"Image": models, code, and papers
Addressing Fairness and Explainability in Image Classification Using Optimal Transport
Aug 22, 2023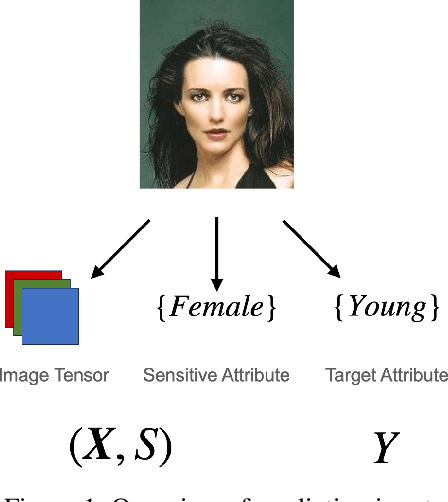
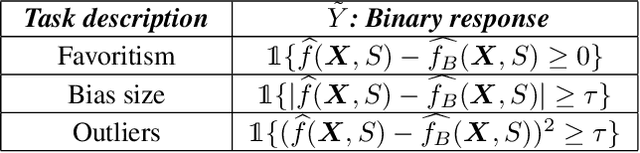
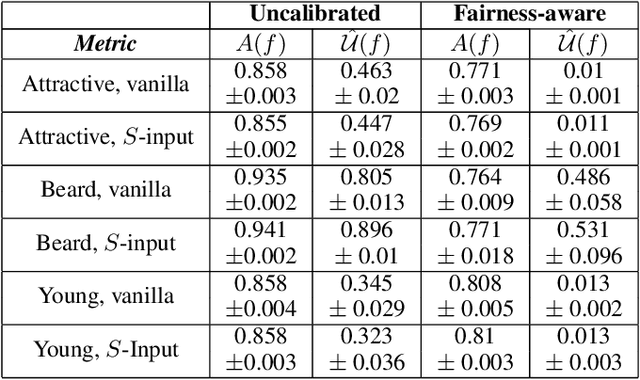
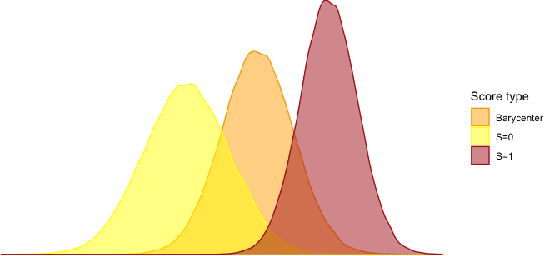

Algorithmic Fairness and the explainability of potentially unfair outcomes are crucial for establishing trust and accountability of Artificial Intelligence systems in domains such as healthcare and policing. Though significant advances have been made in each of the fields separately, achieving explainability in fairness applications remains challenging, particularly so in domains where deep neural networks are used. At the same time, ethical data-mining has become ever more relevant, as it has been shown countless times that fairness-unaware algorithms result in biased outcomes. Current approaches focus on mitigating biases in the outcomes of the model, but few attempts have been made to try to explain \emph{why} a model is biased. To bridge this gap, we propose a comprehensive approach that leverages optimal transport theory to uncover the causes and implications of biased regions in images, which easily extends to tabular data as well. Through the use of Wasserstein barycenters, we obtain scores that are independent of a sensitive variable but keep their marginal orderings. This step ensures predictive accuracy but also helps us to recover the regions most associated with the generation of the biases. Our findings hold significant implications for the development of trustworthy and unbiased AI systems, fostering transparency, accountability, and fairness in critical decision-making scenarios across diverse domains.
Generalized Cross-domain Multi-label Few-shot Learning for Chest X-rays
Sep 08, 2023Real-world application of chest X-ray abnormality classification requires dealing with several challenges: (i) limited training data; (ii) training and evaluation sets that are derived from different domains; and (iii) classes that appear during training may have partial overlap with classes of interest during evaluation. To address these challenges, we present an integrated framework called Generalized Cross-Domain Multi-Label Few-Shot Learning (GenCDML-FSL). The framework supports overlap in classes during training and evaluation, cross-domain transfer, adopts meta-learning to learn using few training samples, and assumes each chest X-ray image is either normal or associated with one or more abnormalities. Furthermore, we propose Generalized Episodic Training (GenET), a training strategy that equips models to operate with multiple challenges observed in the GenCDML-FSL scenario. Comparisons with well-established methods such as transfer learning, hybrid transfer learning, and multi-label meta-learning on multiple datasets show the superiority of our approach.
Active Learning for Classifying 2D Grid-Based Level Completability
Sep 08, 2023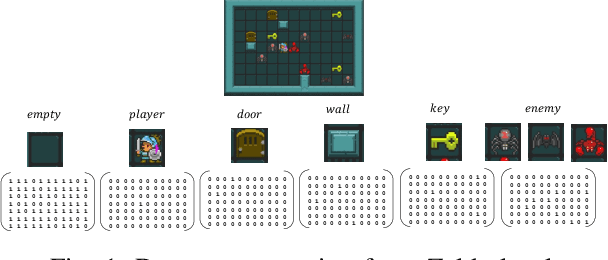
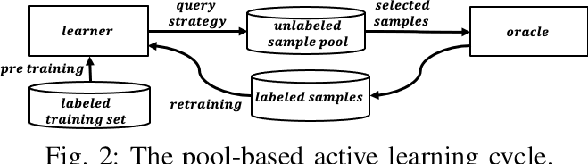
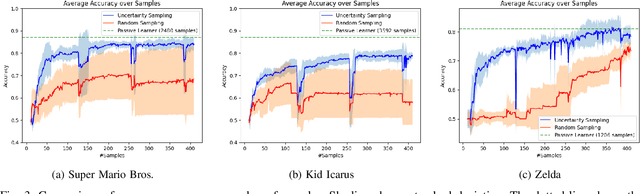
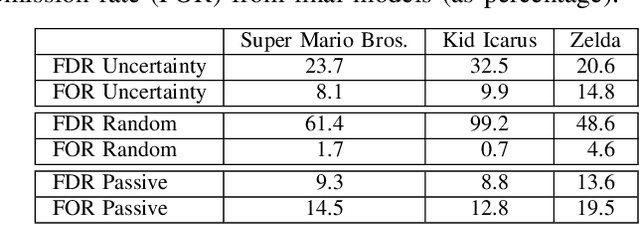
Determining the completability of levels generated by procedural generators such as machine learning models can be challenging, as it can involve the use of solver agents that often require a significant amount of time to analyze and solve levels. Active learning is not yet widely adopted in game evaluations, although it has been used successfully in natural language processing, image and speech recognition, and computer vision, where the availability of labeled data is limited or expensive. In this paper, we propose the use of active learning for learning level completability classification. Through an active learning approach, we train deep-learning models to classify the completability of generated levels for Super Mario Bros., Kid Icarus, and a Zelda-like game. We compare active learning for querying levels to label with completability against random queries. Our results show using an active learning approach to label levels results in better classifier performance with the same amount of labeled data.
* 4 pages, 3 figures
SegmentAnything helps microscopy images based automatic and quantitative organoid detection and analysis
Sep 08, 2023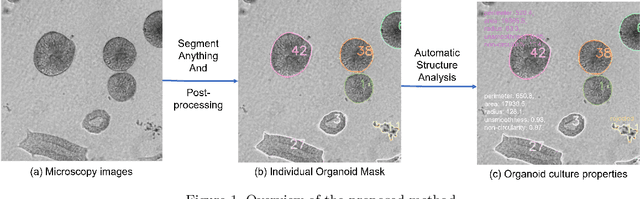
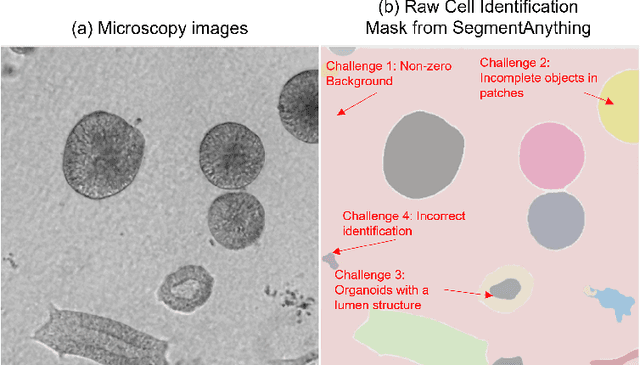
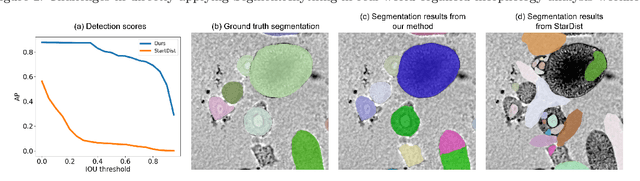
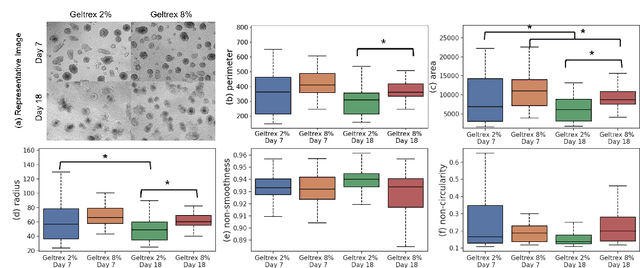
Organoids are self-organized 3D cell clusters that closely mimic the architecture and function of in vivo tissues and organs. Quantification of organoid morphology helps in studying organ development, drug discovery, and toxicity assessment. Recent microscopy techniques provide a potent tool to acquire organoid morphology features, but manual image analysis remains a labor and time-intensive process. Thus, this paper proposes a comprehensive pipeline for microscopy analysis that leverages the SegmentAnything to precisely demarcate individual organoids. Additionally, we introduce a set of morphological properties, including perimeter, area, radius, non-smoothness, and non-circularity, allowing researchers to analyze the organoid structures quantitatively and automatically. To validate the effectiveness of our approach, we conducted tests on bright-field images of human induced pluripotent stem cells (iPSCs) derived neural-epithelial (NE) organoids. The results obtained from our automatic pipeline closely align with manual organoid detection and measurement, showcasing the capability of our proposed method in accelerating organoids morphology analysis.
AdaPlus: Integrating Nesterov Momentum and Precise Stepsize Adjustment on AdamW Basis
Sep 05, 2023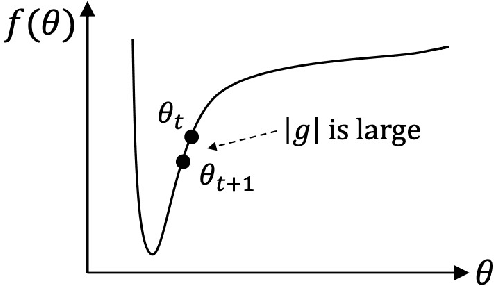
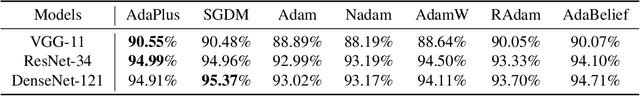
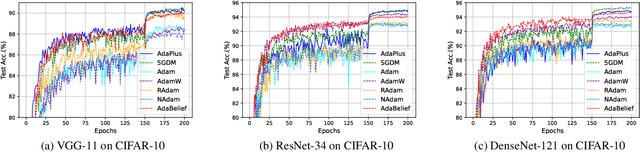
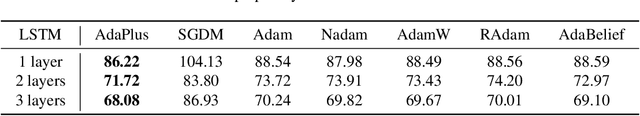
This paper proposes an efficient optimizer called AdaPlus which integrates Nesterov momentum and precise stepsize adjustment on AdamW basis. AdaPlus combines the advantages of AdamW, Nadam, and AdaBelief and, in particular, does not introduce any extra hyper-parameters. We perform extensive experimental evaluations on three machine learning tasks to validate the effectiveness of AdaPlus. The experiment results validate that AdaPlus (i) is the best adaptive method which performs most comparable with (even slightly better than) SGD with momentum on image classification tasks and (ii) outperforms other state-of-the-art optimizers on language modeling tasks and illustrates the highest stability when training GANs. The experiment code of AdaPlus is available at: https://github.com/guanleics/AdaPlus.
EfficientDreamer: High-Fidelity and Robust 3D Creation via Orthogonal-view Diffusion Prior
Aug 25, 2023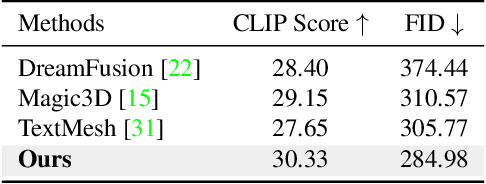
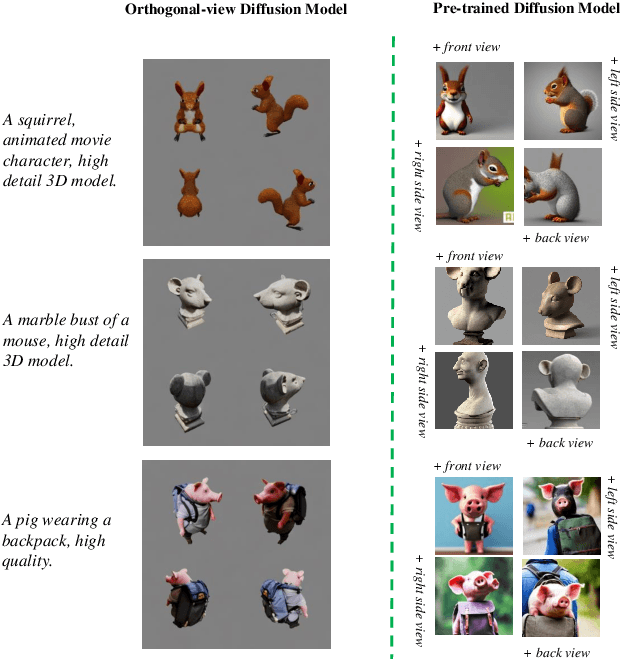
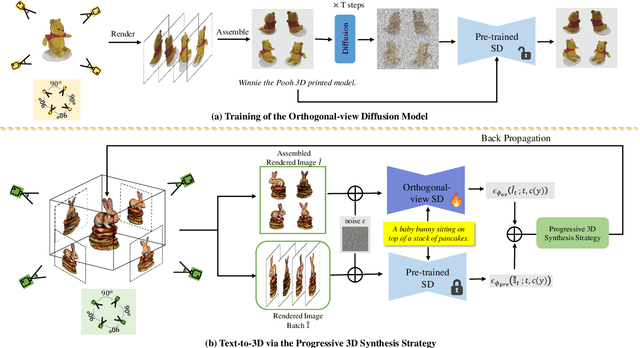
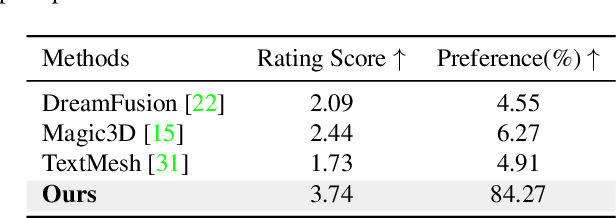

While the image diffusion model has made significant strides in text-driven 3D content creation, it often falls short in accurately capturing the intended meaning of the text prompt, particularly with respect to direction information. This shortcoming gives rise to the Janus problem, where multi-faced 3D models are produced with the guidance of such diffusion models. In this paper, we present a robust pipeline for generating high-fidelity 3D content with orthogonal-view image guidance. Specifically, we introduce a novel 2D diffusion model that generates an image consisting of four orthogonal-view sub-images for the given text prompt. The 3D content is then created with this diffusion model, which enhances 3D consistency and provides strong structured semantic priors. This addresses the infamous Janus problem and significantly promotes generation efficiency. Additionally, we employ a progressive 3D synthesis strategy that results in substantial improvement in the quality of the created 3D contents. Both quantitative and qualitative evaluations show that our method demonstrates a significant improvement over previous text-to-3D techniques.
Mirror-Aware Neural Humans
Sep 09, 2023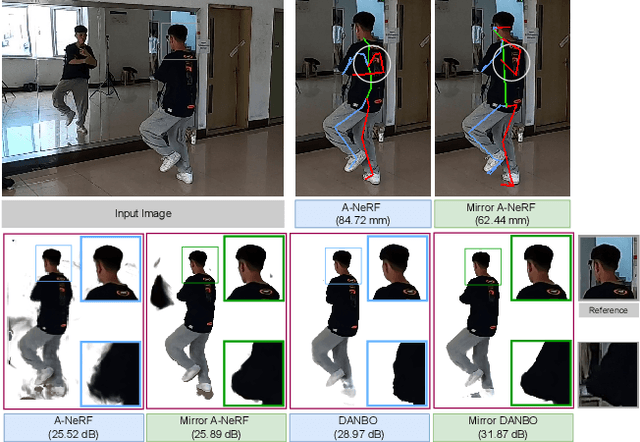
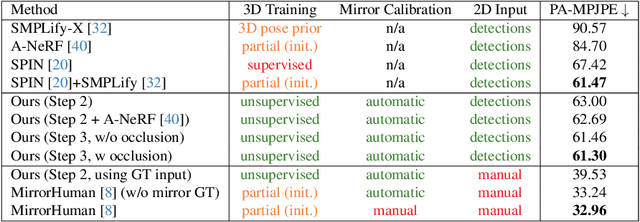
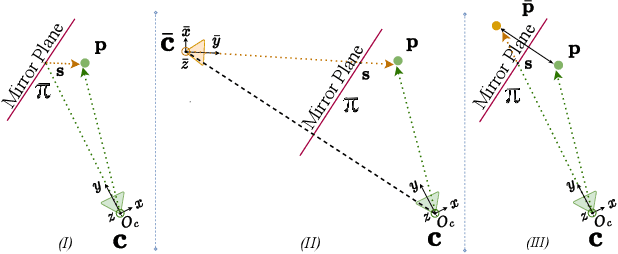

Human motion capture either requires multi-camera systems or is unreliable using single-view input due to depth ambiguities. Meanwhile, mirrors are readily available in urban environments and form an affordable alternative by recording two views with only a single camera. However, the mirror setting poses the additional challenge of handling occlusions of real and mirror image. Going beyond existing mirror approaches for 3D human pose estimation, we utilize mirrors for learning a complete body model, including shape and dense appearance. Our main contributions are extending articulated neural radiance fields to include a notion of a mirror, making it sample-efficient over potential occlusion regions. Together, our contributions realize a consumer-level 3D motion capture system that starts from off-the-shelf 2D poses by automatically calibrating the camera, estimating mirror orientation, and subsequently lifting 2D keypoint detections to 3D skeleton pose that is used to condition the mirror-aware NeRF. We empirically demonstrate the benefit of learning a body model and accounting for occlusion in challenging mirror scenes.
ABC Easy as 123: A Blind Counter for Exemplar-Free Multi-Class Class-agnostic Counting
Sep 09, 2023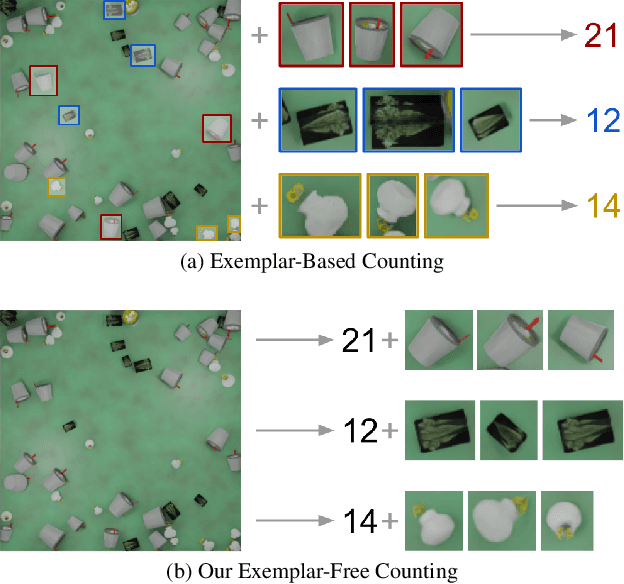
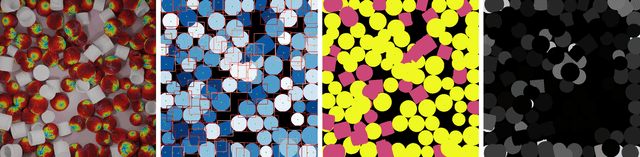
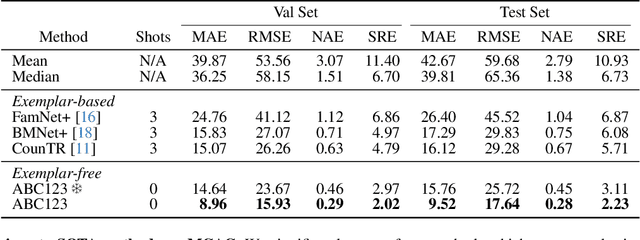

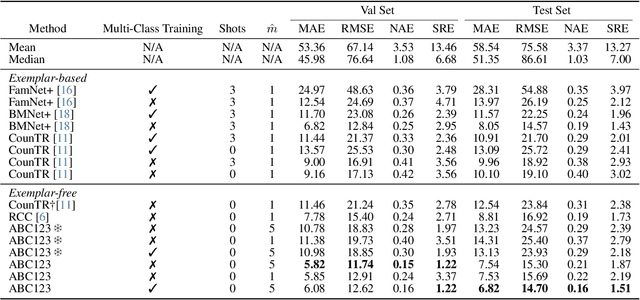
Class-agnostic counting methods enumerate objects of an arbitrary class, providing tremendous utility in many fields. Prior works have limited usefulness as they require either a set of examples of the type to be counted or that the image contains only a single type of object. A significant factor in these shortcomings is the lack of a dataset to properly address counting in settings with more than one kind of object present. To address these issues, we propose the first Multi-class, Class-Agnostic Counting dataset (MCAC) and A Blind Counter (ABC123), a method that can count multiple types of objects simultaneously without using examples of type during training or inference. ABC123 introduces a new paradigm where instead of requiring exemplars to guide the enumeration, examples are found after the counting stage to help a user understand the generated outputs. We show that ABC123 outperforms contemporary methods on MCAC without the requirement of human in-the-loop annotations. We also show that this performance transfers to FSC-147, the standard class-agnostic counting dataset.
Cheap Lunch for Medical Image Segmentation by Fine-tuning SAM on Few Exemplars
Aug 27, 2023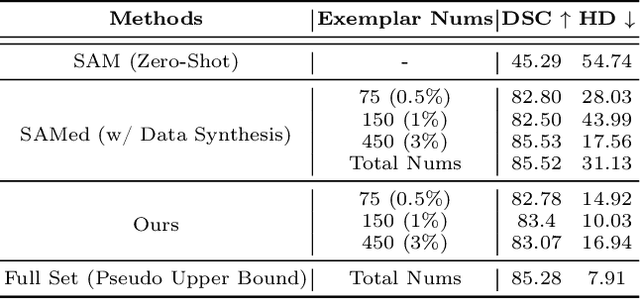
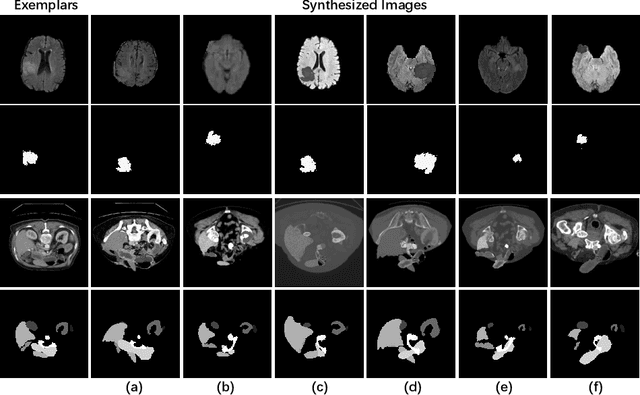
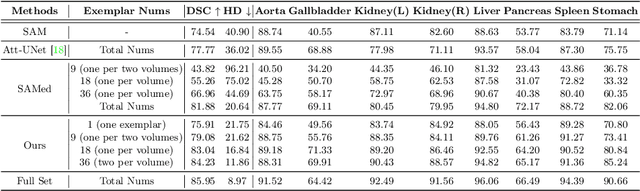
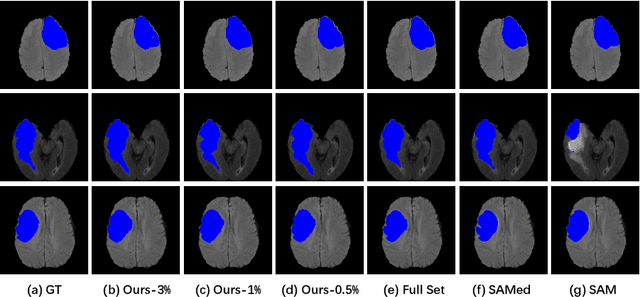
The Segment Anything Model (SAM) has demonstrated remarkable capabilities of scaled-up segmentation models, enabling zero-shot generalization across a variety of domains. By leveraging large-scale foundational models as pre-trained models, it is a natural progression to fine-tune SAM for specific domains to further enhance performances. However, the adoption of foundational models in the medical domain presents a challenge due to the difficulty and expense of labeling sufficient data for adaptation within hospital systems. In this paper, we introduce an efficient and practical approach for fine-tuning SAM using a limited number of exemplars, making it suitable for such scenarios. Our approach combines two established techniques from the literature: an exemplar-guided synthesis module and the widely recognized Low-Rank Adaptation (LoRA) fine-tuning strategy, serving as data-level and model-level attempts respectively. Interestingly, our empirical findings suggest that SAM can be effectively aligned within the medical domain even with few labeled data. We validate our approach through experiments on brain tumor segmentation (BraTS) and multi-organ CT segmentation (Synapse). The comprehensive results underscore the feasibility and effectiveness of such an approach, paving the way for the practical application of SAM in the medical domain.
Disruptive Autoencoders: Leveraging Low-level features for 3D Medical Image Pre-training
Jul 31, 2023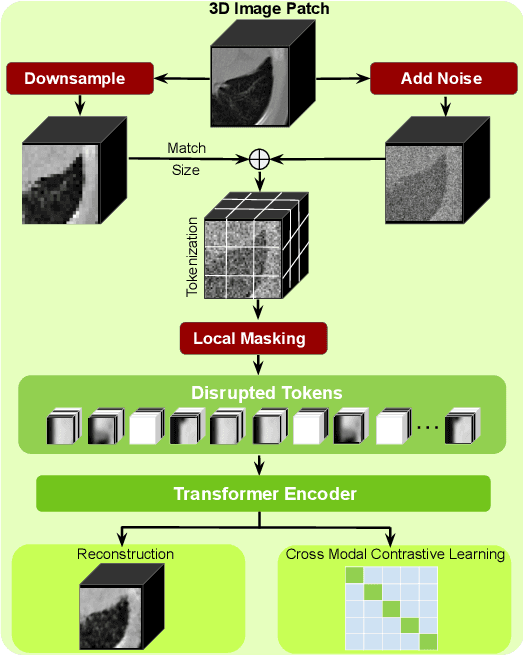
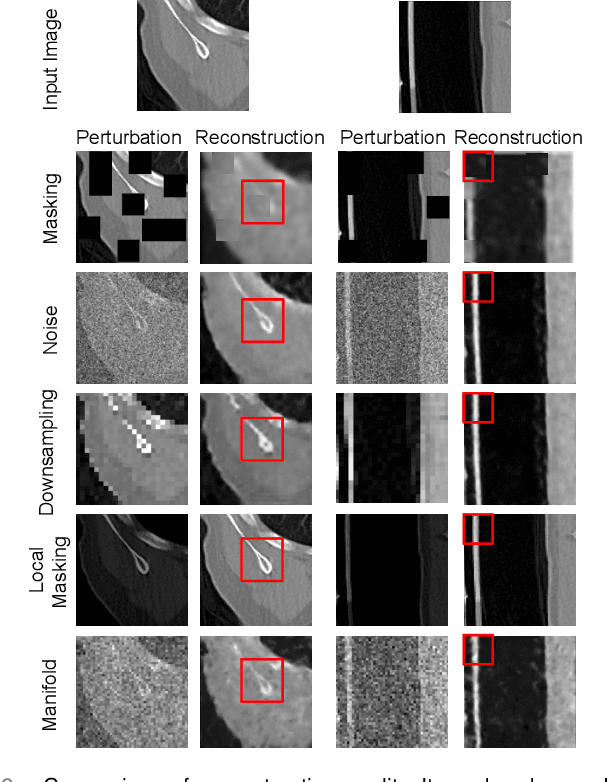

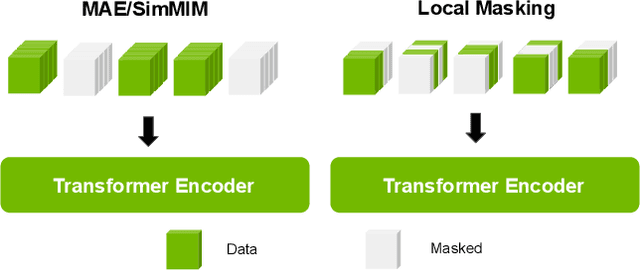
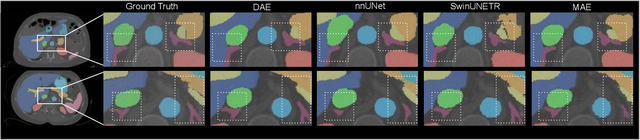
Harnessing the power of pre-training on large-scale datasets like ImageNet forms a fundamental building block for the progress of representation learning-driven solutions in computer vision. Medical images are inherently different from natural images as they are acquired in the form of many modalities (CT, MR, PET, Ultrasound etc.) and contain granulated information like tissue, lesion, organs etc. These characteristics of medical images require special attention towards learning features representative of local context. In this work, we focus on designing an effective pre-training framework for 3D radiology images. First, we propose a new masking strategy called local masking where the masking is performed across channel embeddings instead of tokens to improve the learning of local feature representations. We combine this with classical low-level perturbations like adding noise and downsampling to further enable low-level representation learning. To this end, we introduce Disruptive Autoencoders, a pre-training framework that attempts to reconstruct the original image from disruptions created by a combination of local masking and low-level perturbations. Additionally, we also devise a cross-modal contrastive loss (CMCL) to accommodate the pre-training of multiple modalities in a single framework. We curate a large-scale dataset to enable pre-training of 3D medical radiology images (MRI and CT). The proposed pre-training framework is tested across multiple downstream tasks and achieves state-of-the-art performance. Notably, our proposed method tops the public test leaderboard of BTCV multi-organ segmentation challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge