cancer detection
Cancer detection using Artificial Intelligence (AI) involves leveraging advanced machine learning algorithms and techniques to identify and diagnose cancer from various medical data sources. The goal is to enhance early detection, improve diagnostic accuracy, and potentially reduce the need for invasive procedures.
Papers and Code
Agent-Based Output Drift Detection for Breast Cancer Response Prediction in a Multisite Clinical Decision Support System
Dec 20, 2025Modern clinical decision support systems can concurrently serve multiple, independent medical imaging institutions, but their predictive performance may degrade across sites due to variations in patient populations, imaging hardware, and acquisition protocols. Continuous surveillance of predictive model outputs offers a safe and reliable approach for identifying such distributional shifts without ground truth labels. However, most existing methods rely on centralized monitoring of aggregated predictions, overlooking site-specific drift dynamics. We propose an agent-based framework for detecting drift and assessing its severity in multisite clinical AI systems. To evaluate its effectiveness, we simulate a multi-center environment for output-based drift detection, assigning each site a drift monitoring agent that performs batch-wise comparisons of model outputs against a reference distribution. We analyse several multi-center monitoring schemes, that differ in how the reference is obtained (site-specific, global, production-only and adaptive), alongside a centralized baseline. Results on real-world breast cancer imaging data using a pathological complete response prediction model shows that all multi-center schemes outperform centralized monitoring, with F1-score improvements up to 10.3% in drift detection. In the absence of site-specific references, the adaptive scheme performs best, with F1-scores of 74.3% for drift detection and 83.7% for drift severity classification. These findings suggest that adaptive, site-aware agent-based drift monitoring can enhance reliability of multisite clinical decision support systems.
Prior-Guided DETR for Ultrasound Nodule Detection
Jan 05, 2026Accurate detection of ultrasound nodules is essential for the early diagnosis and treatment of thyroid and breast cancers. However, this task remains challenging due to irregular nodule shapes, indistinct boundaries, substantial scale variations, and the presence of speckle noise that degrades structural visibility. To address these challenges, we propose a prior-guided DETR framework specifically designed for ultrasound nodule detection. Instead of relying on purely data-driven feature learning, the proposed framework progressively incorporates different prior knowledge at multiple stages of the network. First, a Spatially-adaptive Deformable FFN with Prior Regularization (SDFPR) is embedded into the CNN backbone to inject geometric priors into deformable sampling, stabilizing feature extraction for irregular and blurred nodules. Second, a Multi-scale Spatial-Frequency Feature Mixer (MSFFM) is designed to extract multi-scale structural priors, where spatial-domain processing emphasizes contour continuity and boundary cues, while frequency-domain modeling captures global morphology and suppresses speckle noise. Furthermore, a Dense Feature Interaction (DFI) mechanism propagates and exploits these prior-modulated features across all encoder layers, enabling the decoder to enhance query refinement under consistent geometric and structural guidance. Experiments conducted on two clinically collected thyroid ultrasound datasets (Thyroid I and Thyroid II) and two public benchmarks (TN3K and BUSI) for thyroid and breast nodules demonstrate that the proposed method achieves superior accuracy compared with 18 detection methods, particularly in detecting morphologically complex nodules.The source code is publicly available at https://github.com/wjj1wjj/Ultrasound-DETR.
Computational Mapping of Reactive Stroma in Prostate Cancer Yields Interpretable, Prognostic Biomarkers
Jan 10, 2026Current histopathological grading of prostate cancer relies primarily on glandular architecture, largely overlooking the tumor microenvironment. Here, we present PROTAS, a deep learning framework that quantifies reactive stroma (RS) in routine hematoxylin and eosin (H&E) slides and links stromal morphology to underlying biology. PROTAS-defined RS is characterized by nuclear enlargement, collagen disorganization, and transcriptomic enrichment of contractile pathways. PROTAS detects RS robustly in the external Prostate, Lung, Colorectal, and Ovarian (PLCO) dataset and, using domain-adversarial training, generalizes to diagnostic biopsies. In head-to-head comparisons, PROTAS outperforms pathologists for RS detection, and spatial RS features predict biochemical recurrence independently of established prognostic variables (c-index 0.80). By capturing subtle stromal phenotypes associated with tumor progression, PROTAS provides an interpretable, scalable biomarker to refine risk stratification.
LDP: Parameter-Efficient Fine-Tuning of Multimodal LLM for Medical Report Generation
Dec 11, 2025Colonoscopic polyp diagnosis is pivotal for early colorectal cancer detection, yet traditional automated reporting suffers from inconsistencies and hallucinations due to the scarcity of high-quality multimodal medical data. To bridge this gap, we propose LDP, a novel framework leveraging multimodal large language models (MLLMs) for professional polyp diagnosis report generation. Specifically, we curate MMEndo, a multimodal endoscopic dataset comprising expert-annotated colonoscopy image-text pairs. We fine-tune the Qwen2-VL-7B backbone using Parameter-Efficient Fine-Tuning (LoRA) and align it with clinical standards via Direct Preference Optimization (DPO). Extensive experiments show that our LDP outperforms existing baselines on both automated metrics and rigorous clinical expert evaluations (achieving a Physician Score of 7.2/10), significantly reducing training computational costs by 833x compared to full fine-tuning. The proposed solution offers a scalable, clinically viable path for primary healthcare, with additional validation on the IU-XRay dataset confirming its robustness.
DGSAN: Dual-Graph Spatiotemporal Attention Network for Pulmonary Nodule Malignancy Prediction
Dec 24, 2025Lung cancer continues to be the leading cause of cancer-related deaths globally. Early detection and diagnosis of pulmonary nodules are essential for improving patient survival rates. Although previous research has integrated multimodal and multi-temporal information, outperforming single modality and single time point, the fusion methods are limited to inefficient vector concatenation and simple mutual attention, highlighting the need for more effective multimodal information fusion. To address these challenges, we introduce a Dual-Graph Spatiotemporal Attention Network, which leverages temporal variations and multimodal data to enhance the accuracy of predictions. Our methodology involves developing a Global-Local Feature Encoder to better capture the local, global, and fused characteristics of pulmonary nodules. Additionally, a Dual-Graph Construction method organizes multimodal features into inter-modal and intra-modal graphs. Furthermore, a Hierarchical Cross-Modal Graph Fusion Module is introduced to refine feature integration. We also compiled a novel multimodal dataset named the NLST-cmst dataset as a comprehensive source of support for related research. Our extensive experiments, conducted on both the NLST-cmst and curated CSTL-derived datasets, demonstrate that our DGSAN significantly outperforms state-of-the-art methods in classifying pulmonary nodules with exceptional computational efficiency.
Feature Learning with Multi-Stage Vision Transformers on Inter-Modality HER2 Status Scoring and Tumor Classification on Whole Slides
Dec 26, 2025The popular use of histopathology images, such as hematoxylin and eosin (H&E), has proven to be useful in detecting tumors. However, moving such cancer cases forward for treatment requires accurate on the amount of the human epidermal growth factor receptor 2 (HER2) protein expression. Predicting both the lower and higher levels of HER2 can be challenging. Moreover, jointly analyzing H&E and immunohistochemistry (IHC) stained images for HER2 scoring is difficult. Although several deep learning methods have been investigated to address the challenge of HER2 scoring, they suffer from providing a pixel-level localization of HER2 status. In this study, we propose a single end-to-end pipeline using a system of vision transformers with HER2 status scoring on whole slide images of WSIs. The method includes patch-wise processing of H&E WSIs for tumor localization. A novel mapping function is proposed to correspondingly identify correlated IHC WSIs regions with malignant regions on H&E. A clinically inspired HER2 scoring mechanism is embedded in the pipeline and allows for automatic pixel-level annotation of 4-way HER2 scoring (0, 1+, 2+, and 3+). Also, the proposed method accurately returns HER2-negative and HER2-positive. Privately curated datasets were collaboratively extracted from 13 different cases of WSIs of H&E and IHC. A thorough experiment is conducted on the proposed method. Results obtained showed a good classification accuracy during tumor localization. Also, a classification accuracy of 0.94 and a specificity of 0.933 were returned for the prediction of HER2 status, scoring in the 4-way methods. The applicability of the proposed pipeline was investigated using WSIs patches as comparable to human pathologists. Findings from the study showed the usability of jointly evaluated H&E and IHC images on end-to-end ViTs-based models for HER2 scoring
See More, Change Less: Anatomy-Aware Diffusion for Contrast Enhancement
Dec 08, 2025Image enhancement improves visual quality and helps reveal details that are hard to see in the original image. In medical imaging, it can support clinical decision-making, but current models often over-edit. This can distort organs, create false findings, and miss small tumors because these models do not understand anatomy or contrast dynamics. We propose SMILE, an anatomy-aware diffusion model that learns how organs are shaped and how they take up contrast. It enhances only clinically relevant regions while leaving all other areas unchanged. SMILE introduces three key ideas: (1) structure-aware supervision that follows true organ boundaries and contrast patterns; (2) registration-free learning that works directly with unaligned multi-phase CT scans; (3) unified inference that provides fast and consistent enhancement across all contrast phases. Across six external datasets, SMILE outperforms existing methods in image quality (14.2% higher SSIM, 20.6% higher PSNR, 50% better FID) and in clinical usefulness by producing anatomically accurate and diagnostically meaningful images. SMILE also improves cancer detection from non-contrast CT, raising the F1 score by up to 10 percent.
Skin Lesion Classification Using a Soft Voting Ensemble of Convolutional Neural Networks
Dec 23, 2025Skin cancer can be identified by dermoscopic examination and ocular inspection, but early detection significantly increases survival chances. Artificial intelligence (AI), using annotated skin images and Convolutional Neural Networks (CNNs), improves diagnostic accuracy. This paper presents an early skin cancer classification method using a soft voting ensemble of CNNs. In this investigation, three benchmark datasets, namely HAM10000, ISIC 2016, and ISIC 2019, were used. The process involved rebalancing, image augmentation, and filtering techniques, followed by a hybrid dual encoder for segmentation via transfer learning. Accurate segmentation focused classification models on clinically significant features, reducing background artifacts and improving accuracy. Classification was performed through an ensemble of MobileNetV2, VGG19, and InceptionV3, balancing accuracy and speed for real-world deployment. The method achieved lesion recognition accuracies of 96.32\%, 90.86\%, and 93.92\% for the three datasets. The system performance was evaluated using established skin lesion detection metrics, yielding impressive results.
Tumor-anchored deep feature random forests for out-of-distribution detection in lung cancer segmentation
Dec 09, 2025Accurate segmentation of cancerous lesions from 3D computed tomography (CT) scans is essential for automated treatment planning and response assessment. However, even state-of-the-art models combining self-supervised learning (SSL) pretrained transformers with convolutional decoders are susceptible to out-of-distribution (OOD) inputs, generating confidently incorrect tumor segmentations, posing risks for safe clinical deployment. Existing logit-based methods suffer from task-specific model biases, while architectural enhancements to explicitly detect OOD increase parameters and computational costs. Hence, we introduce a plug-and-play and lightweight post-hoc random forests-based OOD detection framework called RF-Deep that leverages deep features with limited outlier exposure. RF-Deep enhances generalization to imaging variations by repurposing the hierarchical features from the pretrained-then-finetuned backbone encoder, providing task-relevant OOD detection by extracting the features from multiple regions of interest anchored to the predicted tumor segmentations. Hence, it scales to images of varying fields-of-view. We compared RF-Deep against existing OOD detection methods using 1,916 CT scans across near-OOD (pulmonary embolism, negative COVID-19) and far-OOD (kidney cancer, healthy pancreas) datasets. RF-Deep achieved AUROC > 93.50 for the challenging near-OOD datasets and near-perfect detection (AUROC > 99.00) for the far-OOD datasets, substantially outperforming logit-based and radiomics approaches. RF-Deep maintained similar performance consistency across networks of different depths and pretraining strategies, demonstrating its effectiveness as a lightweight, architecture-agnostic approach to enhance the reliability of tumor segmentation from CT volumes.
DBT-DINO: Towards Foundation model based analysis of Digital Breast Tomosynthesis
Dec 15, 2025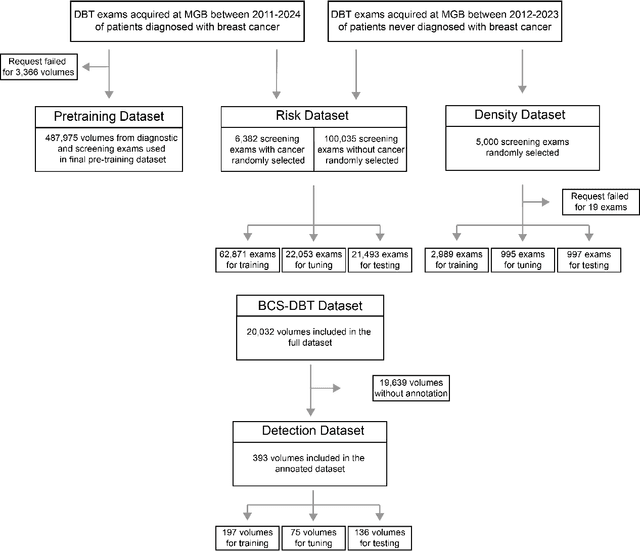
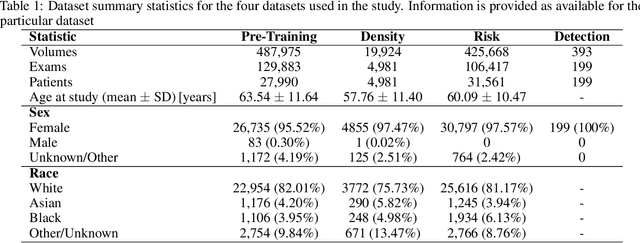
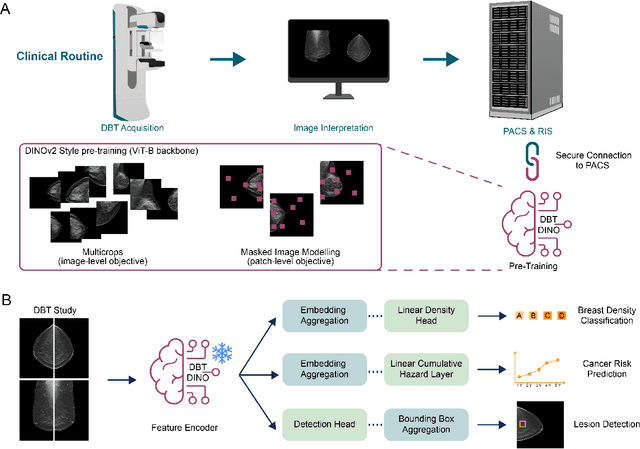
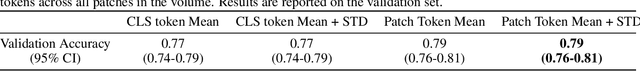
Foundation models have shown promise in medical imaging but remain underexplored for three-dimensional imaging modalities. No foundation model currently exists for Digital Breast Tomosynthesis (DBT), despite its use for breast cancer screening. To develop and evaluate a foundation model for DBT (DBT-DINO) across multiple clinical tasks and assess the impact of domain-specific pre-training. Self-supervised pre-training was performed using the DINOv2 methodology on over 25 million 2D slices from 487,975 DBT volumes from 27,990 patients. Three downstream tasks were evaluated: (1) breast density classification using 5,000 screening exams; (2) 5-year risk of developing breast cancer using 106,417 screening exams; and (3) lesion detection using 393 annotated volumes. For breast density classification, DBT-DINO achieved an accuracy of 0.79 (95\% CI: 0.76--0.81), outperforming both the MetaAI DINOv2 baseline (0.73, 95\% CI: 0.70--0.76, p<.001) and DenseNet-121 (0.74, 95\% CI: 0.71--0.76, p<.001). For 5-year breast cancer risk prediction, DBT-DINO achieved an AUROC of 0.78 (95\% CI: 0.76--0.80) compared to DINOv2's 0.76 (95\% CI: 0.74--0.78, p=.57). For lesion detection, DINOv2 achieved a higher average sensitivity of 0.67 (95\% CI: 0.60--0.74) compared to DBT-DINO with 0.62 (95\% CI: 0.53--0.71, p=.60). DBT-DINO demonstrated better performance on cancerous lesions specifically with a detection rate of 78.8\% compared to Dinov2's 77.3\%. Using a dataset of unprecedented size, we developed DBT-DINO, the first foundation model for DBT. DBT-DINO demonstrated strong performance on breast density classification and cancer risk prediction. However, domain-specific pre-training showed variable benefits on the detection task, with ImageNet baseline outperforming DBT-DINO on general lesion detection, indicating that localized detection tasks require further methodological development.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge