Adam E. Flanders
RSNA Large Language Model Benchmark Dataset for Chest Radiographs of Cardiothoracic Disease: Radiologist Evaluation and Validation Enhanced by AI Labels (REVEAL-CXR)
Jan 21, 2026Abstract:Multimodal large language models have demonstrated comparable performance to that of radiology trainees on multiple-choice board-style exams. However, to develop clinically useful multimodal LLM tools, high-quality benchmarks curated by domain experts are essential. To curate released and holdout datasets of 100 chest radiographic studies each and propose an artificial intelligence (AI)-assisted expert labeling procedure to allow radiologists to label studies more efficiently. A total of 13,735 deidentified chest radiographs and their corresponding reports from the MIDRC were used. GPT-4o extracted abnormal findings from the reports, which were then mapped to 12 benchmark labels with a locally hosted LLM (Phi-4-Reasoning). From these studies, 1,000 were sampled on the basis of the AI-suggested benchmark labels for expert review; the sampling algorithm ensured that the selected studies were clinically relevant and captured a range of difficulty levels. Seventeen chest radiologists participated, and they marked "Agree all", "Agree mostly" or "Disagree" to indicate their assessment of the correctness of the LLM suggested labels. Each chest radiograph was evaluated by three experts. Of these, at least two radiologists selected "Agree All" for 381 radiographs. From this set, 200 were selected, prioritizing those with less common or multiple finding labels, and divided into 100 released radiographs and 100 reserved as the holdout dataset. The holdout dataset is used exclusively by RSNA to independently evaluate different models. A benchmark of 200 chest radiographic studies with 12 benchmark labels was created and made publicly available https://imaging.rsna.org, with each chest radiograph verified by three radiologists. In addition, an AI-assisted labeling procedure was developed to help radiologists label at scale, minimize unnecessary omissions, and support a semicollaborative environment.
A Multi-agent Large Language Model Framework to Automatically Assess Performance of a Clinical AI Triage Tool
Oct 30, 2025Abstract:Purpose: The purpose of this study was to determine if an ensemble of multiple LLM agents could be used collectively to provide a more reliable assessment of a pixel-based AI triage tool than a single LLM. Methods: 29,766 non-contrast CT head exams from fourteen hospitals were processed by a commercial intracranial hemorrhage (ICH) AI detection tool. Radiology reports were analyzed by an ensemble of eight open-source LLM models and a HIPAA compliant internal version of GPT-4o using a single multi-shot prompt that assessed for presence of ICH. 1,726 examples were manually reviewed. Performance characteristics of the eight open-source models and consensus were compared to GPT-4o. Three ideal consensus LLM ensembles were tested for rating the performance of the triage tool. Results: The cohort consisted of 29,766 head CTs exam-report pairs. The highest AUC performance was achieved with llama3.3:70b and GPT-4o (AUC= 0.78). The average precision was highest for Llama3.3:70b and GPT-4o (AP=0.75 & 0.76). Llama3.3:70b had the highest F1 score (0.81) and recall (0.85), greater precision (0.78), specificity (0.72), and MCC (0.57). Using MCC (95% CI) the ideal combination of LLMs were: Full-9 Ensemble 0.571 (0.552-0.591), Top-3 Ensemble 0.558 (0.537-0.579), Consensus 0.556 (0.539-0.574), and GPT4o 0.522 (0.500-0.543). No statistically significant differences were observed between Top-3, Full-9, and Consensus (p > 0.05). Conclusion: An ensemble of medium to large sized open-source LLMs provides a more consistent and reliable method to derive a ground truth retrospective evaluation of a clinical AI triage tool over a single LLM alone.
The RSNA Lumbar Degenerative Imaging Spine Classification (LumbarDISC) Dataset
Jun 10, 2025Abstract:The Radiological Society of North America (RSNA) Lumbar Degenerative Imaging Spine Classification (LumbarDISC) dataset is the largest publicly available dataset of adult MRI lumbar spine examinations annotated for degenerative changes. The dataset includes 2,697 patients with a total of 8,593 image series from 8 institutions across 6 countries and 5 continents. The dataset is available for free for non-commercial use via Kaggle and RSNA Medical Imaging Resource of AI (MIRA). The dataset was created for the RSNA 2024 Lumbar Spine Degenerative Classification competition where competitors developed deep learning models to grade degenerative changes in the lumbar spine. The degree of spinal canal, subarticular recess, and neural foraminal stenosis was graded at each intervertebral disc level in the lumbar spine. The images were annotated by expert volunteer neuroradiologists and musculoskeletal radiologists from the RSNA, American Society of Neuroradiology, and the American Society of Spine Radiology. This dataset aims to facilitate research and development in machine learning and lumbar spine imaging to lead to improved patient care and clinical efficiency.
The RSNA Abdominal Traumatic Injury CT (RATIC) Dataset
May 30, 2024



Abstract:The RSNA Abdominal Traumatic Injury CT (RATIC) dataset is the largest publicly available collection of adult abdominal CT studies annotated for traumatic injuries. This dataset includes 4,274 studies from 23 institutions across 14 countries. The dataset is freely available for non-commercial use via Kaggle at https://www.kaggle.com/competitions/rsna-2023-abdominal-trauma-detection. Created for the RSNA 2023 Abdominal Trauma Detection competition, the dataset encourages the development of advanced machine learning models for detecting abdominal injuries on CT scans. The dataset encompasses detection and classification of traumatic injuries across multiple organs, including the liver, spleen, kidneys, bowel, and mesentery. Annotations were created by expert radiologists from the American Society of Emergency Radiology (ASER) and Society of Abdominal Radiology (SAR). The dataset is annotated at multiple levels, including the presence of injuries in three solid organs with injury grading, image-level annotations for active extravasations and bowel injury, and voxelwise segmentations of each of the potentially injured organs. With the release of this dataset, we hope to facilitate research and development in machine learning and abdominal trauma that can lead to improved patient care and outcomes.
Analysis of the BraTS 2023 Intracranial Meningioma Segmentation Challenge
May 16, 2024



Abstract:We describe the design and results from the BraTS 2023 Intracranial Meningioma Segmentation Challenge. The BraTS Meningioma Challenge differed from prior BraTS Glioma challenges in that it focused on meningiomas, which are typically benign extra-axial tumors with diverse radiologic and anatomical presentation and a propensity for multiplicity. Nine participating teams each developed deep-learning automated segmentation models using image data from the largest multi-institutional systematically expert annotated multilabel multi-sequence meningioma MRI dataset to date, which included 1000 training set cases, 141 validation set cases, and 283 hidden test set cases. Each case included T2, T2/FLAIR, T1, and T1Gd brain MRI sequences with associated tumor compartment labels delineating enhancing tumor, non-enhancing tumor, and surrounding non-enhancing T2/FLAIR hyperintensity. Participant automated segmentation models were evaluated and ranked based on a scoring system evaluating lesion-wise metrics including dice similarity coefficient (DSC) and 95% Hausdorff Distance. The top ranked team had a lesion-wise median dice similarity coefficient (DSC) of 0.976, 0.976, and 0.964 for enhancing tumor, tumor core, and whole tumor, respectively and a corresponding average DSC of 0.899, 0.904, and 0.871, respectively. These results serve as state-of-the-art benchmarks for future pre-operative meningioma automated segmentation algorithms. Additionally, we found that 1286 of 1424 cases (90.3%) had at least 1 compartment voxel abutting the edge of the skull-stripped image edge, which requires further investigation into optimal pre-processing face anonymization steps.
The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification
Jul 05, 2021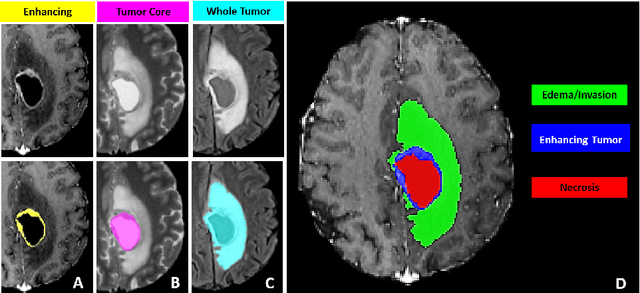
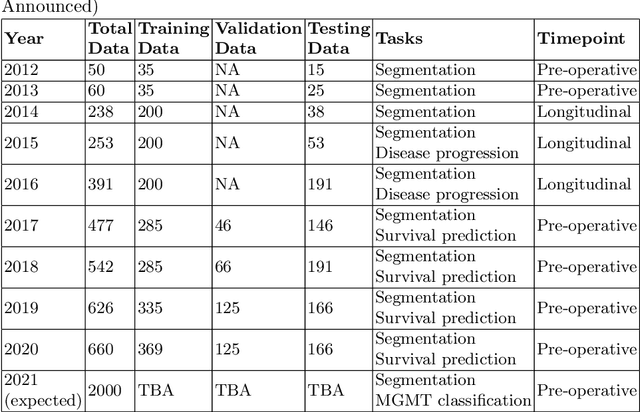
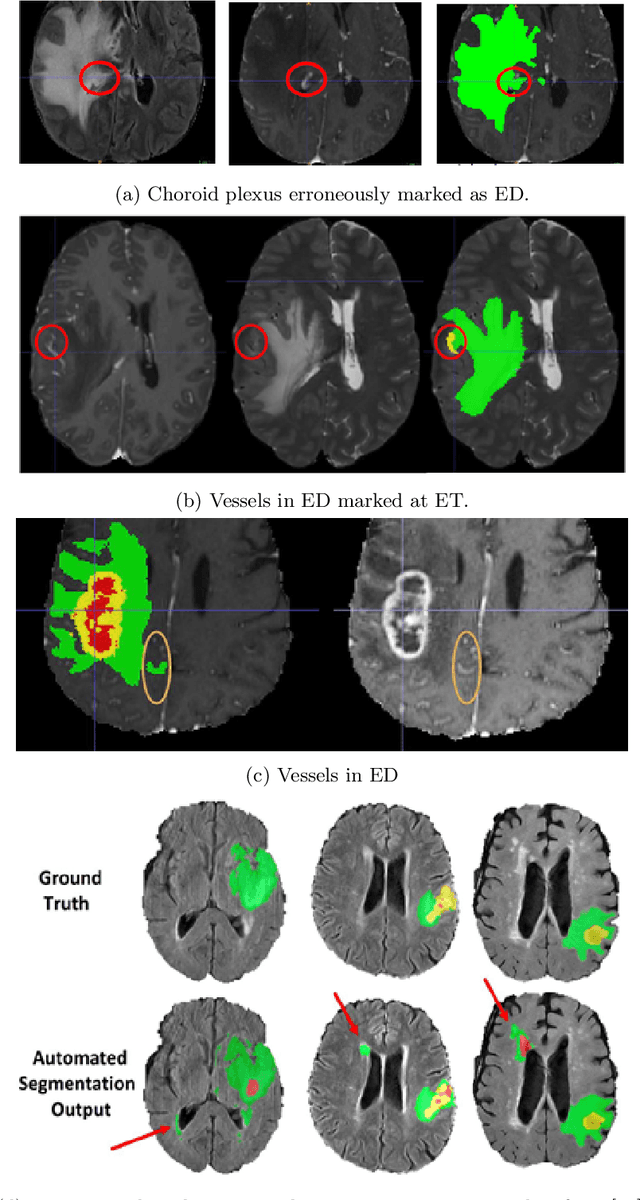
Abstract:The BraTS 2021 challenge celebrates its 10th anniversary and is jointly organized by the Radiological Society of North America (RSNA), the American Society of Neuroradiology (ASNR), and the Medical Image Computing and Computer Assisted Interventions (MICCAI) society. Since its inception, BraTS has been focusing on being a common benchmarking venue for brain glioma segmentation algorithms, with well-curated multi-institutional multi-parametric magnetic resonance imaging (mpMRI) data. Gliomas are the most common primary malignancies of the central nervous system, with varying degrees of aggressiveness and prognosis. The RSNA-ASNR-MICCAI BraTS 2021 challenge targets the evaluation of computational algorithms assessing the same tumor compartmentalization, as well as the underlying tumor's molecular characterization, in pre-operative baseline mpMRI data from 2,000 patients. Specifically, the two tasks that BraTS 2021 focuses on are: a) the segmentation of the histologically distinct brain tumor sub-regions, and b) the classification of the tumor's O[6]-methylguanine-DNA methyltransferase (MGMT) promoter methylation status. The performance evaluation of all participating algorithms in BraTS 2021 will be conducted through the Sage Bionetworks Synapse platform (Task 1) and Kaggle (Task 2), concluding in distributing to the top ranked participants monetary awards of $60,000 collectively.
Application of Deep Learning in Neuroradiology: Automated Detection of Basal Ganglia Hemorrhage using 2D-Convolutional Neural Networks
Oct 27, 2017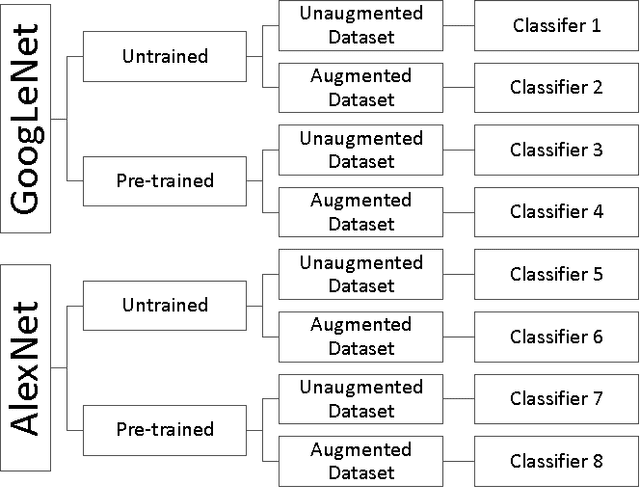
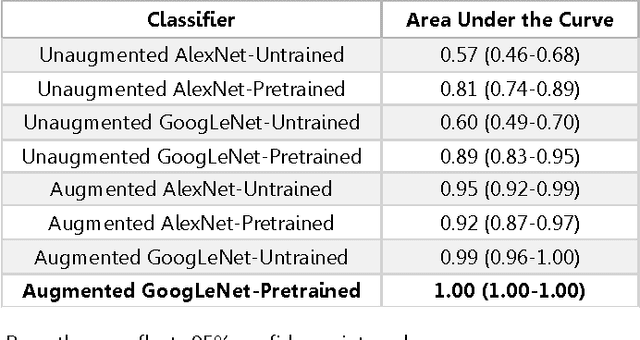
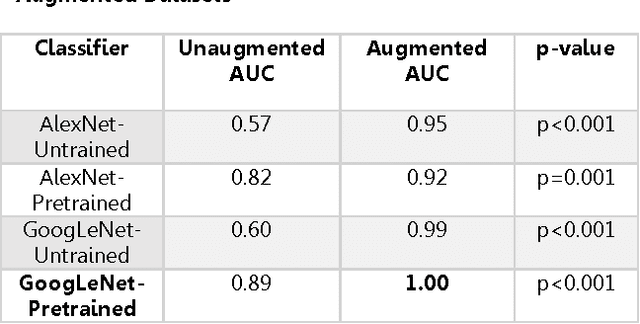
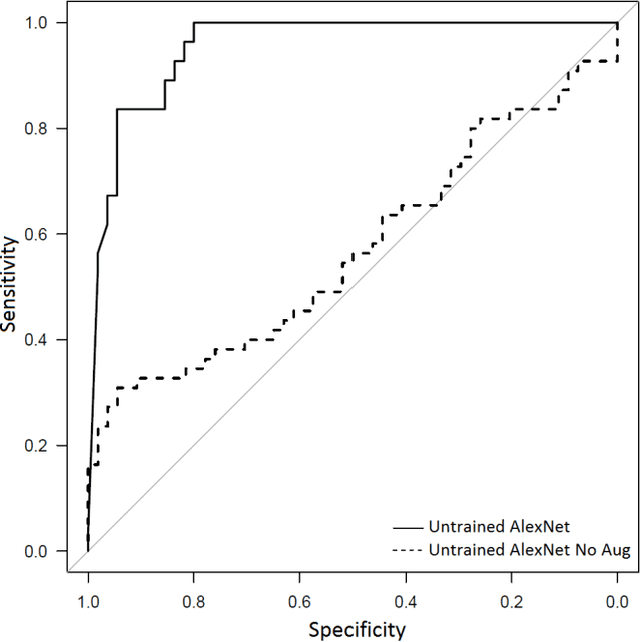
Abstract:Background: Deep learning techniques have achieved high accuracy in image classification tasks, and there is interest in applicability to neuroimaging critical findings. This study evaluates the efficacy of 2D deep convolutional neural networks (DCNNs) for detecting basal ganglia (BG) hemorrhage on noncontrast head CT. Materials and Methods: 170 unique de-identified HIPAA-compliant noncontrast head CTs were obtained, those with and without BG hemorrhage. 110 cases were held-out for test, and 60 were split into training (45) and validation (15), consisting of 20 right, 20 left, and 20 no BG hemorrhage. Data augmentation was performed to increase size and variation of the training dataset by 48-fold. Two DCNNs were used to classify the images-AlexNet and GoogLeNet-using untrained networks and those pre-trained on ImageNet. Area under the curves (AUC) for the receiver-operator characteristic (ROC) curves were calculated, using the DeLong method for statistical comparison of ROCs. Results: The best performing model was the pre-trained augmented GoogLeNet, which had an AUC of 1.00 in classification of hemorrhage. Preprocessing augmentation increased accuracy for all networks (p<0.001), and pretrained networks outperformed untrained ones (p<0.001) for the unaugmented models. The best performing GoogLeNet model (AUC 1.00) outperformed the best performing AlexNet model (AUC 0.95)(p=0.01). Conclusion: For this dataset, the best performing DCNN identified BG hemorrhage on noncontrast head CT with an AUC of 1.00. Pretrained networks and data augmentation increased classifier accuracy. Future prospective research would be important to determine if the accuracy can be maintained on a larger cohort of patients and for very small hemorrhages.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge