cancer detection
Cancer detection using Artificial Intelligence (AI) involves leveraging advanced machine learning algorithms and techniques to identify and diagnose cancer from various medical data sources. The goal is to enhance early detection, improve diagnostic accuracy, and potentially reduce the need for invasive procedures.
Papers and Code
Lung Nodule-SSM: Self-Supervised Lung Nodule Detection and Classification in Thoracic CT Images
May 21, 2025
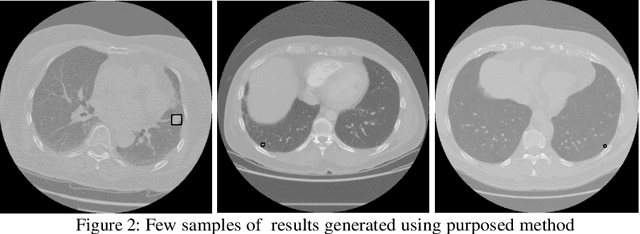
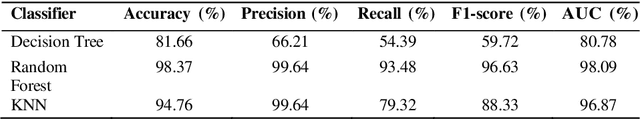
Lung cancer remains among the deadliest types of cancer in recent decades, and early lung nodule detection is crucial for improving patient outcomes. The limited availability of annotated medical imaging data remains a bottleneck in developing accurate computer-aided diagnosis (CAD) systems. Self-supervised learning can help leverage large amounts of unlabeled data to develop more robust CAD systems. With the recent advent of transformer-based architecture and their ability to generalize to unseen tasks, there has been an effort within the healthcare community to adapt them to various medical downstream tasks. Thus, we propose a novel "LungNodule-SSM" method, which utilizes selfsupervised learning with DINOv2 as a backbone to enhance lung nodule detection and classification without annotated data. Our methodology has two stages: firstly, the DINOv2 model is pre-trained on unlabeled CT scans to learn robust feature representations, then secondly, these features are fine-tuned using transformer-based architectures for lesionlevel detection and accurate lung nodule diagnosis. The proposed method has been evaluated on the challenging LUNA 16 dataset, consisting of 888 CT scans, and compared with SOTA methods. Our experimental results show the superiority of our proposed method with an accuracy of 98.37%, explaining its effectiveness in lung nodule detection. The source code, datasets, and pre-processed data can be accessed using the link:https://github.com/EMeRALDsNRPU/Lung-Nodule-SSM-Self-Supervised-Lung-Nodule-Detection-and-Classification/tree/main
Deep Learning Enabled Segmentation, Classification and Risk Assessment of Cervical Cancer
May 21, 2025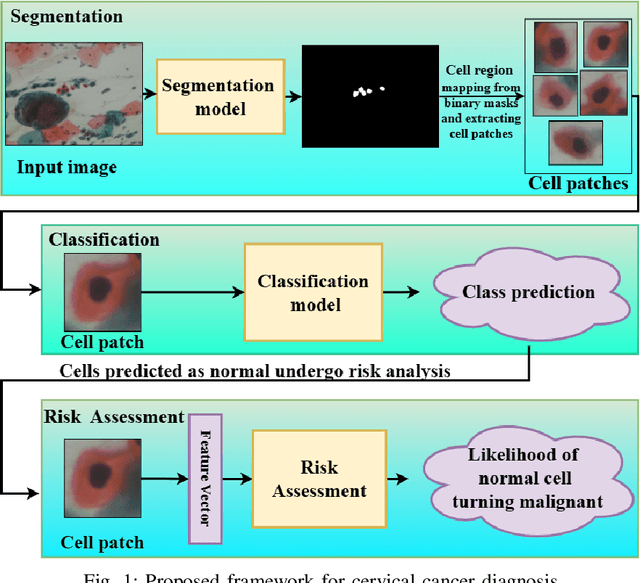
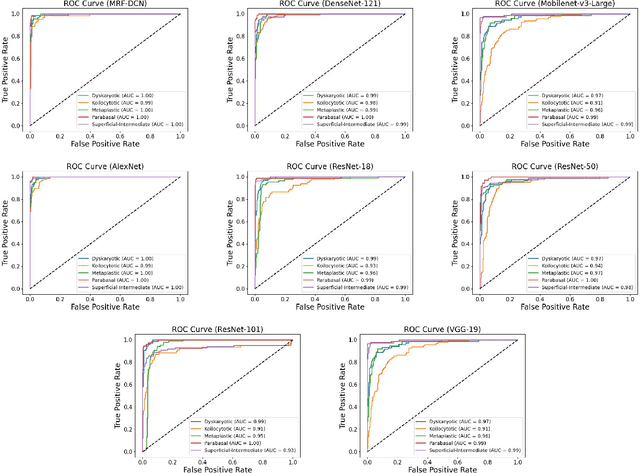
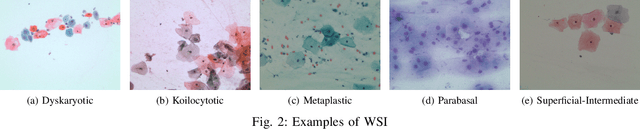
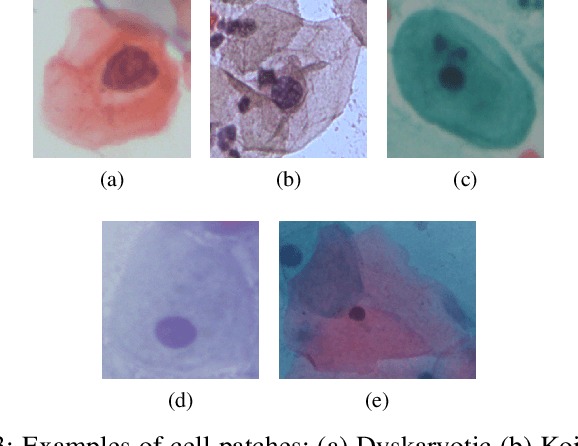
Cervical cancer, the fourth leading cause of cancer in women globally, requires early detection through Pap smear tests to identify precancerous changes and prevent disease progression. In this study, we performed a focused analysis by segmenting the cellular boundaries and drawing bounding boxes to isolate the cancer cells. A novel Deep Learning (DL) architecture, the ``Multi-Resolution Fusion Deep Convolutional Network", was proposed to effectively handle images with varying resolutions and aspect ratios, with its efficacy showcased using the SIPaKMeD dataset. The performance of this DL model was observed to be similar to the state-of-the-art models, with accuracy variations of a mere 2\% to 3\%, achieved using just 1.7 million learnable parameters, which is approximately 85 times less than the VGG-19 model. Furthermore, we introduced a multi-task learning technique that simultaneously performs segmentation and classification tasks and begets an Intersection over Union score of 0.83 and a classification accuracy of 90\%. The final stage of the workflow employs a probabilistic approach for risk assessment, extracting feature vectors to predict the likelihood of normal cells progressing to malignant states, which can be utilized for the prognosis of cervical cancer.
Towards Facilitated Fairness Assessment of AI-based Skin Lesion Classifiers Through GenAI-based Image Synthesis
Jul 23, 2025Recent advancements in Deep Learning and its application on the edge hold great potential for the revolution of routine screenings for skin cancers like Melanoma. Along with the anticipated benefits of this technology, potential dangers arise from unforseen and inherent biases. Thus, assessing and improving the fairness of such systems is of utmost importance. A key challenge in fairness assessment is to ensure that the evaluation dataset is sufficiently representative of different Personal Identifiable Information (PII) (sex, age, and race) and other minority groups. Against the backdrop of this challenge, this study leverages the state-of-the-art Generative AI (GenAI) LightningDiT model to assess the fairness of publicly available melanoma classifiers. The results suggest that fairness assessment using highly realistic synthetic data is a promising direction. Yet, our findings indicate that verifying fairness becomes difficult when the melanoma-detection model used for evaluation is trained on data that differ from the dataset underpinning the synthetic images. Nonetheless, we propose that our approach offers a valuable new avenue for employing synthetic data to gauge and enhance fairness in medical-imaging GenAI systems.
Optimizing Neuro-Fuzzy and Colonial Competition Algorithms for Skin Cancer Diagnosis in Dermatoscopic Images
May 13, 2025



The rising incidence of skin cancer, coupled with limited public awareness and a shortfall in clinical expertise, underscores an urgent need for advanced diagnostic aids. Artificial Intelligence (AI) has emerged as a promising tool in this domain, particularly for distinguishing malignant from benign skin lesions. Leveraging publicly available datasets of skin lesions, researchers have been developing AI-based diagnostic solutions. However, the integration of such computer systems in clinical settings is still nascent. This study aims to bridge this gap by employing a fusion of image processing techniques and machine learning algorithms, specifically neuro-fuzzy and colonial competition approaches. Applied to dermoscopic images from the ISIC database, our method achieved a notable accuracy of 94% on a dataset of 560 images. These results underscore the potential of our approach in aiding clinicians in the early detection of melanoma, thereby contributing significantly to skin cancer diagnostics.
* 7 pages, 10 figures. Accepted at the 2nd Asia Pacific Computer Systems Conference (APCS 2024), March 15-17, 2024
Learning from Anatomy: Supervised Anatomical Pretraining (SAP) for Improved Metastatic Bone Disease Segmentation in Whole-Body MRI
Jun 24, 2025The segmentation of metastatic bone disease (MBD) in whole-body MRI (WB-MRI) is a challenging problem. Due to varying appearances and anatomical locations of lesions, ambiguous boundaries, and severe class imbalance, obtaining reliable segmentations requires large, well-annotated datasets capturing lesion variability. Generating such datasets requires substantial time and expertise, and is prone to error. While self-supervised learning (SSL) can leverage large unlabeled datasets, learned generic representations often fail to capture the nuanced features needed for accurate lesion detection. In this work, we propose a Supervised Anatomical Pretraining (SAP) method that learns from a limited dataset of anatomical labels. First, an MRI-based skeletal segmentation model is developed and trained on WB-MRI scans from healthy individuals for high-quality skeletal delineation. Then, we compare its downstream efficacy in segmenting MBD on a cohort of 44 patients with metastatic prostate cancer, against both a baseline random initialization and a state-of-the-art SSL method. SAP significantly outperforms both the baseline and SSL-pretrained models, achieving a normalized surface Dice of 0.76 and a Dice coefficient of 0.64. The method achieved a lesion detection F2 score of 0.44, improving on 0.24 (baseline) and 0.31 (SSL). When considering only clinically relevant lesions larger than 1~ml, SAP achieves a detection sensitivity of 100% in 28 out of 32 patients. Learning bone morphology from anatomy yields an effective and domain-relevant inductive bias that can be leveraged for the downstream segmentation task of bone lesions. All code and models are made publicly available.
OpenPros: A Large-Scale Dataset for Limited View Prostate Ultrasound Computed Tomography
May 18, 2025Prostate cancer is one of the most common and lethal cancers among men, making its early detection critically important. Although ultrasound imaging offers greater accessibility and cost-effectiveness compared to MRI, traditional transrectal ultrasound methods suffer from low sensitivity, especially in detecting anteriorly located tumors. Ultrasound computed tomography provides quantitative tissue characterization, but its clinical implementation faces significant challenges, particularly under anatomically constrained limited-angle acquisition conditions specific to prostate imaging. To address these unmet needs, we introduce OpenPros, the first large-scale benchmark dataset explicitly developed for limited-view prostate USCT. Our dataset includes over 280,000 paired samples of realistic 2D speed-of-sound (SOS) phantoms and corresponding ultrasound full-waveform data, generated from anatomically accurate 3D digital prostate models derived from real clinical MRI/CT scans and ex vivo ultrasound measurements, annotated by medical experts. Simulations are conducted under clinically realistic configurations using advanced finite-difference time-domain and Runge-Kutta acoustic wave solvers, both provided as open-source components. Through comprehensive baseline experiments, we demonstrate that state-of-the-art deep learning methods surpass traditional physics-based approaches in both inference efficiency and reconstruction accuracy. Nevertheless, current deep learning models still fall short of delivering clinically acceptable high-resolution images with sufficient accuracy. By publicly releasing OpenPros, we aim to encourage the development of advanced machine learning algorithms capable of bridging this performance gap and producing clinically usable, high-resolution, and highly accurate prostate ultrasound images. The dataset is publicly accessible at https://open-pros.github.io/.
Anomaly Detection and Improvement of Clusters using Enhanced K-Means Algorithm
May 30, 2025This paper introduces a unified approach to cluster refinement and anomaly detection in datasets. We propose a novel algorithm that iteratively reduces the intra-cluster variance of N clusters until a global minimum is reached, yielding tighter clusters than the standard k-means algorithm. We evaluate the method using intrinsic measures for unsupervised learning, including the silhouette coefficient, Calinski-Harabasz index, and Davies-Bouldin index, and extend it to anomaly detection by identifying points whose assignment causes a significant variance increase. External validation on synthetic data and the UCI Breast Cancer and UCI Wine Quality datasets employs the Jaccard similarity score, V-measure, and F1 score. Results show variance reductions of 18.7% and 88.1% on the synthetic and Wine Quality datasets, respectively, along with accuracy and F1 score improvements of 22.5% and 20.8% on the Wine Quality dataset.
Generalizing imaging biomarker repeatability studies using Bayesian inference: Applications in detecting heterogeneous treatment response in whole-body diffusion-weighted MRI of metastatic prostate cancer
May 14, 2025The assessment of imaging biomarkers is critical for advancing precision medicine and improving disease characterization. Despite the availability of methods to derive disease heterogeneity metrics in imaging studies, a robust framework for evaluating measurement uncertainty remains underdeveloped. To address this gap, we propose a novel Bayesian framework to assess the precision of disease heterogeneity measures in biomarker studies. Our approach extends traditional methods for evaluating biomarker precision by providing greater flexibility in statistical assumptions and enabling the analysis of biomarkers beyond univariate or multivariate normally-distributed variables. Using Hamiltonian Monte Carlo sampling, the framework supports both, for example, normally-distributed and Dirichlet-Multinomial distributed variables, enabling the derivation of posterior distributions for biomarker parameters under diverse model assumptions. Designed to be broadly applicable across various imaging modalities and biomarker types, the framework builds a foundation for generalizing reproducible and objective biomarker evaluation. To demonstrate utility, we apply the framework to whole-body diffusion-weighted MRI (WBDWI) to assess heterogeneous therapeutic responses in metastatic bone disease. Specifically, we analyze data from two patient studies investigating treatments for metastatic castrate-resistant prostate cancer (mCRPC). Our results reveal an approximately 70% response rate among individual tumors across both studies, objectively characterizing differential responses to systemic therapies and validating the clinical relevance of the proposed methodology. This Bayesian framework provides a powerful tool for advancing biomarker research across diverse imaging-based studies while offering valuable insights into specific clinical applications, such as mCRPC treatment response.
Improving Oral Cancer Outcomes Through Machine Learning and Dimensionality Reduction
Jun 11, 2025Oral cancer presents a formidable challenge in oncology, necessitating early diagnosis and accurate prognosis to enhance patient survival rates. Recent advancements in machine learning and data mining have revolutionized traditional diagnostic methodologies, providing sophisticated and automated tools for differentiating between benign and malignant oral lesions. This study presents a comprehensive review of cutting-edge data mining methodologies, including Neural Networks, K-Nearest Neighbors (KNN), Support Vector Machines (SVM), and ensemble learning techniques, specifically applied to the diagnosis and prognosis of oral cancer. Through a rigorous comparative analysis, our findings reveal that Neural Networks surpass other models, achieving an impressive classification accuracy of 93,6 % in predicting oral cancer. Furthermore, we underscore the potential benefits of integrating feature selection and dimensionality reduction techniques to enhance model performance. These insights underscore the significant promise of advanced data mining techniques in bolstering early detection, optimizing treatment strategies, and ultimately improving patient outcomes in the realm of oral oncology.
A Narrative Review on Large AI Models in Lung Cancer Screening, Diagnosis, and Treatment Planning
Jun 08, 2025Lung cancer remains one of the most prevalent and fatal diseases worldwide, demanding accurate and timely diagnosis and treatment. Recent advancements in large AI models have significantly enhanced medical image understanding and clinical decision-making. This review systematically surveys the state-of-the-art in applying large AI models to lung cancer screening, diagnosis, prognosis, and treatment. We categorize existing models into modality-specific encoders, encoder-decoder frameworks, and joint encoder architectures, highlighting key examples such as CLIP, BLIP, Flamingo, BioViL-T, and GLoRIA. We further examine their performance in multimodal learning tasks using benchmark datasets like LIDC-IDRI, NLST, and MIMIC-CXR. Applications span pulmonary nodule detection, gene mutation prediction, multi-omics integration, and personalized treatment planning, with emerging evidence of clinical deployment and validation. Finally, we discuss current limitations in generalizability, interpretability, and regulatory compliance, proposing future directions for building scalable, explainable, and clinically integrated AI systems. Our review underscores the transformative potential of large AI models to personalize and optimize lung cancer care.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge