unsupervised parsers.
SmartPatch: Improving Handwritten Word Imitation with Patch Discriminators
May 21, 2021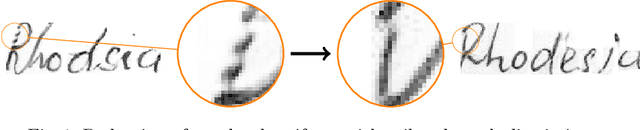
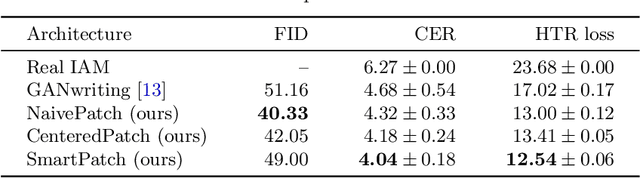
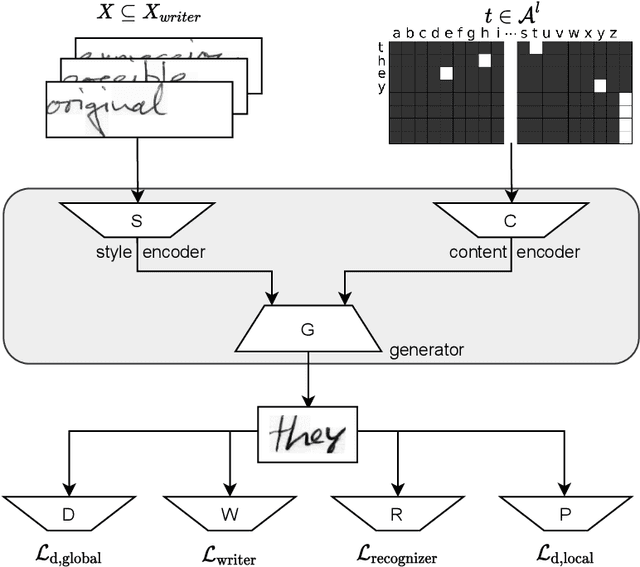

Abstract:As of recent generative adversarial networks have allowed for big leaps in the realism of generated images in diverse domains, not the least of which being handwritten text generation. The generation of realistic-looking hand-written text is important because it can be used for data augmentation in handwritten text recognition (HTR) systems or human-computer interaction. We propose SmartPatch, a new technique increasing the performance of current state-of-the-art methods by augmenting the training feedback with a tailored solution to mitigate pen-level artifacts. We combine the well-known patch loss with information gathered from the parallel trained handwritten text recognition system and the separate characters of the word. This leads to a more enhanced local discriminator and results in more realistic and higher-quality generated handwritten words.
Robust partial Fourier reconstruction for diffusion-weighted imaging using a recurrent convolutional neural network
May 19, 2021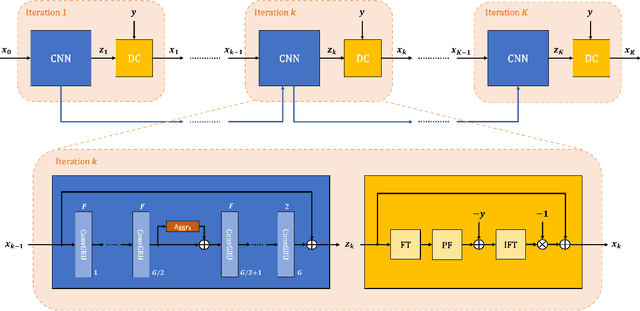
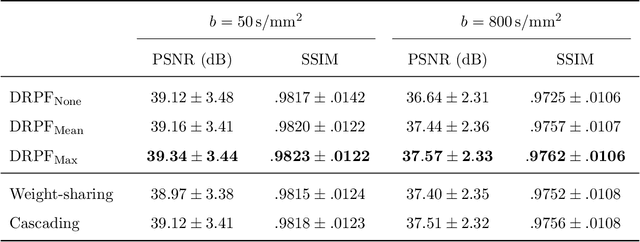
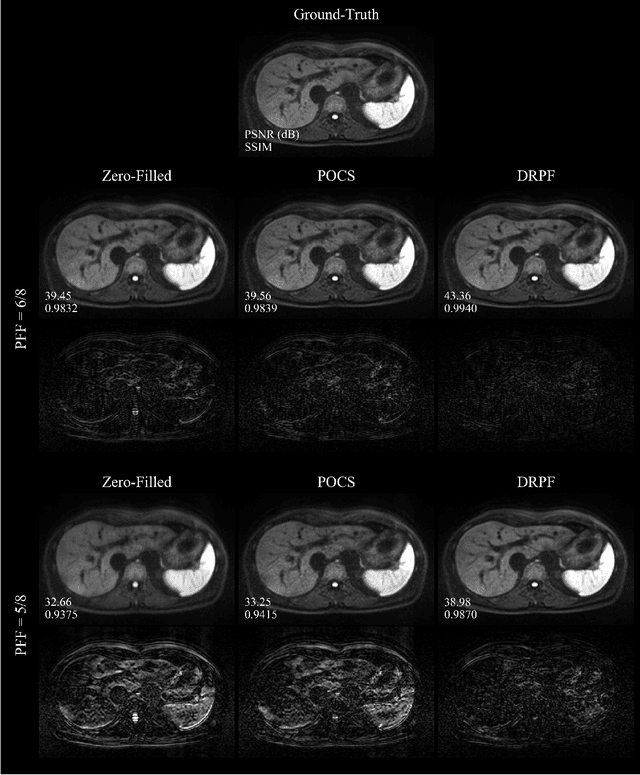
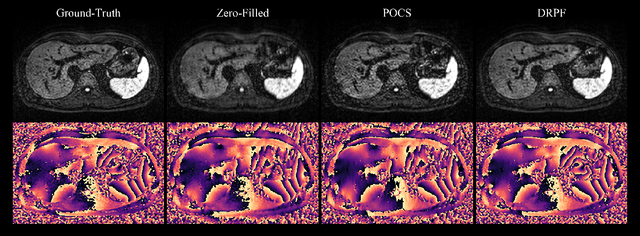
Abstract:Purpose: To develop an algorithm for robust partial Fourier (PF) reconstruction applicable to diffusion-weighted (DW) images with non-smooth phase variations. Methods: Based on an unrolled proximal splitting algorithm, a neural network architecture is derived which alternates between data consistency operations and regularization implemented by recurrent convolutions. In order to exploit correlations, multiple repetitions of the same slice are jointly reconstructed under consideration of permutation-equivariance. The proposed method is trained on DW liver data of 60 volunteers and evaluated on retrospectively and prospectively sub-sampled data of different anatomies and resolutions. In addition, the benefits of using a recurrent network over other unrolling strategies is investigated. Results: Conventional PF techniques can be significantly outperformed in terms of quantitative measures as well as perceptual image quality. The proposed method is able to generalize well to brain data with contrasts and resolution not present in the training set. The reduction in echo time (TE) associated with prospective PF-sampling enables DW imaging with higher signal. Also, the TE increase in acquisitions with higher resolution can be compensated for. It can be shown that unrolling by means of a recurrent network produced better results than using a weight-shared network or a cascade of networks. Conclusion: This work demonstrates that robust PF reconstruction of DW data is feasible even at strong PF factors in applications with severe phase variations. Since the proposed method does not rely on smoothness priors of the phase but uses learned recurrent convolutions instead, artifacts of conventional PF methods can be avoided.
Robust Classification from Noisy Labels: Integrating Additional Knowledge for Chest Radiography Abnormality Assessment
Apr 21, 2021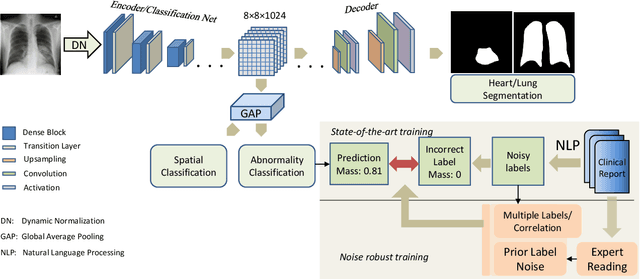
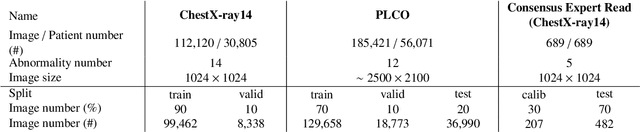
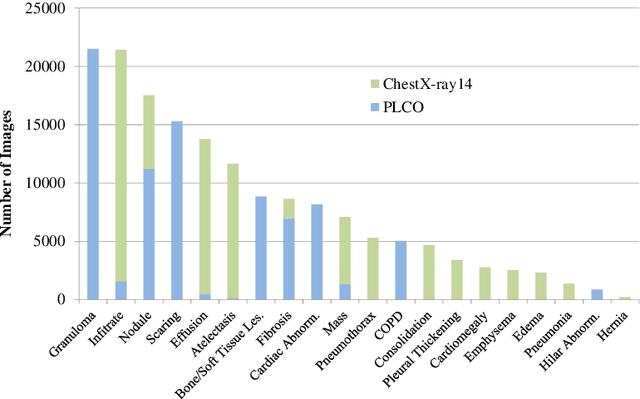
Abstract:Chest radiography is the most common radiographic examination performed in daily clinical practice for the detection of various heart and lung abnormalities. The large amount of data to be read and reported, with more than 100 studies per day for a single radiologist, poses a challenge in consistently maintaining high interpretation accuracy. The introduction of large-scale public datasets has led to a series of novel systems for automated abnormality classification. However, the labels of these datasets were obtained using natural language processed medical reports, yielding a large degree of label noise that can impact the performance. In this study, we propose novel training strategies that handle label noise from such suboptimal data. Prior label probabilities were measured on a subset of training data re-read by 4 board-certified radiologists and were used during training to increase the robustness of the training model to the label noise. Furthermore, we exploit the high comorbidity of abnormalities observed in chest radiography and incorporate this information to further reduce the impact of label noise. Additionally, anatomical knowledge is incorporated by training the system to predict lung and heart segmentation, as well as spatial knowledge labels. To deal with multiple datasets and images derived from various scanners that apply different post-processing techniques, we introduce a novel image normalization strategy. Experiments were performed on an extensive collection of 297,541 chest radiographs from 86,876 patients, leading to a state-of-the-art performance level for 17 abnormalities from 2 datasets. With an average AUC score of 0.880 across all abnormalities, our proposed training strategies can be used to significantly improve performance scores.
MR-Contrast-Aware Image-to-Image Translations with Generative Adversarial Networks
Apr 03, 2021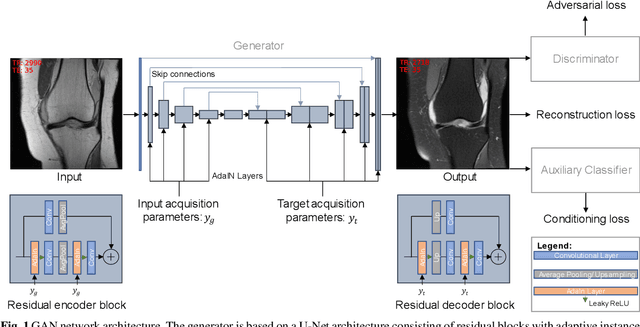
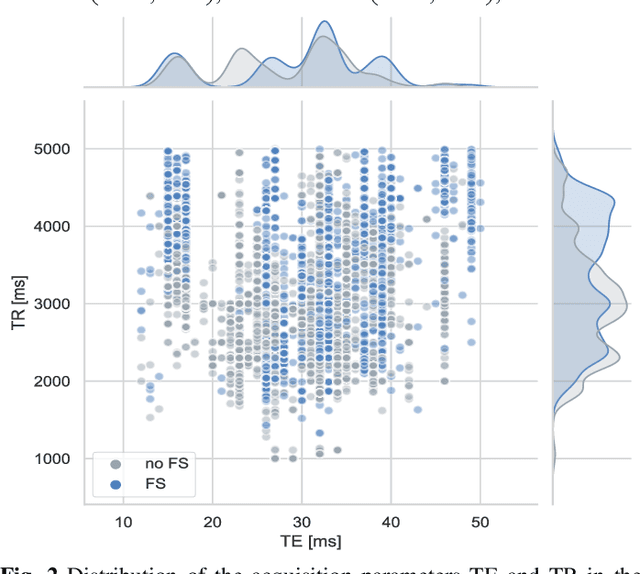
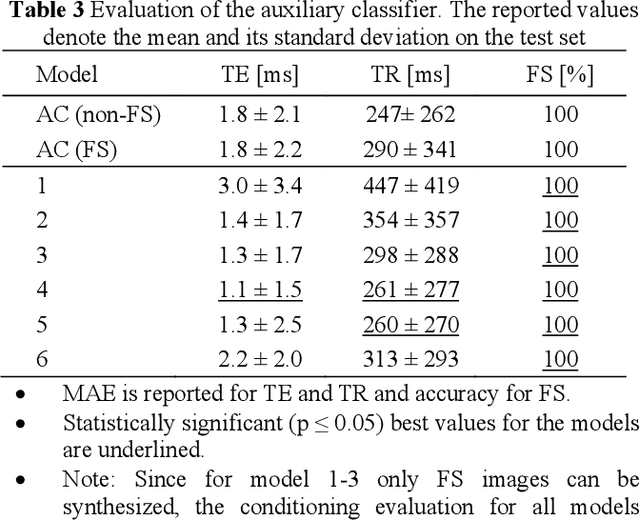
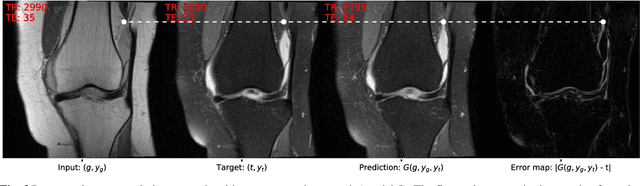
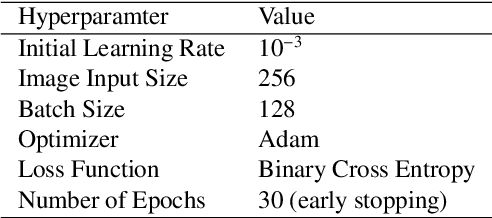
Abstract:Purpose A Magnetic Resonance Imaging (MRI) exam typically consists of several sequences that yield different image contrasts. Each sequence is parameterized through multiple acquisition parameters that influence image contrast, signal-to-noise ratio, acquisition time, and/or resolution. Depending on the clinical indication, different contrasts are required by the radiologist to make a diagnosis. As MR sequence acquisition is time consuming and acquired images may be corrupted due to motion, a method to synthesize MR images with adjustable contrast properties is required. Methods Therefore, we trained an image-to-image generative adversarial network conditioned on the MR acquisition parameters repetition time and echo time. Our approach is motivated by style transfer networks, whereas the "style" for an image is explicitly given in our case, as it is determined by the MR acquisition parameters our network is conditioned on. Results This enables us to synthesize MR images with adjustable image contrast. We evaluated our approach on the fastMRI dataset, a large set of publicly available MR knee images, and show that our method outperforms a benchmark pix2pix approach in the translation of non-fat-saturated MR images to fat-saturated images. Our approach yields a peak signal-to-noise ratio and structural similarity of 24.48 and 0.66, surpassing the pix2pix benchmark model significantly. Conclusion Our model is the first that enables fine-tuned contrast synthesis, which can be used to synthesize missing MR contrasts or as a data augmentation technique for AI training in MRI.
Quantifying the Scanner-Induced Domain Gap in Mitosis Detection
Mar 30, 2021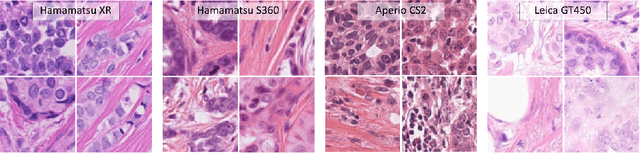
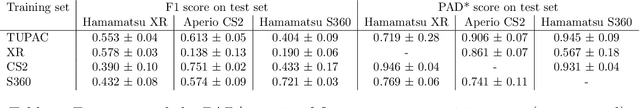

Abstract:Automated detection of mitotic figures in histopathology images has seen vast improvements, thanks to modern deep learning-based pipelines. Application of these methods, however, is in practice limited by strong variability of images between labs. This results in a domain shift of the images, which causes a performance drop of the models. Hypothesizing that the scanner device plays a decisive role in this effect, we evaluated the susceptibility of a standard mitosis detection approach to the domain shift introduced by using a different whole slide scanner. Our work is based on the MICCAI-MIDOG challenge 2021 data set, which includes 200 tumor cases of human breast cancer and four scanners. Our work indicates that the domain shift induced not by biochemical variability but purely by the choice of acquisition device is underestimated so far. Models trained on images of the same scanner yielded an average F1 score of 0.683, while models trained on a single other scanner only yielded an average F1 score of 0.325. Training on another multi-domain mitosis dataset led to mean F1 scores of 0.52. We found this not to be reflected by domain-shifts measured as proxy A distance-derived metric.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge