"Image": models, code, and papers
Multiple resolution residual network for automatic thoracic organs-at-risk segmentation from CT
May 27, 2020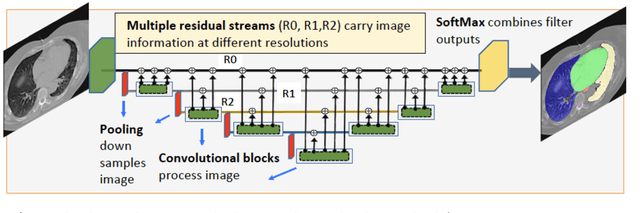
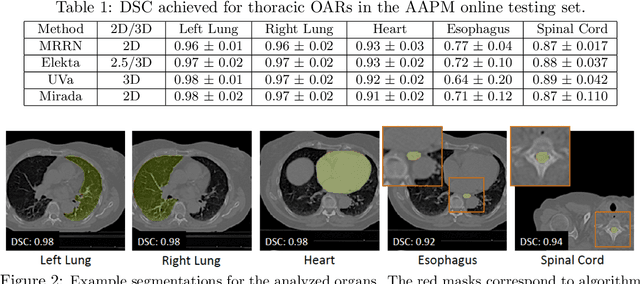
We implemented and evaluated a multiple resolution residual network (MRRN) for multiple normal organs-at-risk (OAR) segmentation from computed tomography (CT) images for thoracic radiotherapy treatment (RT) planning. Our approach simultaneously combines feature streams computed at multiple image resolutions and feature levels through residual connections. The feature streams at each level are updated as the images are passed through various feature levels. We trained our approach using 206 thoracic CT scans of lung cancer patients with 35 scans held out for validation to segment the left and right lungs, heart, esophagus, and spinal cord. This approach was tested on 60 CT scans from the open-source AAPM Thoracic Auto-Segmentation Challenge dataset. Performance was measured using the Dice Similarity Coefficient (DSC). Our approach outperformed the best-performing method in the grand challenge for hard-to-segment structures like the esophagus and achieved comparable results for all other structures. Median DSC using our method was 0.97 (interquartile range [IQR]: 0.97-0.98) for the left and right lungs, 0.93 (IQR: 0.93-0.95) for the heart, 0.78 (IQR: 0.76-0.80) for the esophagus, and 0.88 (IQR: 0.86-0.89) for the spinal cord.
Learning crystal plasticity using digital image correlation: Examples from discrete dislocation dynamics
Sep 24, 2017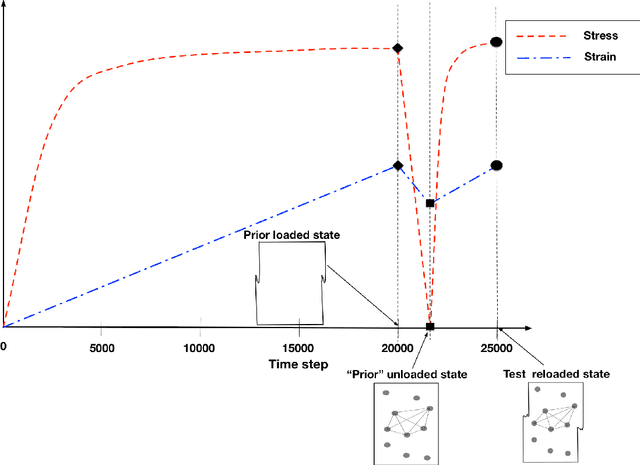
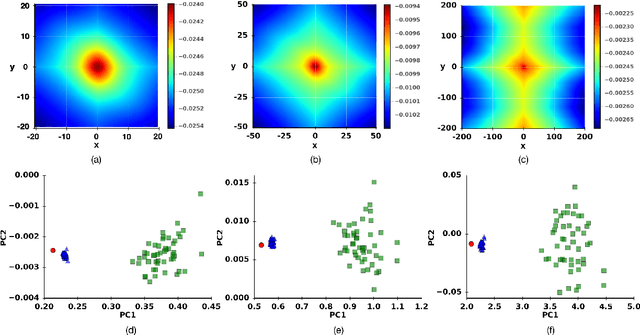
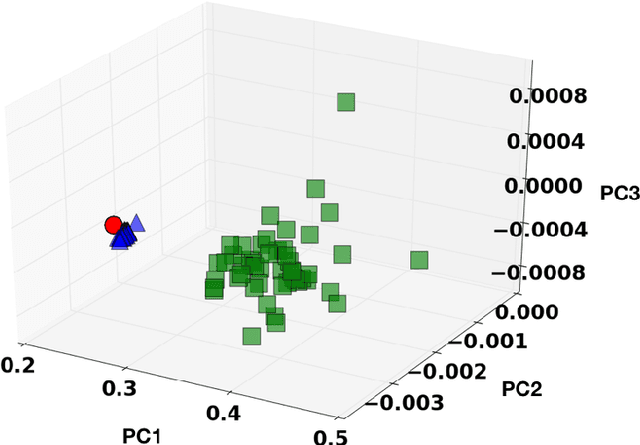
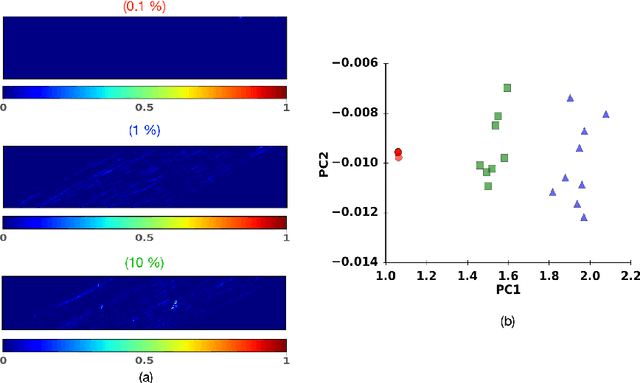
Digital image correlation (DIC) is a well-established, non-invasive technique for tracking and quantifying the deformation of mechanical samples under strain. While it provides an obvious way to observe incremental and aggregate displacement information, it seems likely that DIC data sets, which after all reflect the spatially-resolved response of a microstructure to loads, contain much richer information than has generally been extracted from them. In this paper, we demonstrate a machine-learning approach to quantifying the prior deformation history of a crystalline sample based on its response to a subsequent DIC test. This prior deformation history is encoded in the microstructure through the inhomogeneity of the dislocation microstructure, and in the spatial correlations of the dislocation patterns, which mediate the system's response to the DIC test load. Our domain consists of deformed crystalline thin films generated by a discrete dislocation plasticity simulation. We explore the range of applicability of machine learning (ML) for typical experimental protocols, and as a function of possible size effects and stochasticity. Plasticity size effects may directly influence the data, rendering unsupervised techniques unable to distinguish different plasticity regimes.
Discriminative Nonlinear Analysis Operator Learning: When Cosparse Model Meets Image Classification
Apr 30, 2017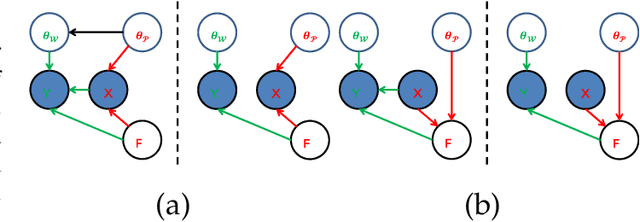
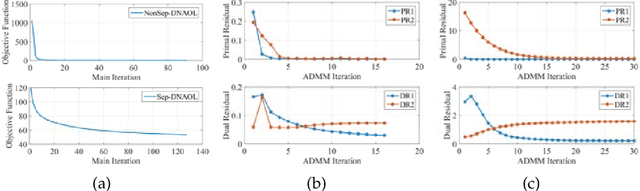
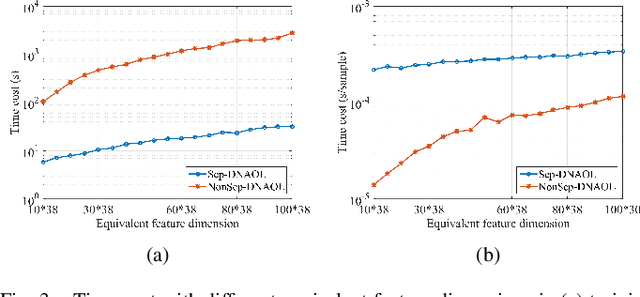
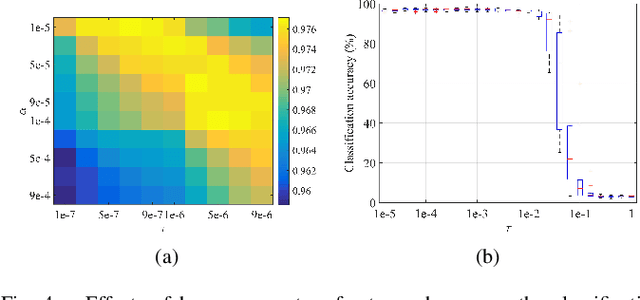
Linear synthesis model based dictionary learning framework has achieved remarkable performances in image classification in the last decade. Behaved as a generative feature model, it however suffers from some intrinsic deficiencies. In this paper, we propose a novel parametric nonlinear analysis cosparse model (NACM) with which a unique feature vector will be much more efficiently extracted. Additionally, we derive a deep insight to demonstrate that NACM is capable of simultaneously learning the task adapted feature transformation and regularization to encode our preferences, domain prior knowledge and task oriented supervised information into the features. The proposed NACM is devoted to the classification task as a discriminative feature model and yield a novel discriminative nonlinear analysis operator learning framework (DNAOL). The theoretical analysis and experimental performances clearly demonstrate that DNAOL will not only achieve the better or at least competitive classification accuracies than the state-of-the-art algorithms but it can also dramatically reduce the time complexities in both training and testing phases.
Increasing Shape Bias in ImageNet-Trained Networks Using Transfer Learning and Domain-Adversarial Methods
Jul 30, 2019
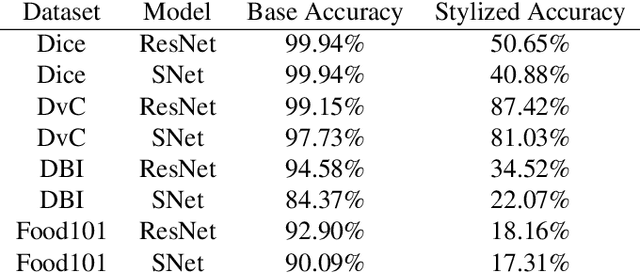

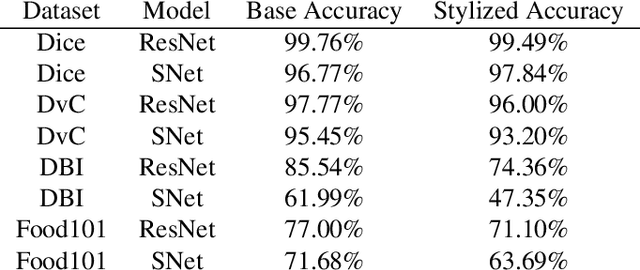
Convolutional Neural Networks (CNNs) have become the state-of-the-art method to learn from image data. However, recent research shows that they may include a texture and colour bias in their representation, contrary to the intuition that they learn the shapes of the image content and to human biological learning. Thus, recent works have attempted to increase the shape bias in CNNs in order to train more robust and accurate networks on tasks. One such approach uses style-transfer in order to remove texture clues from the data. This work reproduces this methodology on four image classification datasets, as well as extends the method to use domain-adversarial training in order to further increase the shape bias in the learned representation. The results show the proposed method increases the robustness and shape bias of the CNNs, while it does not provide a gain in accuracy.
GIFT: Learning Transformation-Invariant Dense Visual Descriptors via Group CNNs
Nov 14, 2019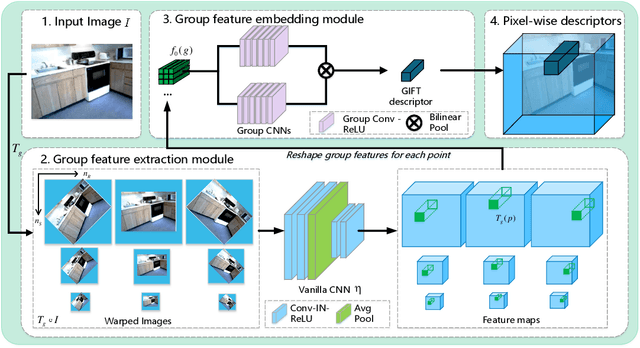
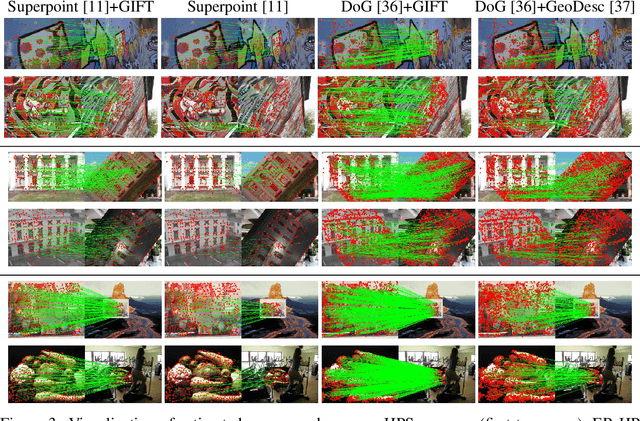
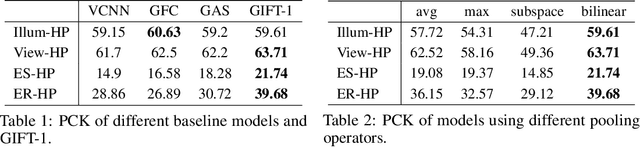
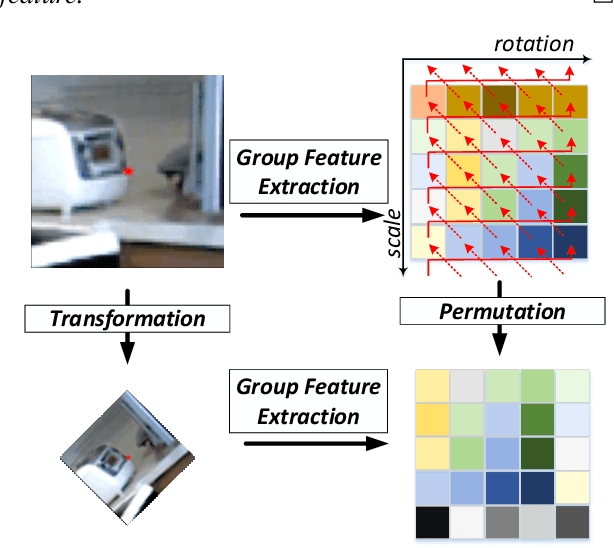

Finding local correspondences between images with different viewpoints requires local descriptors that are robust against geometric transformations. An approach for transformation invariance is to integrate out the transformations by pooling the features extracted from transformed versions of an image. However, the feature pooling may sacrifice the distinctiveness of the resulting descriptors. In this paper, we introduce a novel visual descriptor named Group Invariant Feature Transform (GIFT), which is both discriminative and robust to geometric transformations. The key idea is that the features extracted from the transformed versions of an image can be viewed as a function defined on the group of the transformations. Instead of feature pooling, we use group convolutions to exploit underlying structures of the extracted features on the group, resulting in descriptors that are both discriminative and provably invariant to the group of transformations. Extensive experiments show that GIFT outperforms state-of-the-art methods on several benchmark datasets and practically improves the performance of relative pose estimation.
Virtual organelle self-coding for fluorescence imaging via adversarial learning
Sep 10, 2019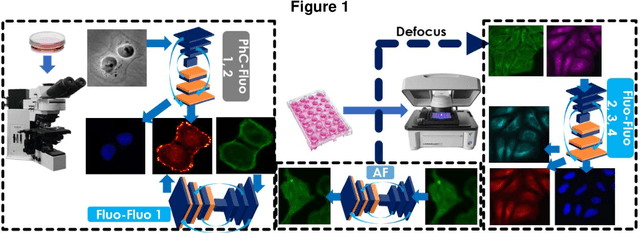
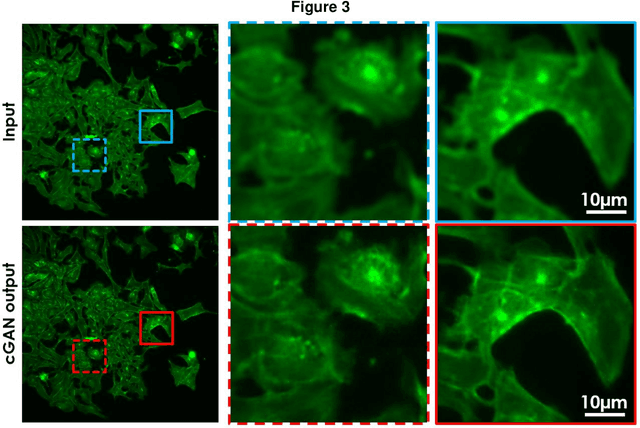
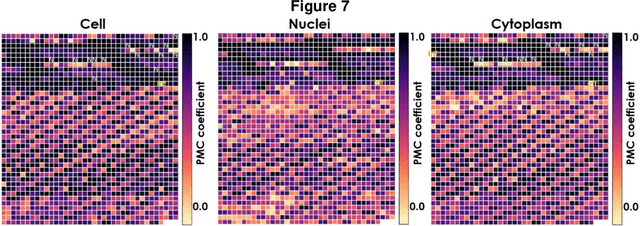
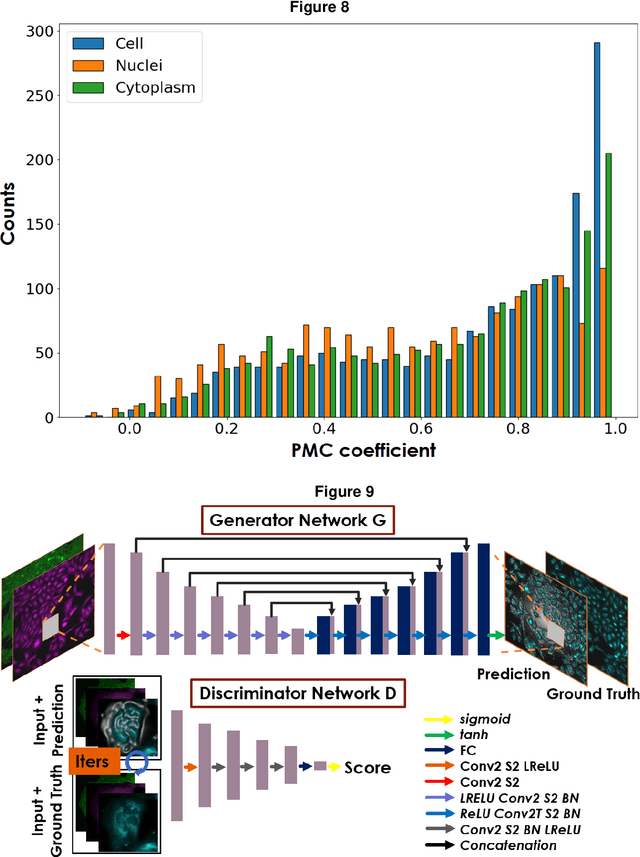
Fluorescence microscopy plays a vital role in understanding the subcellular structures of living cells. However, it requires considerable effort in sample preparation related to chemical fixation, staining, cost, and time. To reduce those factors, we present a virtual fluorescence staining method based on deep neural networks (VirFluoNet) to transform fluorescence images of molecular labels into other molecular fluorescence labels in the same field-of-view. To achieve this goal, we develop and train a conditional generative adversarial network (cGAN) to perform digital fluorescence imaging demonstrated on human osteosarcoma U2OS cell fluorescence images captured under Cell Painting staining protocol. A detailed comparative analysis is also conducted on the performance of the cGAN network between predicting fluorescence channels based on phase contrast or based on another fluorescence channel using human breast cancer MDA-MB-231 cell line as a test case. In addition, we implement a deep learning model to perform autofocusing on another human U2OS fluorescence dataset as a preprocessing step to defocus an out-focus channel in U2OS dataset. A quantitative index of image prediction error is introduced based on signal pixel-wise spatial and intensity differences with ground truth to evaluate the performance of prediction to high-complex and throughput fluorescence. This index provides a rational way to perform image segmentation on error signals and to understand the likelihood of mis-interpreting biology from the predicted image. In total, these findings contribute to the utility of deep learning image regression for fluorescence microscopy datasets of biological cells, balanced against savings of cost, time, and experimental effort. Furthermore, the approach introduced here holds promise for modeling the internal relationships between organelles and biomolecules within living cells.
Superpixel-Based Background Recovery from Multiple Images
Nov 04, 2019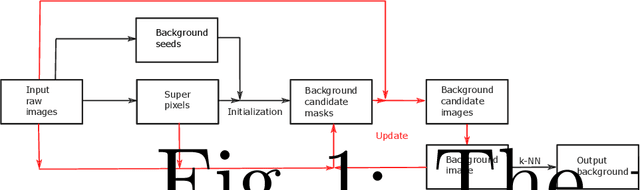


In this paper, we propose an intuitive method to recover background from multiple images. The implementation consists of three stages: model initialization, model update, and background output. We consider the pixels whose values change little in all input images as background seeds. Images are then segmented into superpixels with simple linear iterative clustering. When the number of pixels labelled as background in a superpixel is bigger than a predefined threshold, we label the superpixel as background to initialize the background candidate masks. Background candidate images are obtained from input raw images with the masks. Combining all candidate images, a background image is produced. The background candidate masks, candidate images, and the background image are then updated alternately until convergence. Finally, ghosting artifacts is removed with the k-nearest neighbour method. An experiment on an outdoor dataset demonstrates that the proposed algorithm can achieve promising results.
NeurIPS 2019 Disentanglement Challenge: Improved Disentanglement through Learned Aggregation of Convolutional Feature Maps
Feb 27, 2020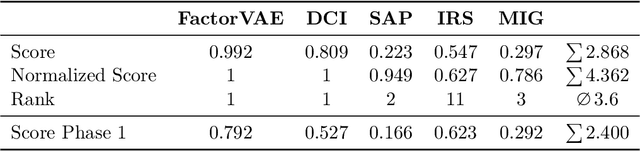
This report to our stage 2 submission to the NeurIPS 2019 disentanglement challenge presents a simple image preprocessing method for learning disentangled latent factors. We propose to train a variational autoencoder on regionally aggregated feature maps obtained from networks pretrained on the ImageNet database, utilizing the implicit inductive bias contained in those features for disentanglement. This bias can be further enhanced by explicitly fine-tuning the feature maps on auxiliary tasks useful for the challenge, such as angle, position estimation, or color classification. Our approach achieved the 2nd place in stage 2 of the challenge. Code is available at https://github.com/mseitzer/neurips2019-disentanglement-challenge.
Natural Color Image Enhancement based on Modified Multiscale Retinex Algorithm and Performance Evaluation usingWavelet Energy
Jun 22, 2014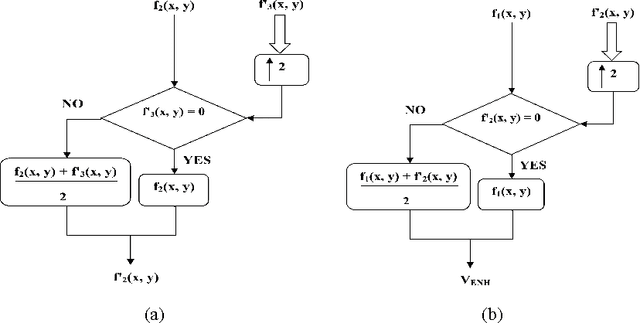
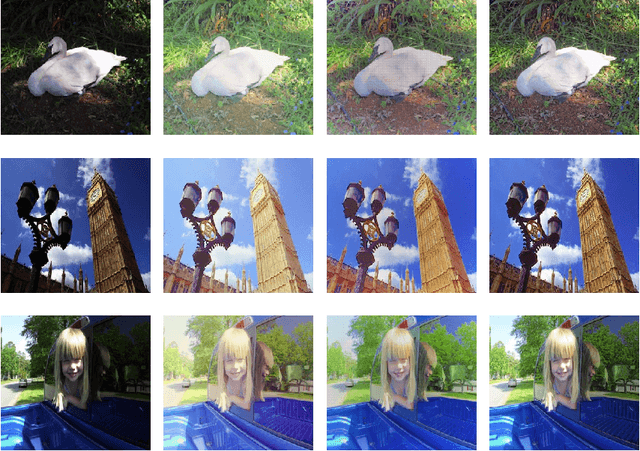
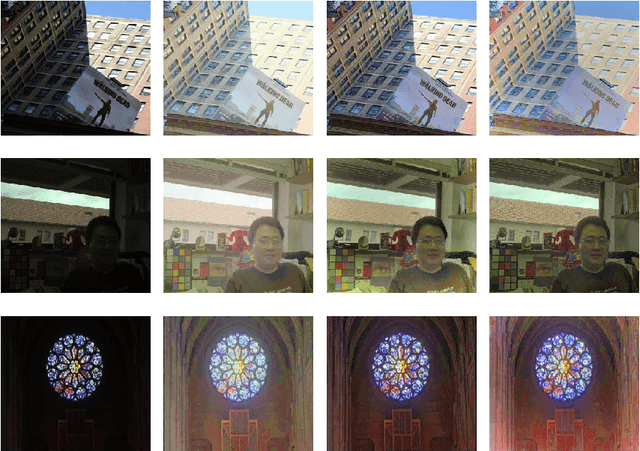
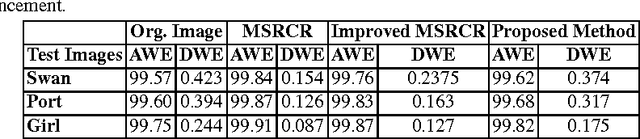


This paper presents a new color image enhancement technique based on modified MultiScale Retinex(MSR) algorithm and visual quality of the enhanced images are evaluated using a new metric, namely, wavelet energy. The color image enhancement is achieved by down sampling the value component of HSV color space converted image into three scales (normal, medium and fine) following the contrast stretching operation. These down sampled value components are enhanced using the MSR algorithm. The value component is reconstructed by averaging each pixels of the lower scale image with that of the upper scale image subsequent to up sampling the lower scale image. This process replaces dark pixel by the average pixels of both the lower scale and upper scale, while retaining the bright pixels. The quality of the reconstructed images in the proposed method is found to be good and far better then the other researchers method. The performance of the proposed scheme is evaluated using new wavelet domain based assessment criterion, referred as wavelet energy. This scheme computes the energy of both original and enhanced image in wavelet domain. The number of edge details as well as wavelet energy is less in a poor quality image compared with naturally enhanced image. Experimental results presented confirms that the proposed wavelet energy based color image quality assessment technique efficiently characterizes both the local and global details of enhanced image.
Adaptive Weighting Multi-Field-of-View CNN for Semantic Segmentation in Pathology
Apr 12, 2019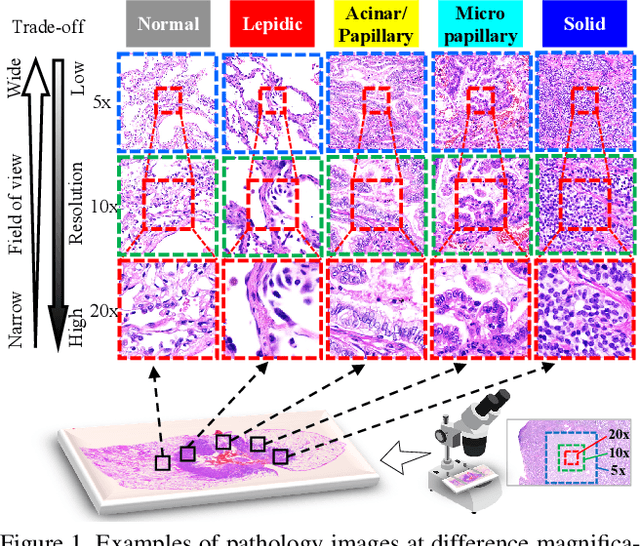
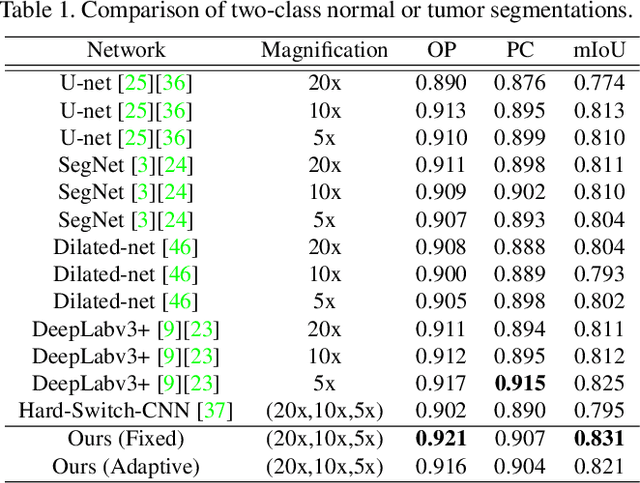
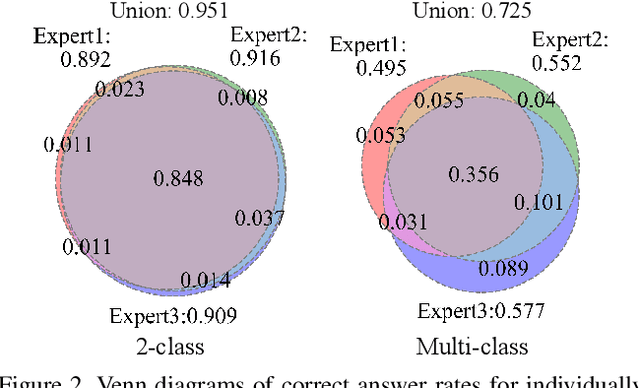
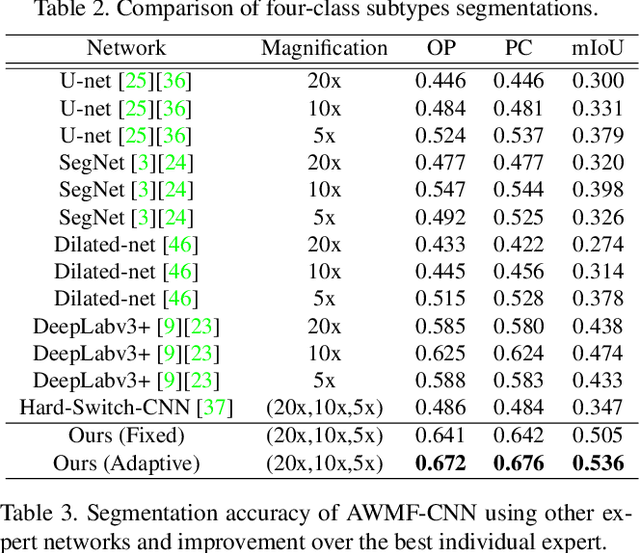
Automated digital histopathology image segmentation is an important task to help pathologists diagnose tumors and cancer subtypes. For pathological diagnosis of cancer subtypes, pathologists usually change the magnification of whole-slide images (WSI) viewers. A key assumption is that the importance of the magnifications depends on the characteristics of the input image, such as cancer subtypes. In this paper, we propose a novel semantic segmentation method, called Adaptive-Weighting-Multi-Field-of-View-CNN (AWMF-CNN), that can adaptively use image features from images with different magnifications to segment multiple cancer subtype regions in the input image. The proposed method aggregates several expert CNNs for images of different magnifications by adaptively changing the weight of each expert depending on the input image. It leverages information in the images with different magnifications that might be useful for identifying the subtypes. It outperformed other state-of-the-art methods in experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge