Alan L. Yuille
Elastic Boundary Projection for 3D Medical Imaging Segmentation
Dec 03, 2018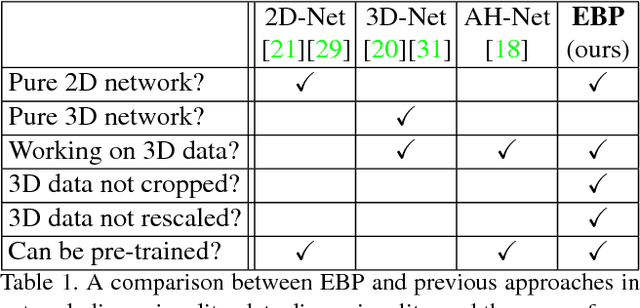
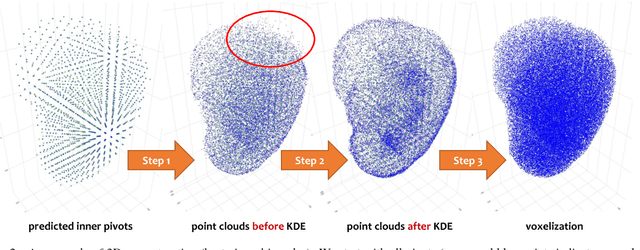
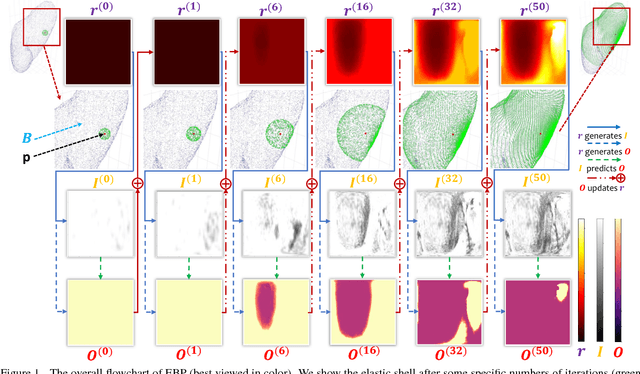

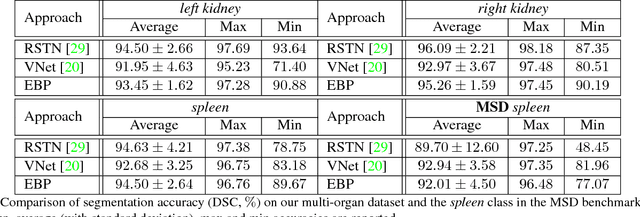
Abstract:We focus on an important yet challenging problem: using a 2D deep network to deal with 3D segmentation for medical imaging analysis. Existing approaches either applied multi-view planar (2D) networks or directly used volumetric (3D) networks for this purpose, but both of them are not ideal: 2D networks cannot capture 3D contexts effectively, and 3D networks are both memory-consuming and less stable arguably due to the lack of pre-trained models. In this paper, we bridge the gap between 2D and 3D using a novel approach named Elastic Boundary Projection (EBP). The key observation is that, although the object is a 3D volume, what we really need in segmentation is to find its boundary which is a 2D surface. Therefore, we place a number of pivot points in the 3D space, and for each pivot, we determine its distance to the object boundary along a dense set of directions. This creates an elastic shell around each pivot which is initialized as a perfect sphere. We train a 2D deep network to determine whether each ending point falls within the object, and gradually adjust the shell so that it gradually converges to the actual shape of the boundary and thus achieves the goal of segmentation. EBP allows 3D segmentation without cutting the volume into slices or small patches, which stands out from conventional 2D and 3D approaches. EBP achieves promising accuracy in segmenting several abdominal organs from CT scans.
Iterative Reorganization with Weak Spatial Constraints: Solving Arbitrary Jigsaw Puzzles for Unsupervised Representation Learning
Dec 02, 2018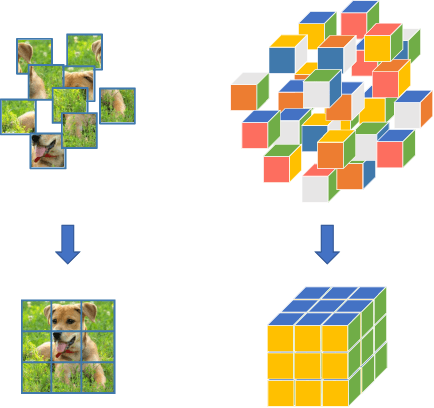
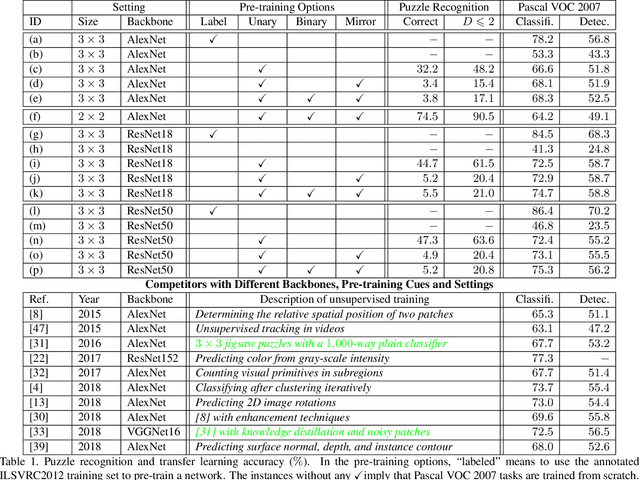
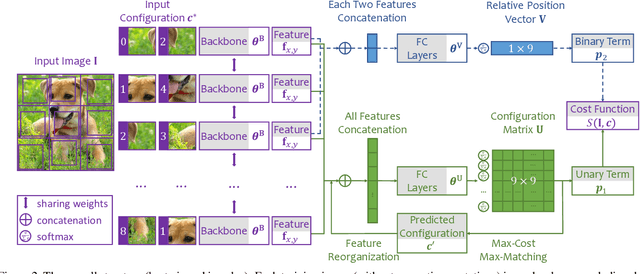
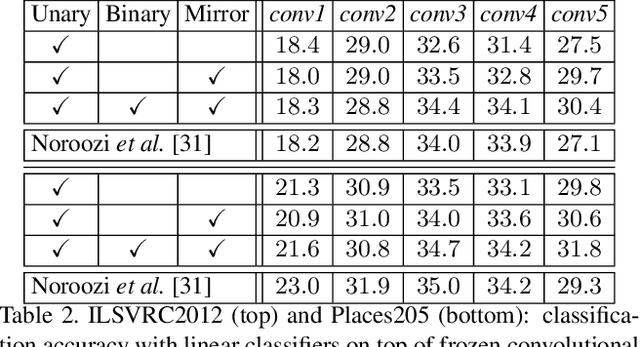
Abstract:Learning visual features from unlabeled image data is an important yet challenging task, which is often achieved by training a model on some annotation-free information. We consider spatial contexts, for which we solve so-called jigsaw puzzles, i.e., each image is cut into grids and then disordered, and the goal is to recover the correct configuration. Existing approaches formulated it as a classification task by defining a fixed mapping from a small subset of configurations to a class set, but these approaches ignore the underlying relationship between different configurations and also limit their application to more complex scenarios. This paper presents a novel approach which applies to jigsaw puzzles with an arbitrary grid size and dimensionality. We provide a fundamental and generalized principle, that weaker cues are easier to be learned in an unsupervised manner and also transfer better. In the context of puzzle recognition, we use an iterative manner which, instead of solving the puzzle all at once, adjusts the order of the patches in each step until convergence. In each step, we combine both unary and binary features on each patch into a cost function judging the correctness of the current configuration. Our approach, by taking similarity between puzzles into consideration, enjoys a more reasonable way of learning visual knowledge. We verify the effectiveness of our approach in two aspects. First, it is able to solve arbitrarily complex puzzles, including high-dimensional puzzles, that prior methods are difficult to handle. Second, it serves as a reliable way of network initialization, which leads to better transfer performance in a few visual recognition tasks including image classification, object detection, and semantic segmentation.
Snapshot Distillation: Teacher-Student Optimization in One Generation
Dec 01, 2018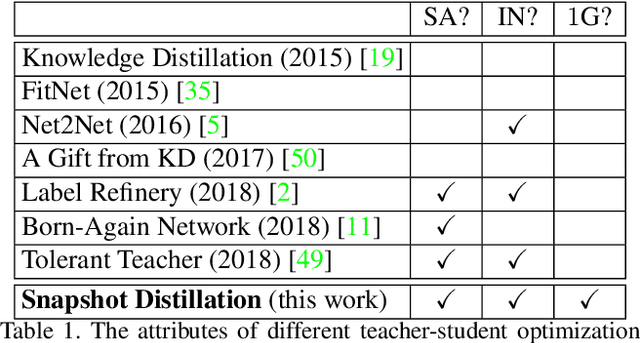
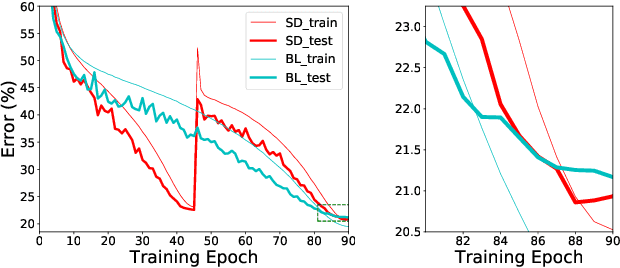
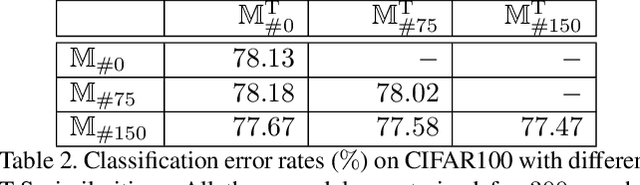
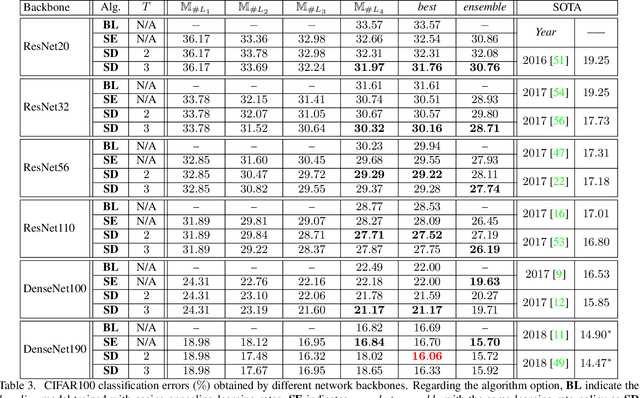
Abstract:Optimizing a deep neural network is a fundamental task in computer vision, yet direct training methods often suffer from over-fitting. Teacher-student optimization aims at providing complementary cues from a model trained previously, but these approaches are often considerably slow due to the pipeline of training a few generations in sequence, i.e., time complexity is increased by several times. This paper presents snapshot distillation (SD), the first framework which enables teacher-student optimization in one generation. The idea of SD is very simple: instead of borrowing supervision signals from previous generations, we extract such information from earlier epochs in the same generation, meanwhile make sure that the difference between teacher and student is sufficiently large so as to prevent under-fitting. To achieve this goal, we implement SD in a cyclic learning rate policy, in which the last snapshot of each cycle is used as the teacher for all iterations in the next cycle, and the teacher signal is smoothed to provide richer information. In standard image classification benchmarks such as CIFAR100 and ILSVRC2012, SD achieves consistent accuracy gain without heavy computational overheads. We also verify that models pre-trained with SD transfers well to object detection and semantic segmentation in the PascalVOC dataset.
Progressive Recurrent Learning for Visual Recognition
Nov 29, 2018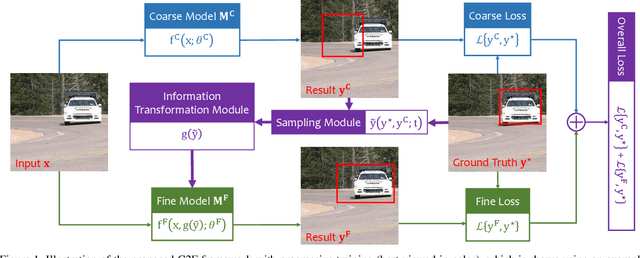
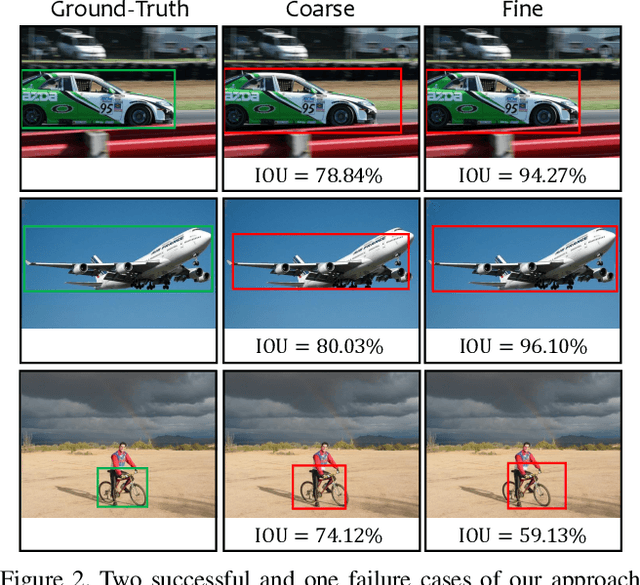
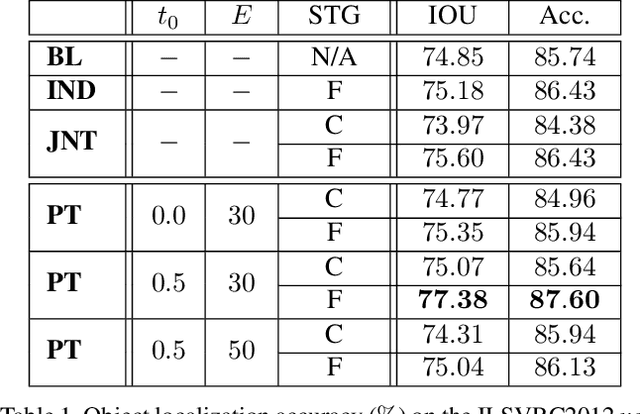

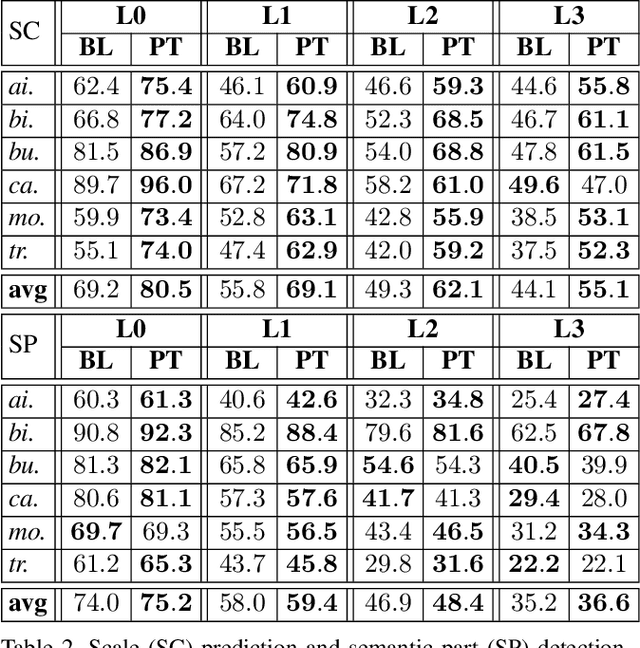
Abstract:Computer vision is difficult, partly because the mathematical function connecting input and output data is often complex, fuzzy and thus hard to learn. A currently popular solution is to design a deep neural network and optimize it on a large-scale dataset. However, as the number of parameters increases, the generalization ability is often not guaranteed, e.g., the model can over-fit due to the limited amount of training data, or fail to converge because the desired function is too difficult to learn. This paper presents an effective framework named progressive recurrent learning (PRL). The core idea is similar to curriculum learning which gradually increases the difficulty of training data. We generalize it to a wide range of vision problems that were previously considered less proper to apply curriculum learning. PRL starts with inserting a recurrent prediction scheme, based on the motivation of feeding the prediction of a vision model to the same model iteratively, so that the auxiliary cues contained in it can be exploited to improve the quality of itself. In order to better optimize this framework, we start with providing perfect prediction, i.e., ground-truth, to the second stage, but gradually replace it with the prediction of the first stage. In the final status, the ground-truth information is not needed any more, so that the entire model works on the real data distribution as in the testing process. We apply PRL to two challenging visual recognition tasks, namely, object localization and semantic segmentation, and demonstrate consistent accuracy gain compared to the baseline training strategy, especially in the scenarios of more difficult vision tasks.
PreCo: A Large-scale Dataset in Preschool Vocabulary for Coreference Resolution
Oct 23, 2018
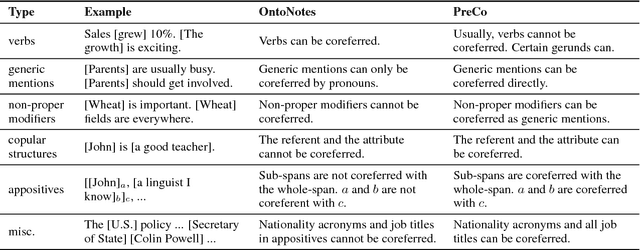
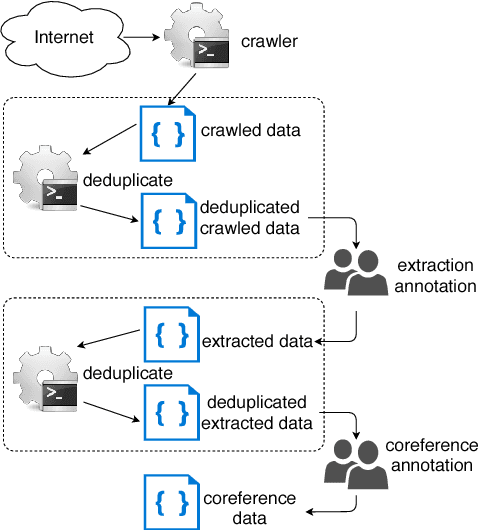
Abstract:We introduce PreCo, a large-scale English dataset for coreference resolution. The dataset is designed to embody the core challenges in coreference, such as entity representation, by alleviating the challenge of low overlap between training and test sets and enabling separated analysis of mention detection and mention clustering. To strengthen the training-test overlap, we collect a large corpus of about 38K documents and 12.4M words which are mostly from the vocabulary of English-speaking preschoolers. Experiments show that with higher training-test overlap, error analysis on PreCo is more efficient than the one on OntoNotes, a popular existing dataset. Furthermore, we annotate singleton mentions making it possible for the first time to quantify the influence that a mention detector makes on coreference resolution performance. The dataset is freely available at https://preschool-lab.github.io/PreCo/.
Adversarial Attacks Beyond the Image Space
Sep 10, 2018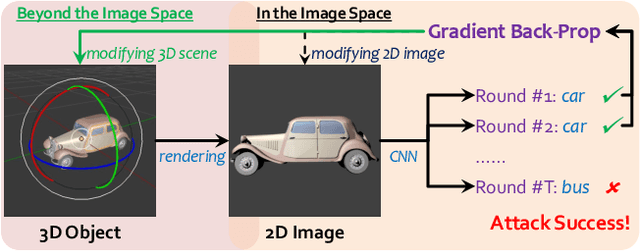
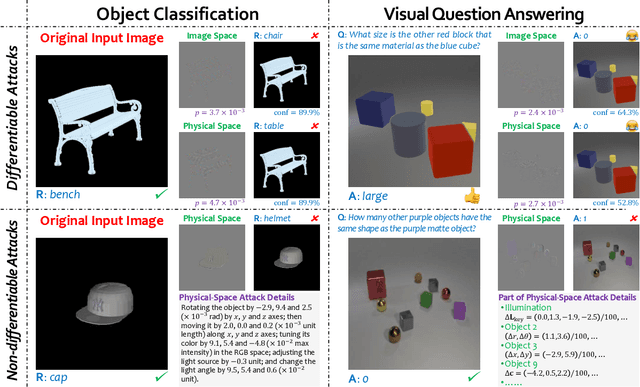
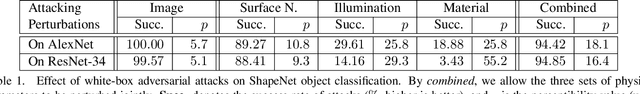

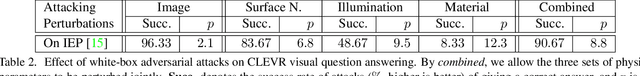
Abstract:Generating adversarial examples is an intriguing problem and an important way of understanding the working mechanism of deep neural networks. Most existing approaches generated perturbations in the image space, i.e., each pixel can be modified independently. However, in this paper we pay special attention to the subset of adversarial examples that are physically authentic -- those corresponding to actual changes in 3D physical properties (like surface normals, illumination condition, etc.). These adversaries arguably pose a more serious concern, as they demonstrate the possibility of causing neural network failure by small perturbations of real-world 3D objects and scenes. In the contexts of object classification and visual question answering, we augment state-of-the-art deep neural networks that receive 2D input images with a rendering module (either differentiable or not) in front, so that a 3D scene (in the physical space) is rendered into a 2D image (in the image space), and then mapped to a prediction (in the output space). The adversarial perturbations can now go beyond the image space, and have clear meanings in the 3D physical world. Through extensive experiments, we found that a vast majority of image-space adversaries cannot be explained by adjusting parameters in the physical space, i.e., they are usually physically inauthentic. But it is still possible to successfully attack beyond the image space on the physical space (such that authenticity is enforced), though this is more difficult than image-space attacks, reflected in lower success rates and heavier perturbations required.
A 3D Coarse-to-Fine Framework for Volumetric Medical Image Segmentation
Aug 02, 2018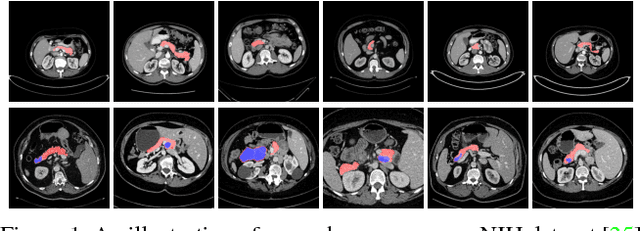
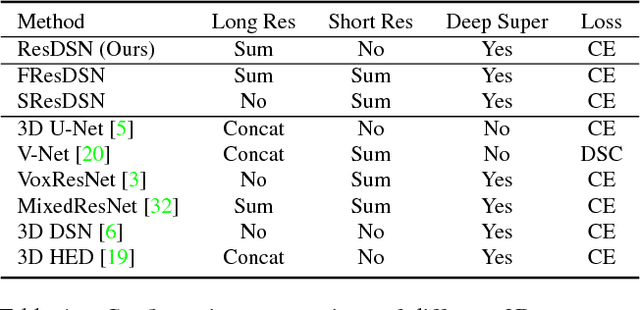

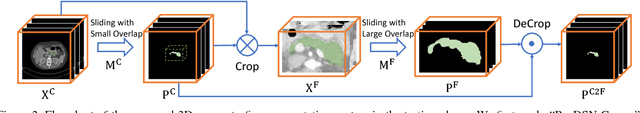
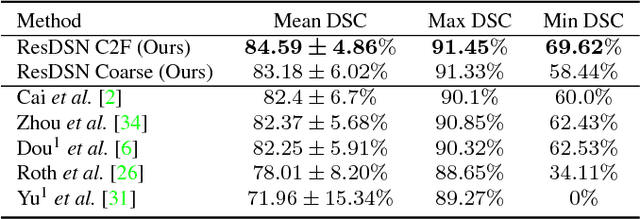
Abstract:In this paper, we adopt 3D Convolutional Neural Networks to segment volumetric medical images. Although deep neural networks have been proven to be very effective on many 2D vision tasks, it is still challenging to apply them to 3D tasks due to the limited amount of annotated 3D data and limited computational resources. We propose a novel 3D-based coarse-to-fine framework to effectively and efficiently tackle these challenges. The proposed 3D-based framework outperforms the 2D counterpart to a large margin since it can leverage the rich spatial infor- mation along all three axes. We conduct experiments on two datasets which include healthy and pathological pancreases respectively, and achieve the current state-of-the-art in terms of Dice-S{\o}rensen Coefficient (DSC). On the NIH pancreas segmentation dataset, we outperform the previous best by an average of over 2%, and the worst case is improved by 7% to reach almost 70%, which indicates the reliability of our framework in clinical applications.
Multi-Scale Coarse-to-Fine Segmentation for Screening Pancreatic Ductal Adenocarcinoma
Jul 09, 2018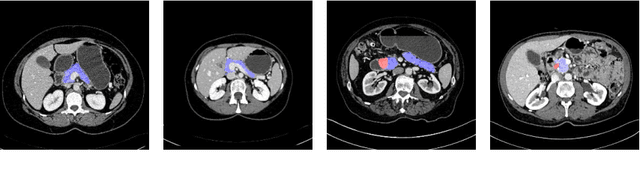

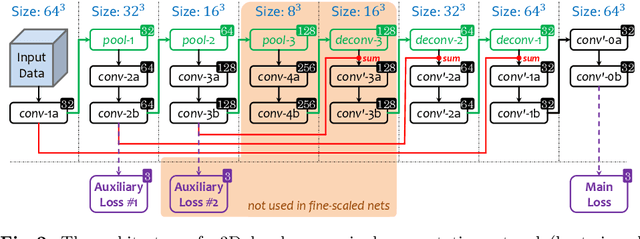
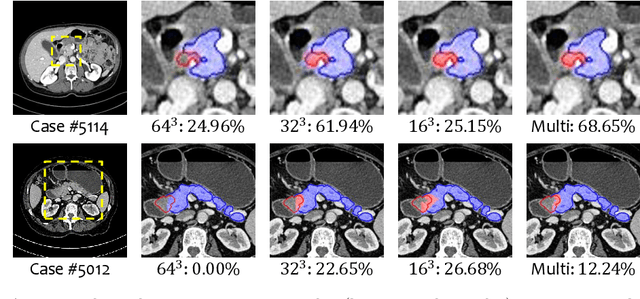
Abstract:This paper proposes an intuitive approach to finding pancreatic ductal adenocarcinoma (PDAC), the most common type of pancreatic cancer, by checking abdominal CT scans. Our idea is named segmentation-for-classification (S4C), which classifies a volume by checking if at least a sufficient number of voxels is segmented as the tumor. In order to deal with tumors with different scales, we train volumetric segmentation networks with multi-scale inputs, and test them in a coarse-to-fine flowchart. A post-processing module is used to filter out outliers and reduce false alarms. We perform a case study on our dataset containing 439 CT scans, in which 136 cases were diagnosed with PDAC and 303 cases are normal. Our approach reports a sensitivity of 94.1% at a specificity of 98.5%, with an average tumor segmentation accuracy of 56.46% over all PDAC cases.
Bridging the Gap Between 2D and 3D Organ Segmentation with Volumetric Fusion Net
Jun 09, 2018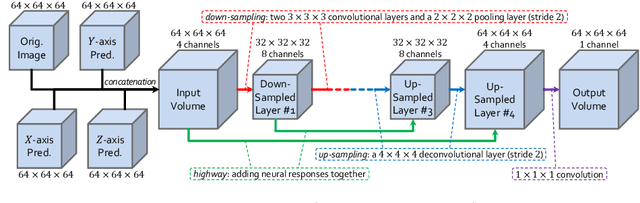
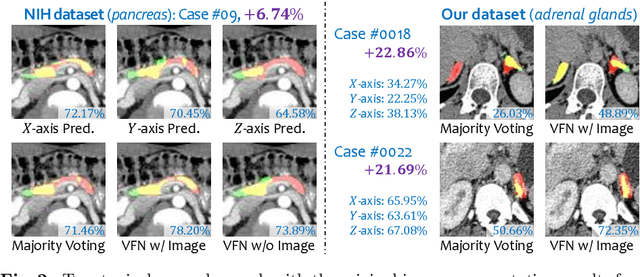
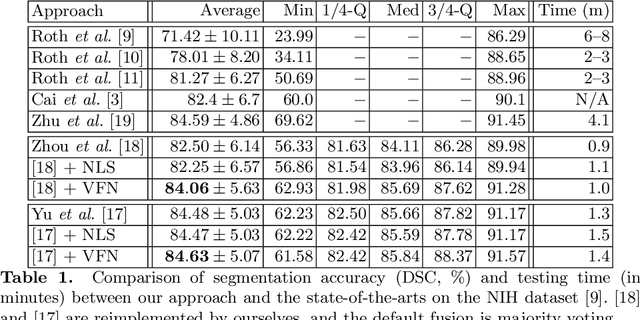

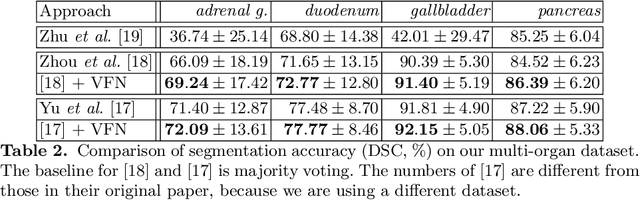
Abstract:There has been a debate on whether to use 2D or 3D deep neural networks for volumetric organ segmentation. Both 2D and 3D models have their advantages and disadvantages. In this paper, we present an alternative framework, which trains 2D networks on different viewpoints for segmentation, and builds a 3D Volumetric Fusion Net (VFN) to fuse the 2D segmentation results. VFN is relatively shallow and contains much fewer parameters than most 3D networks, making our framework more efficient at integrating 3D information for segmentation. We train and test the segmentation and fusion modules individually, and propose a novel strategy, named cross-cross-augmentation, to make full use of the limited training data. We evaluate our framework on several challenging abdominal organs, and verify its superiority in segmentation accuracy and stability over existing 2D and 3D approaches.
Semi-Supervised Multi-Organ Segmentation via Deep Multi-Planar Co-Training
May 12, 2018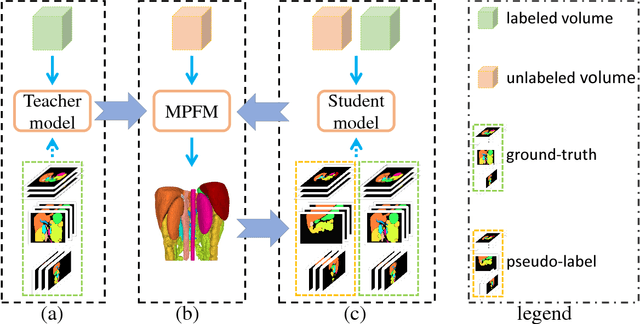
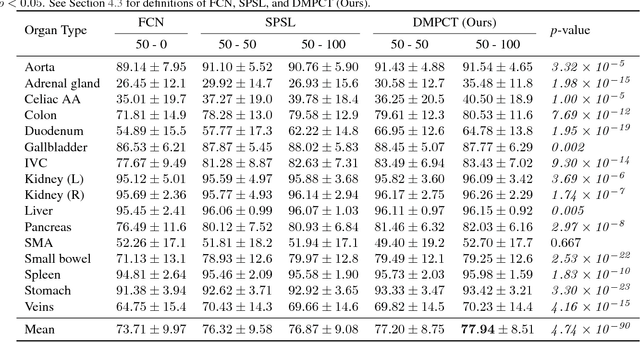
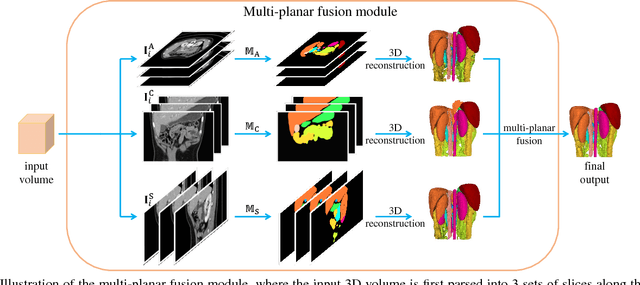
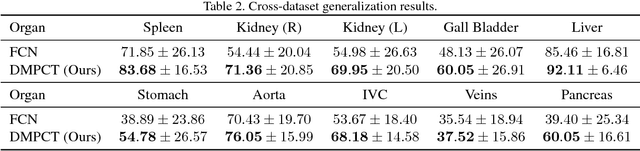
Abstract:In multi-organ segmentation of abdominal CT scans, most existing fully supervised deep learning algorithms require lots of voxel-wise annotations, which are usually difficult, expensive, and slow to obtain. In comparison, massive unlabeled 3D CT volumes are usually easily accessible. Current mainstream works to address the semi-supervised biomedical image segmentation problem are mostly graph-based. By contrast, deep network based semi-supervised learning methods have not drawn much attention in this field. In this work, we propose Deep Multi-Planar Co-Training (DMPCT), whose contributions can be divided into two folds: 1) The deep model is learned in a co-training style which can mine consensus information from multiple planes like the sagittal, coronal, and axial planes; 2) Multi-planar fusion is applied to generate more reliable pseudo-labels, which alleviates the errors occurring in the pseudo-labels and thus can help to train better segmentation networks. Experiments are done on our newly collected large dataset with 100 unlabeled cases as well as 210 labeled cases where 16 anatomical structures are manually annotated by four radiologists and confirmed by a senior expert. The results suggest that DMPCT significantly outperforms the fully supervised method by more than 4% especially when only a small set of annotations is used.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge