"Image": models, code, and papers
A salt and pepper noise image denoising method based on the generative classification
Jul 15, 2018
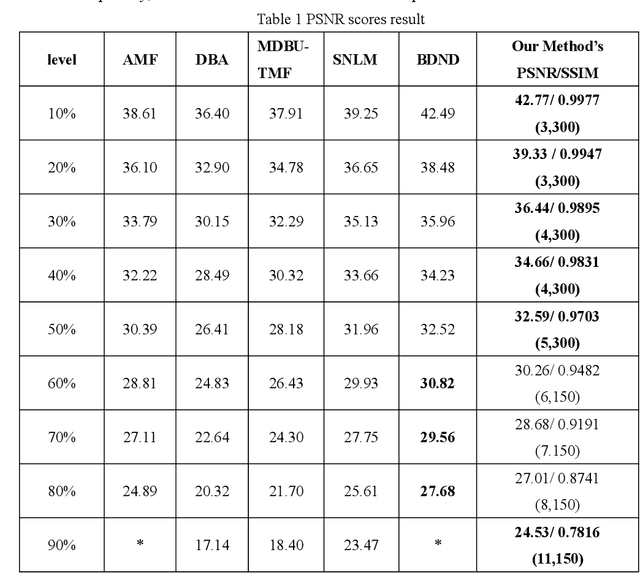
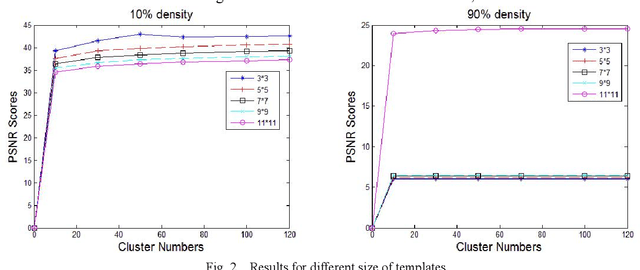
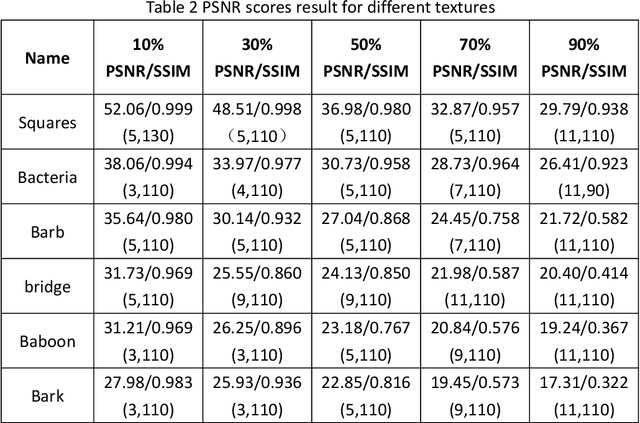
In this paper, an image denoising algorithm is proposed for salt and pepper noise. First, a generative model is built on a patch as a basic unit and then the algorithm locates the image noise within that patch in order to better describe the patch and obtain better subsequent clustering. Second, the algorithm classifies patches using a generative clustering method, thus providing additional similarity information for noise repair and suppressing the interference of noise, abandoning those categories that consist of a smaller number of patches. Finally, the algorithm builds a non-local switching filter to remove the salt and pepper noise. Simulation results show that the proposed algorithm effectively denoises salt and pepper noise of various densities. It obtains a better visual quality and higher peak signal-to-noise ratio score than several state-of-the-art algorithms. In short, our algorithm uses a noisy patch as the basic unit, a patch clustering method to optimize the repair data set as well as obtain a better denoising effect, and provides a guideline for future denoising and repair methods.
Spatiotemporal tomography based on scattered multiangular signals and its application for resolving evolving clouds using moving platforms
Dec 06, 2020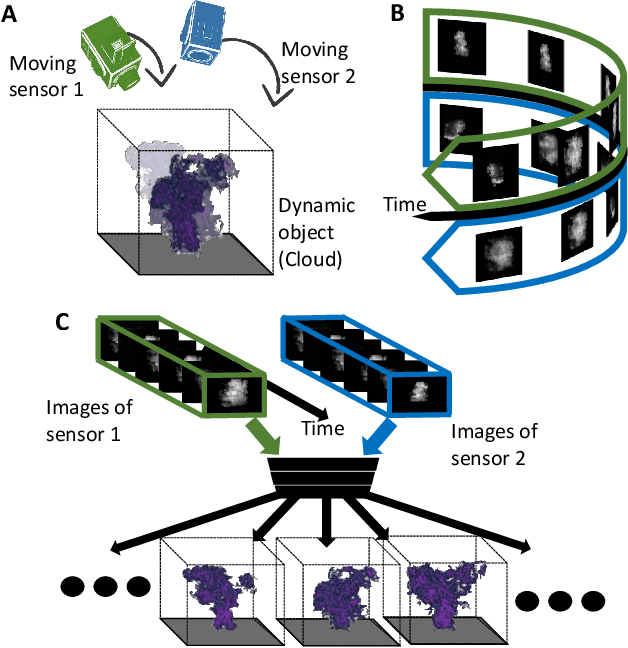
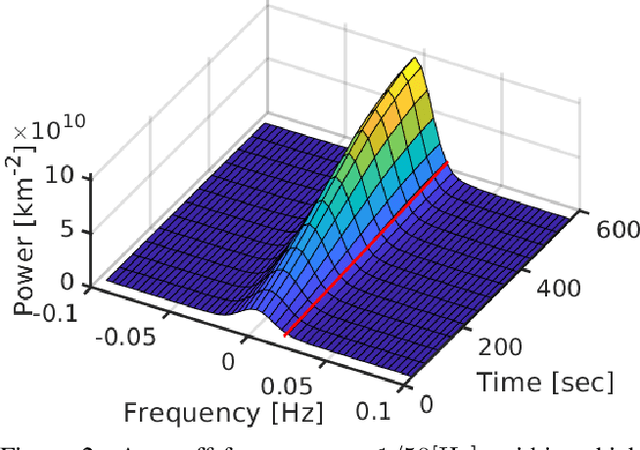
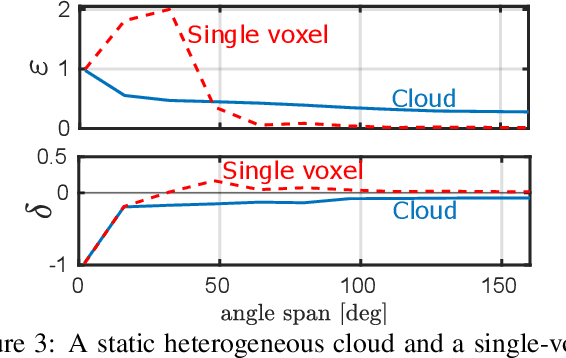
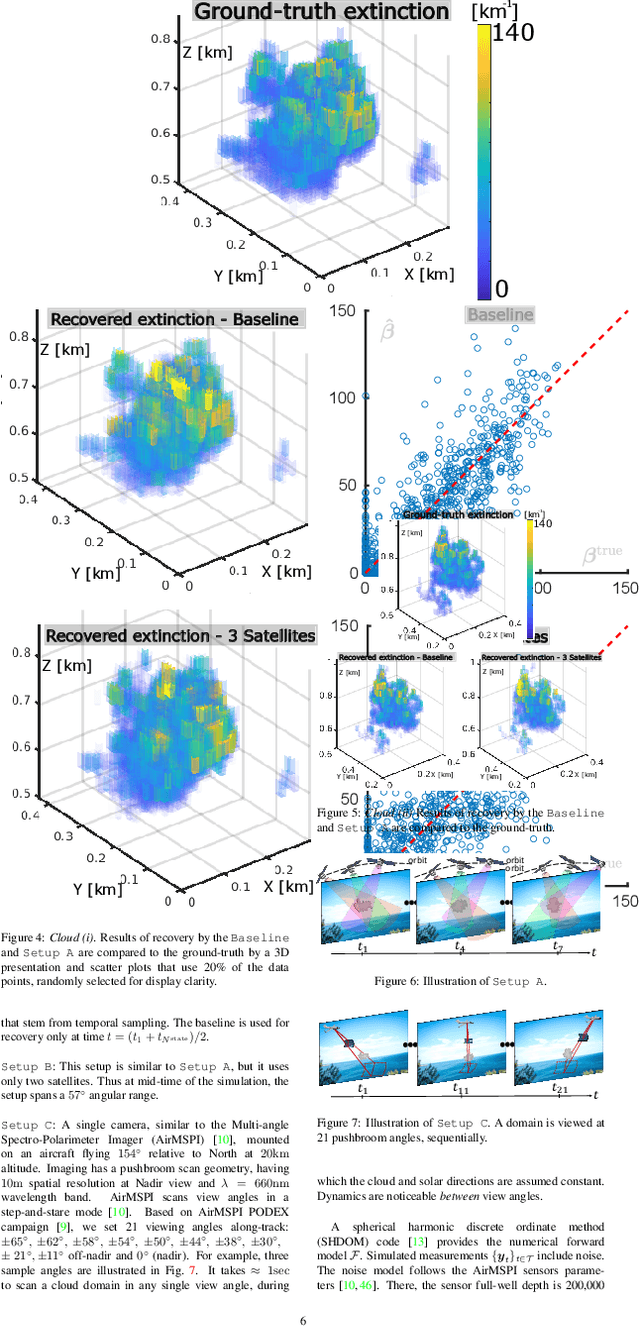
We derive computed tomography (CT) of a time-varying volumetric translucent object, using a small number of moving cameras. We particularly focus on passive scattering tomography, which is a non-linear problem. We demonstrate the approach on dynamic clouds, as clouds have a major effect on Earth's climate. State of the art scattering CT assumes a static object. Existing 4D CT methods rely on a linear image formation model and often on significant priors. In this paper, the angular and temporal sampling rates needed for a proper recovery are discussed. If these rates are used, the paper leads to a representation of the time-varying object, which simplifies 4D CT tomography. The task is achieved using gradient-based optimization. We demonstrate this in physics-based simulations and in an experiment that had yielded real-world data.
Constraint-free Natural Image Reconstruction from fMRI Signals Based on Convolutional Neural Network
Jan 16, 2018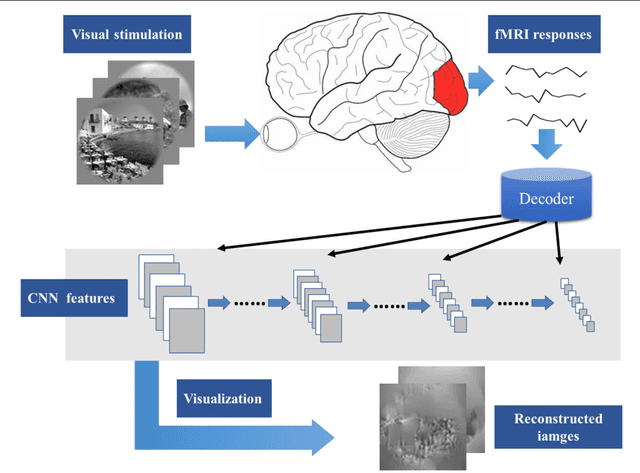
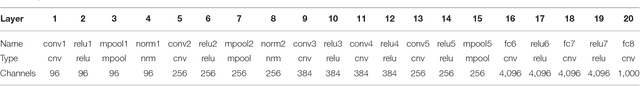
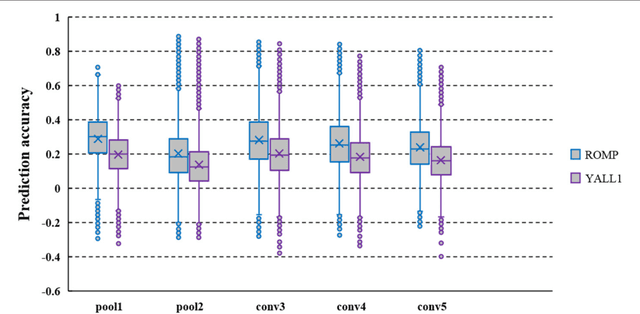
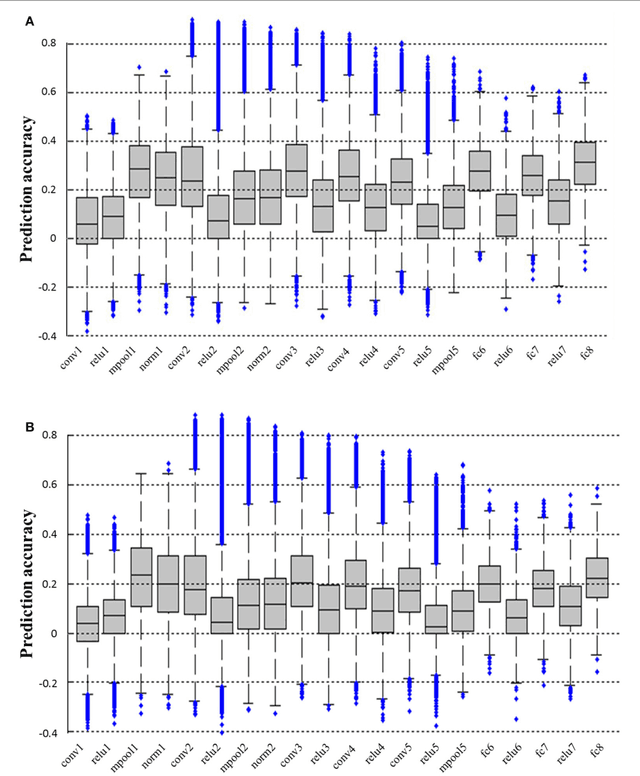
In recent years, research on decoding brain activity based on functional magnetic resonance imaging (fMRI) has made remarkable achievements. However, constraint-free natural image reconstruction from brain activity is still a challenge. The existing methods simplified the problem by using semantic prior information or just reconstructing simple images such as letters and digitals. Without semantic prior information, we present a novel method to reconstruct nature images from fMRI signals of human visual cortex based on the computation model of convolutional neural network (CNN). Firstly, we extracted the units output of viewed natural images in each layer of a pre-trained CNN as CNN features. Secondly, we transformed image reconstruction from fMRI signals into the problem of CNN feature visualizations by training a sparse linear regression to map from the fMRI patterns to CNN features. By iteratively optimization to find the matched image, whose CNN unit features become most similar to those predicted from the brain activity, we finally achieved the promising results for the challenging constraint-free natural image reconstruction. As there was no use of semantic prior information of the stimuli when training decoding model, any category of images (not constraint by the training set) could be reconstructed theoretically. We found that the reconstructed images resembled the natural stimuli, especially in position and shape. The experimental results suggest that hierarchical visual features can effectively express the visual perception process of human brain.
Quantum Superposition Spiking Neural Network
Oct 27, 2020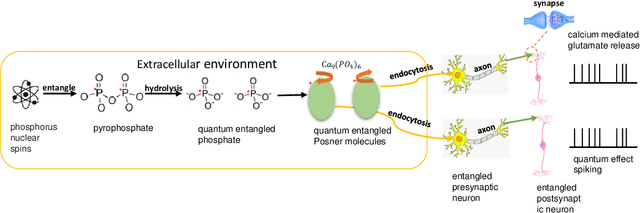
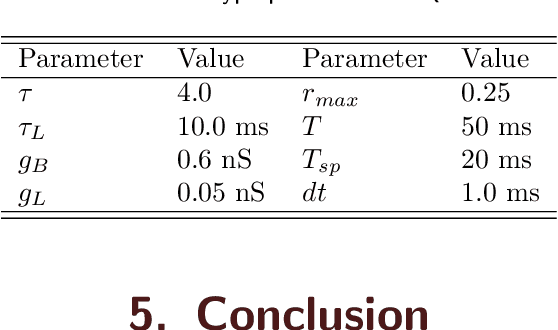

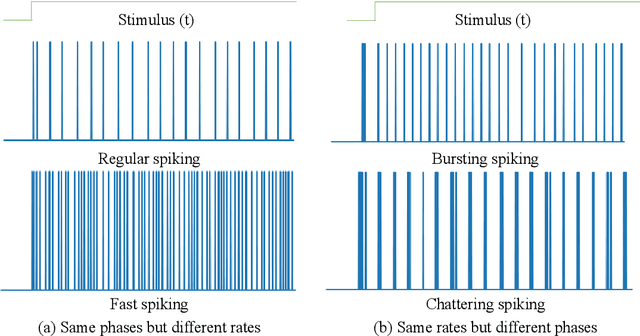
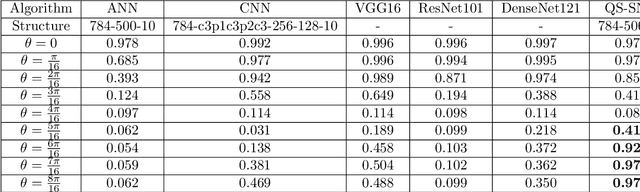
Quantum brain as a novel hypothesis states that some non-trivial mechanisms in quantum computation, such as superposition and entanglement, may have important influence for the formation of brain functions. Inspired by this idea, we propose Quantum Superposition Spiking Neural Network (QS-SNN), which introduce quantum superposition to spiking neural network models to handel challenges which are hard for other state-of-the-art machine learning models. For human brain, grasping the main information no matter how the background changes is necessary to interact efficiently with diverse environments. As an example, it is easy for human to recognize the digits whether it is white character with black background or inversely black character with white background. While if the current machine learning models are trained with one of the cases (e.g. white character with black background), it will be nearly impossible for them to recognize the color inverted version. To handel this challenge, we propose two-compartment spiking neural network with superposition states encoding, which is inspired by quantum information theory and spatial-temporal spiking property from neuron information encoding in the brain. Typical network structures like fully-connected ANN, VGG, ResNet and DenseNet are challenged with the same task. We train these networks on original image dataset and then invert the background color to test their generalization. Result shows that artificial neural network can not deal with this condition while the quantum superposition spiking neural network(QS-SNN) which we proposed in this paper recognizes the color-inverse image successfully. Further the QS-SNN shows its robustness when noises are added on inputs.
An Inductive Transfer Learning Approach using Cycle-consistent Adversarial Domain Adaptation with Application to Brain Tumor Segmentation
May 11, 2020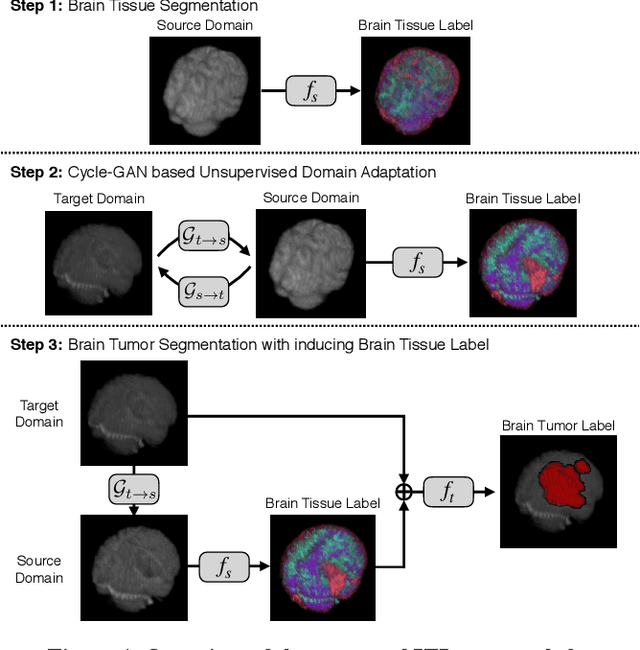
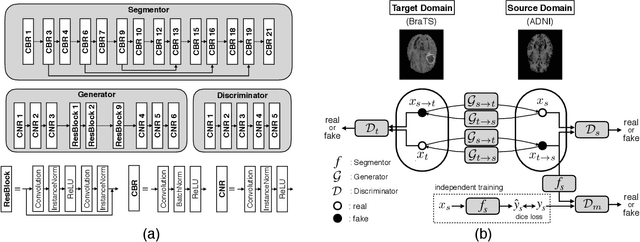
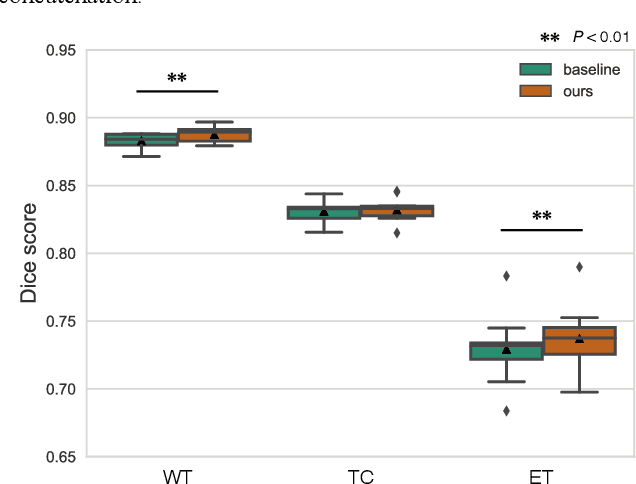
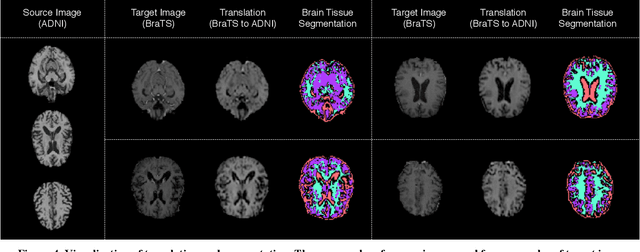
With recent advances in supervised machine learning for medical image analysis applications, the annotated medical image datasets of various domains are being shared extensively. Given that the annotation labelling requires medical expertise, such labels should be applied to as many learning tasks as possible. However, the multi-modal nature of each annotated image renders it difficult to share the annotation label among diverse tasks. In this work, we provide an inductive transfer learning (ITL) approach to adopt the annotation label of the source domain datasets to tasks of the target domain datasets using Cycle-GAN based unsupervised domain adaptation (UDA). To evaluate the applicability of the ITL approach, we adopted the brain tissue annotation label on the source domain dataset of Magnetic Resonance Imaging (MRI) images to the task of brain tumor segmentation on the target domain dataset of MRI. The results confirm that the segmentation accuracy of brain tumor segmentation improved significantly. The proposed ITL approach can make significant contribution to the field of medical image analysis, as we develop a fundamental tool to improve and promote various tasks using medical images.
Stacked Attention Networks for Image Question Answering
Jan 26, 2016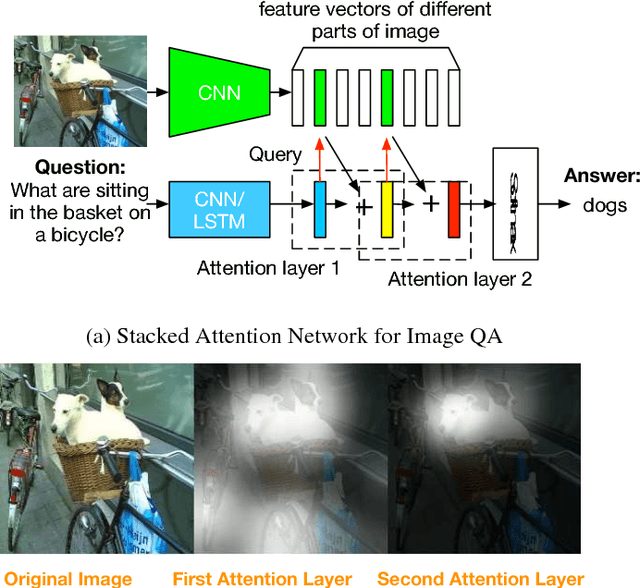
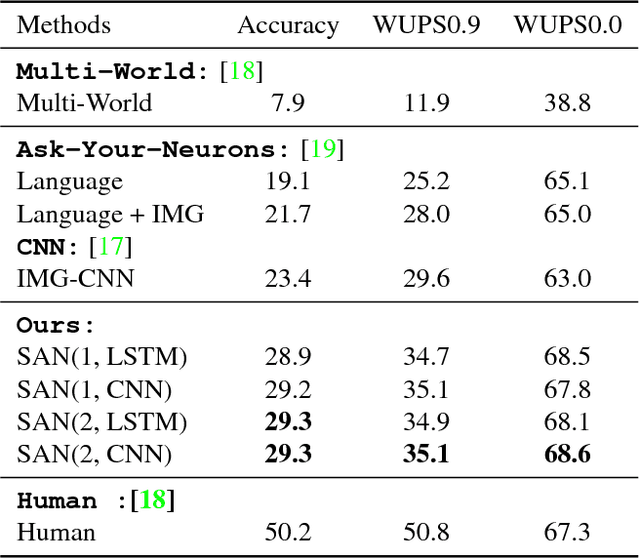
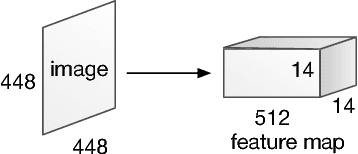
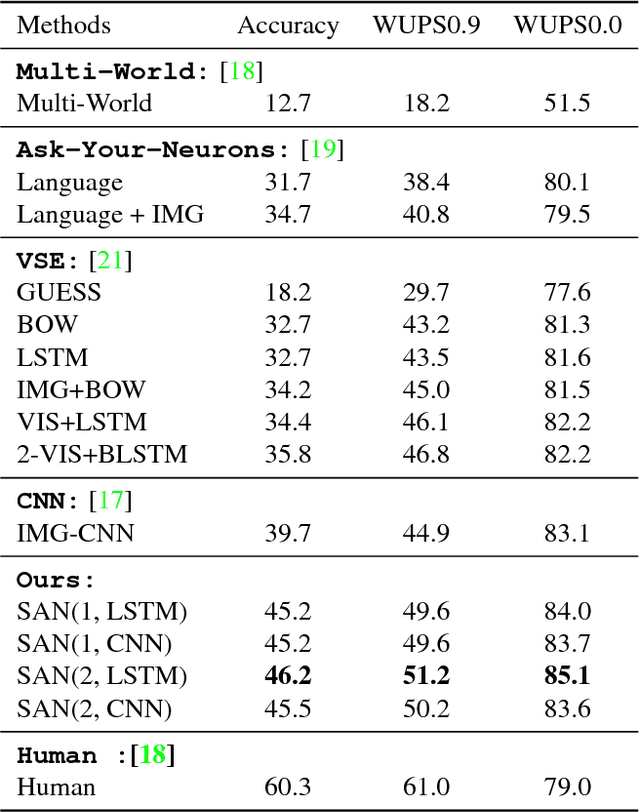
This paper presents stacked attention networks (SANs) that learn to answer natural language questions from images. SANs use semantic representation of a question as query to search for the regions in an image that are related to the answer. We argue that image question answering (QA) often requires multiple steps of reasoning. Thus, we develop a multiple-layer SAN in which we query an image multiple times to infer the answer progressively. Experiments conducted on four image QA data sets demonstrate that the proposed SANs significantly outperform previous state-of-the-art approaches. The visualization of the attention layers illustrates the progress that the SAN locates the relevant visual clues that lead to the answer of the question layer-by-layer.
Target Detection and Segmentation in Circular-Scan Synthetic-Aperture-Sonar Images using Semi-Supervised Convolutional Encoder-Decoders
Jan 10, 2021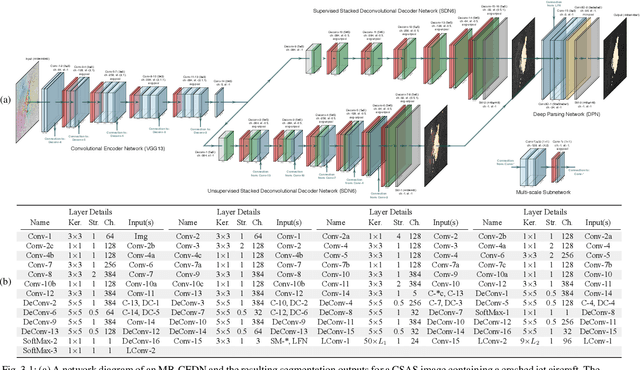
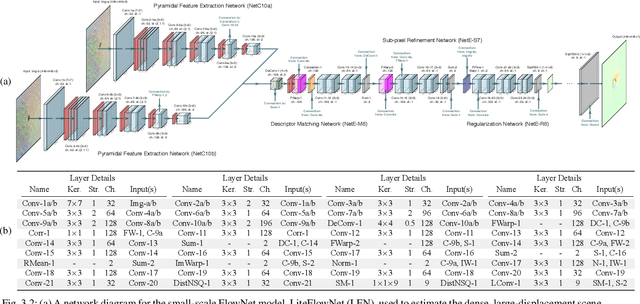
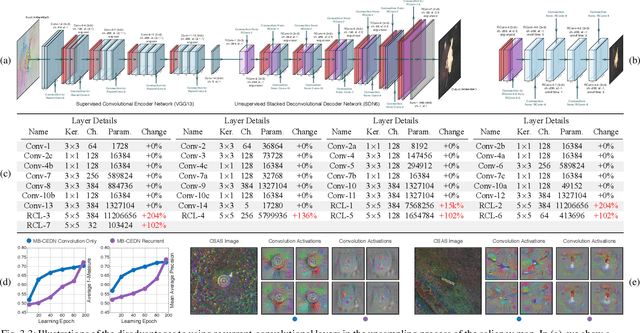
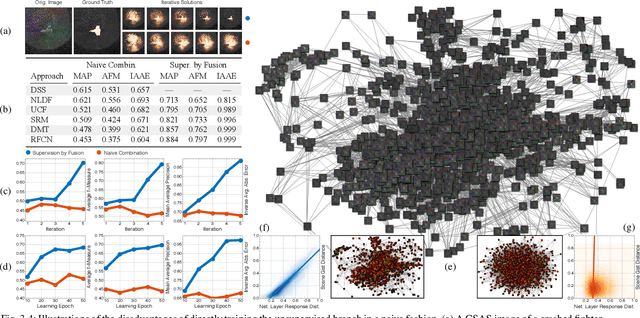
We propose a saliency-based, multi-target detection and segmentation framework for multi-aspect, semi-coherent imagery formed from circular-scan, synthetic-aperture sonar (CSAS). Our framework relies on a multi-branch, convolutional encoder-decoder network (MB-CEDN). The encoder portion extracts features from one or more CSAS images of the targets. These features are then split off and fed into multiple decoders that perform pixel-level classification on the extracted features to roughly mask the target in an unsupervised-trained manner and detect foreground and background pixels in a supervised-trained manner. Each of these target-detection estimates provide different perspectives as to what constitute a target. These opinions are cascaded into a deep-parsing network to model contextual and spatial constraints that help isolate targets better than either solution estimate alone. We evaluate our framework using real-world CSAS data with five broad target classes. Since we are the first to consider both CSAS target detection and segmentation, we adapt existing image and video-processing network topologies from the literature for comparative purposes. We show that our framework outperforms supervised deep networks. It greatly outperforms state-of-the-art unsupervised approaches for diverse target and seafloor types.
An image representation based convolutional network for DNA classification
Jun 13, 2018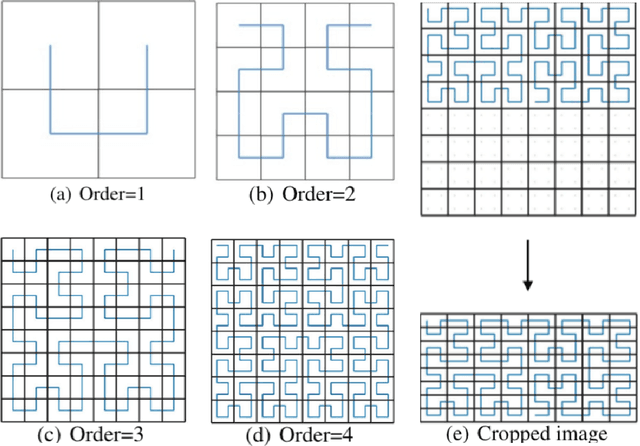
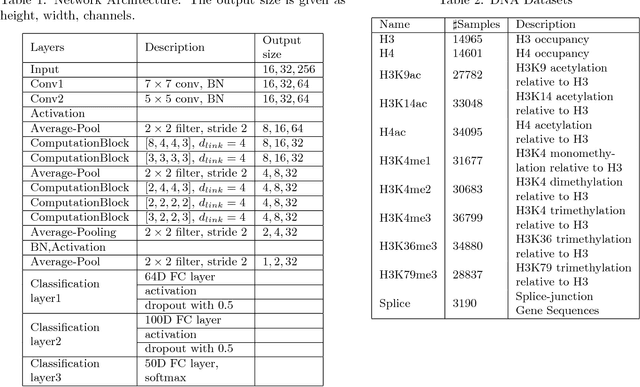
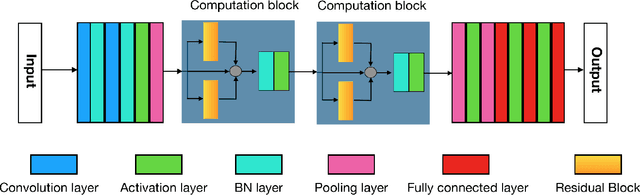
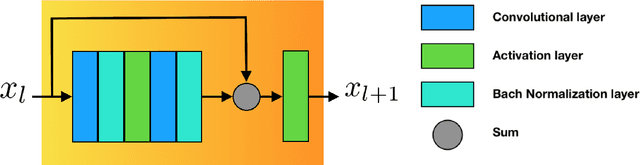
The folding structure of the DNA molecule combined with helper molecules, also referred to as the chromatin, is highly relevant for the functional properties of DNA. The chromatin structure is largely determined by the underlying primary DNA sequence, though the interaction is not yet fully understood. In this paper we develop a convolutional neural network that takes an image-representation of primary DNA sequence as its input, and predicts key determinants of chromatin structure. The method is developed such that it is capable of detecting interactions between distal elements in the DNA sequence, which are known to be highly relevant. Our experiments show that the method outperforms several existing methods both in terms of prediction accuracy and training time.
Graph Laplacian Regularization for Image Denoising: Analysis in the Continuous Domain
Aug 30, 2017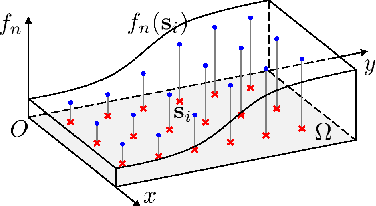
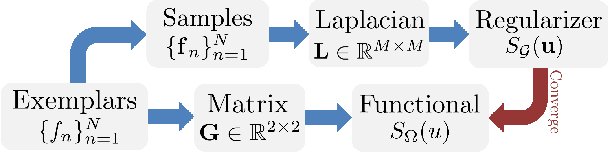
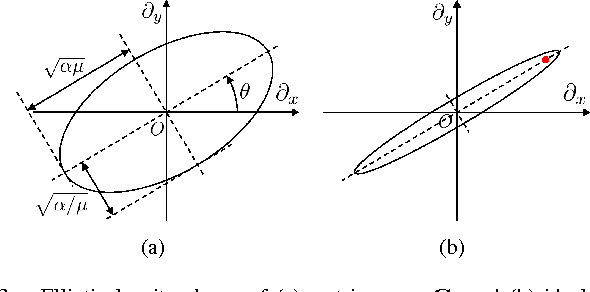
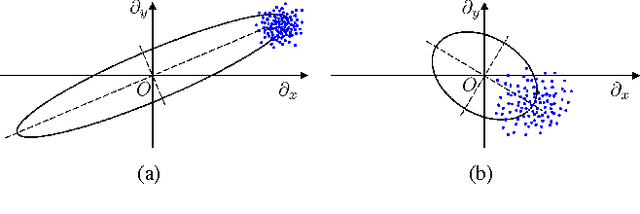
Inverse imaging problems are inherently under-determined, and hence it is important to employ appropriate image priors for regularization. One recent popular prior---the graph Laplacian regularizer---assumes that the target pixel patch is smooth with respect to an appropriately chosen graph. However, the mechanisms and implications of imposing the graph Laplacian regularizer on the original inverse problem are not well understood. To address this problem, in this paper we interpret neighborhood graphs of pixel patches as discrete counterparts of Riemannian manifolds and perform analysis in the continuous domain, providing insights into several fundamental aspects of graph Laplacian regularization for image denoising. Specifically, we first show the convergence of the graph Laplacian regularizer to a continuous-domain functional, integrating a norm measured in a locally adaptive metric space. Focusing on image denoising, we derive an optimal metric space assuming non-local self-similarity of pixel patches, leading to an optimal graph Laplacian regularizer for denoising in the discrete domain. We then interpret graph Laplacian regularization as an anisotropic diffusion scheme to explain its behavior during iterations, e.g., its tendency to promote piecewise smooth signals under certain settings. To verify our analysis, an iterative image denoising algorithm is developed. Experimental results show that our algorithm performs competitively with state-of-the-art denoising methods such as BM3D for natural images, and outperforms them significantly for piecewise smooth images.
An Interactive Medical Image Segmentation Framework Using Iterative Refinement
Jun 05, 2016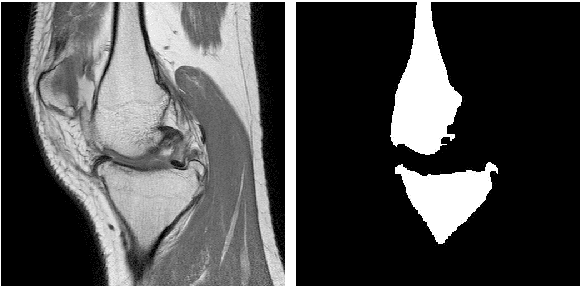
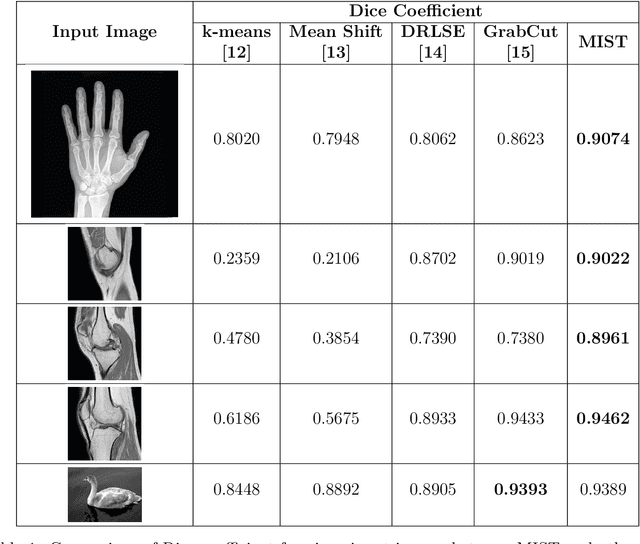
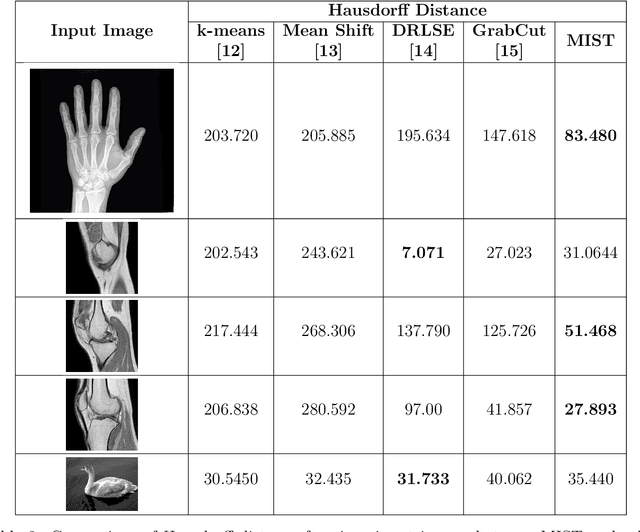
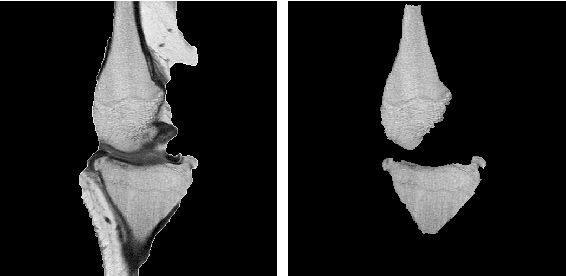
Image segmentation is often performed on medical images for identifying diseases in clinical evaluation. Hence it has become one of the major research areas. Conventional image segmentation techniques are unable to provide satisfactory segmentation results for medical images as they contain irregularities. They need to be pre-processed before segmentation. In order to obtain the most suitable method for medical image segmentation, we propose a two stage algorithm. The first stage automatically generates a binary marker image of the region of interest using mathematical morphology. This marker serves as the mask image for the second stage which uses GrabCut on the input image thus resulting in an efficient segmented result. The obtained result can be further refined by user interaction which can be done using the Graphical User Interface (GUI). Experimental results show that the proposed method is accurate and provides satisfactory segmentation results with minimum user interaction on medical as well as natural images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge