Mr Apex
Papers and Code
APEX-MR: Multi-Robot Asynchronous Planning and Execution for Cooperative Assembly
Mar 20, 2025Compared to a single-robot workstation, a multi-robot system offers several advantages: 1) it expands the system's workspace, 2) improves task efficiency, and more importantly, 3) enables robots to achieve significantly more complex and dexterous tasks, such as cooperative assembly. However, coordinating the tasks and motions of multiple robots is challenging due to issues, e.g. system uncertainty, task efficiency, algorithm scalability, and safety concerns. To address these challenges, this paper studies multi-robot coordination and proposes APEX-MR, an asynchronous planning and execution framework designed to safely and efficiently coordinate multiple robots to achieve cooperative assembly, e.g. LEGO assembly. In particular, APEX-MR provides a systematic approach to post-process multi-robot tasks and motion plans to enable robust asynchronous execution under uncertainty. Experimental results demonstrate that APEX-MR can significantly speed up the execution time of many long-horizon LEGO assembly tasks by 48% compared to sequential planning and 36% compared to synchronous planning on average. To further demonstrate the performance, we deploy APEX-MR to a dual-arm system to perform physical LEGO assembly. To our knowledge, this is the first robotic system capable of performing customized LEGO assembly using commercial LEGO bricks. The experiment results demonstrate that the dual-arm system, with APEX-MR, can safely coordinate robot motions, efficiently collaborate, and construct complex LEGO structures. Our project website is available at https://intelligent-control-lab.github.io/APEX-MR/
Automatic Segmentation of the Prostate on 3D Trans-rectal Ultrasound Images using Statistical Shape Models and Convolutional Neural Networks
Jun 17, 2021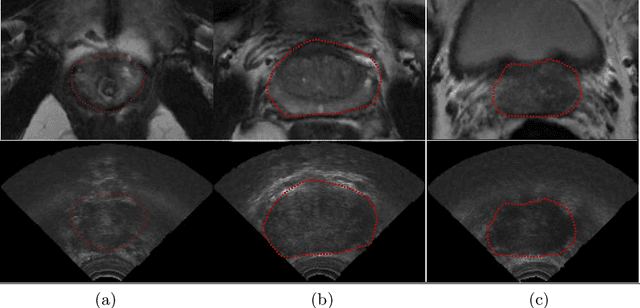
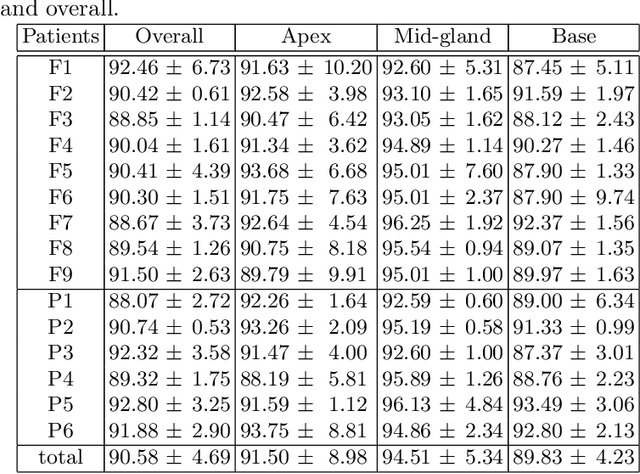
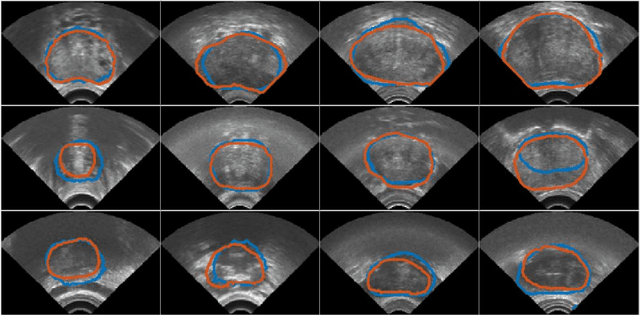
In this work we propose to segment the prostate on a challenging dataset of trans-rectal ultrasound (TRUS) images using convolutional neural networks (CNNs) and statistical shape models (SSMs). TRUS is commonly used for a number of image-guided interventions on the prostate. Fast and accurate segmentation on the organ in these images is crucial to planning and fusion with other modalities such as magnetic resonance images (MRIs) . However, TRUS has limited soft tissue contrast and signal to noise ratio which makes the task of segmenting the prostate challenging and subject to inter-observer and intra-observer variability. This is especially problematic at the base and apex where the gland boundary is hard to define. In this paper, we aim to tackle this problem by taking advantage of shape priors learnt on an MR dataset which has higher soft tissue contrast allowing the prostate to be contoured more accurately. We use this shape prior in combination with a prostate tissue probability map computed by a CNN for segmentation.
Shape Constrained CNN for Cardiac MR Segmentation with Simultaneous Prediction of Shape and Pose Parameters
Oct 18, 2020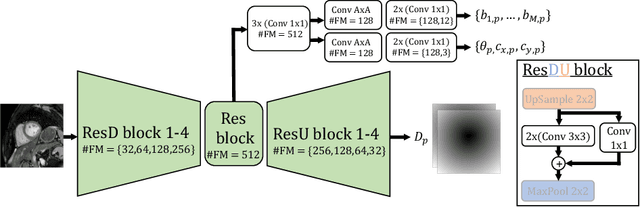
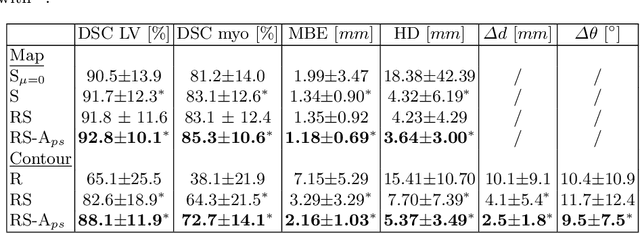
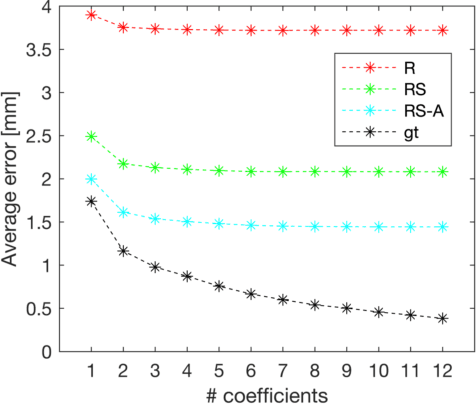
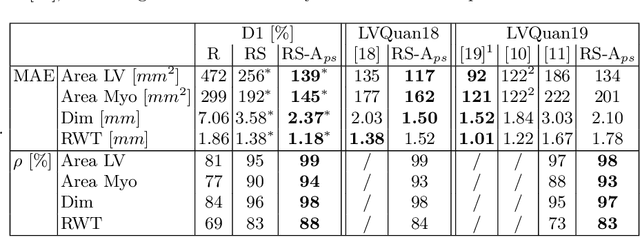
Semantic segmentation using convolutional neural networks (CNNs) is the state-of-the-art for many medical segmentation tasks including left ventricle (LV) segmentation in cardiac MR images. However, a drawback is that these CNNs lack explicit shape constraints, occasionally resulting in unrealistic segmentations. In this paper, we perform LV and myocardial segmentation by regression of pose and shape parameters derived from a statistical shape model. The integrated shape model regularizes predicted segmentations and guarantees realistic shapes. Furthermore, in contrast to semantic segmentation, it allows direct calculation of regional measures such as myocardial thickness. We enforce robustness of shape and pose prediction by simultaneously constructing a segmentation distance map during training. We evaluated the proposed method in a fivefold cross validation on a in-house clinical dataset with 75 subjects containing a total of 1539 delineated short-axis slices covering LV from apex to base, and achieved a correlation of 99% for LV area, 94% for myocardial area, 98% for LV dimensions and 88% for regional wall thicknesses. The method was additionally validated on the LVQuan18 and LVQuan19 public datasets and achieved state-of-the-art results.
Anisotropic 3D Multi-Stream CNN for Accurate Prostate Segmentation from Multi-Planar MRI
Sep 23, 2020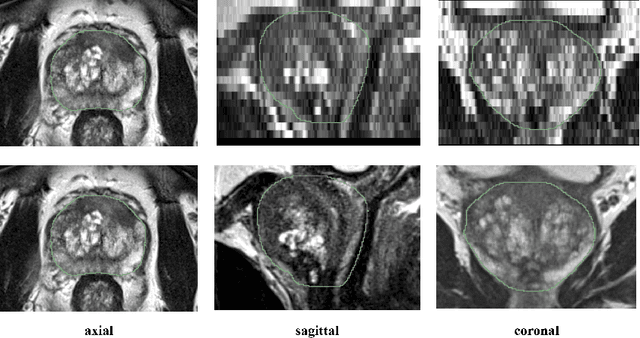
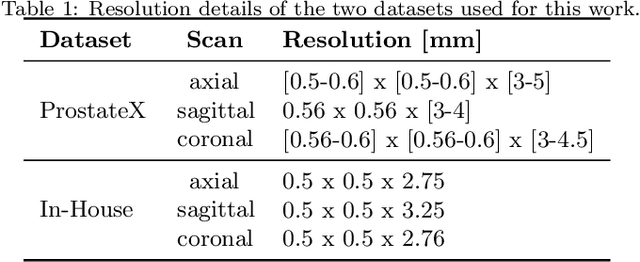
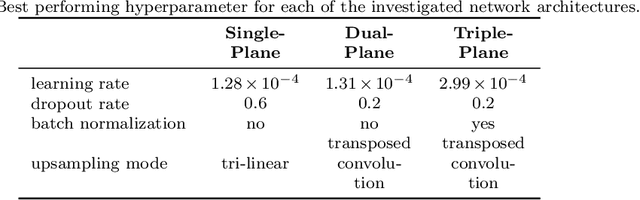

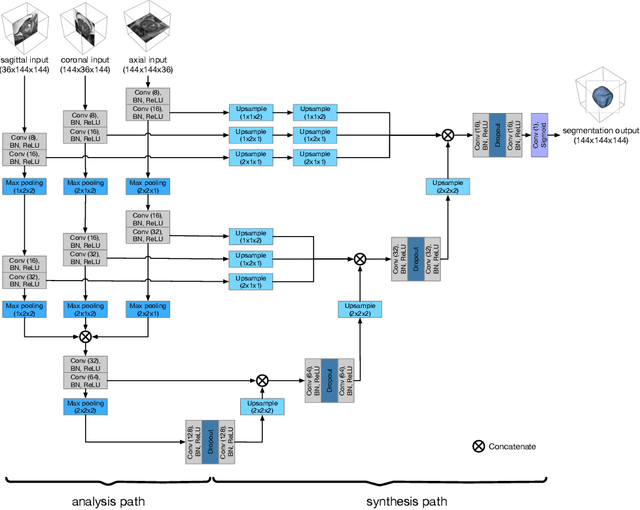

Background and Objective: Accurate and reliable segmentation of the prostate gland in MR images can support the clinical assessment of prostate cancer, as well as the planning and monitoring of focal and loco-regional therapeutic interventions. Despite the availability of multi-planar MR scans due to standardized protocols, the majority of segmentation approaches presented in the literature consider the axial scans only. Methods: We propose an anisotropic 3D multi-stream CNN architecture, which processes additional scan directions to produce a higher-resolution isotropic prostate segmentation. We investigate two variants of our architecture, which work on two (dual-plane) and three (triple-plane) image orientations, respectively. We compare them with the standard baseline (single-plane) used in literature, i.e., plain axial segmentation. To realize a fair comparison, we employ a hyperparameter optimization strategy to select optimal configurations for the individual approaches. Results: Training and evaluation on two datasets spanning multiple sites obtain statistical significant improvement over the plain axial segmentation ($p<0.05$ on the Dice similarity coefficient). The improvement can be observed especially at the base ($0.898$ single-plane vs. $0.906$ triple-plane) and apex ($0.888$ single-plane vs. $0.901$ dual-plane). Conclusion: This study indicates that models employing two or three scan directions are superior to plain axial segmentation. The knowledge of precise boundaries of the prostate is crucial for the conservation of risk structures. Thus, the proposed models have the potential to improve the outcome of prostate cancer diagnosis and therapies.
Learning Shape Priors for Robust Cardiac MR Segmentation from Multi-view Images
Jul 23, 2019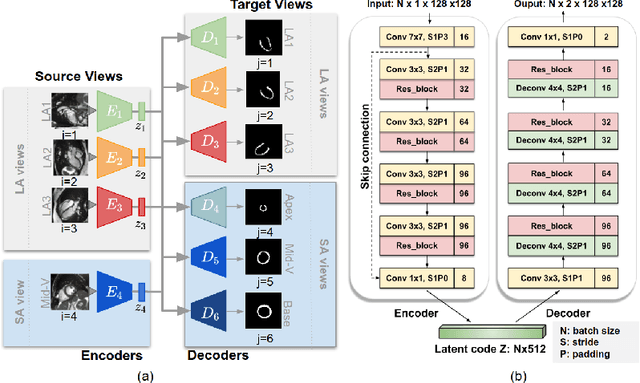
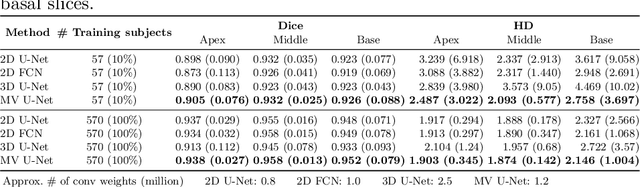
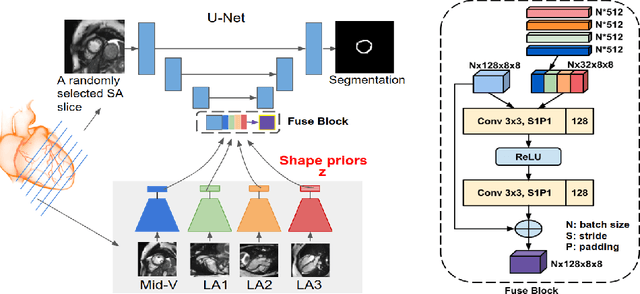
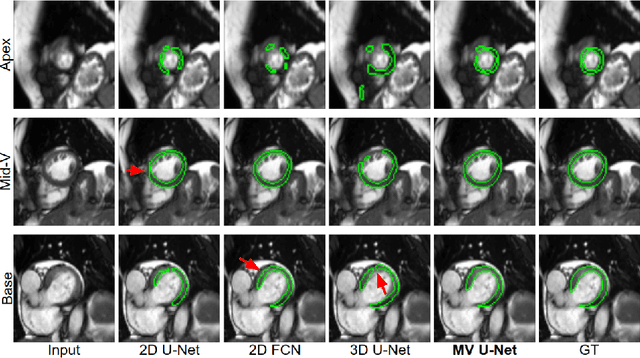
Cardiac MR image segmentation is essential for the morphological and functional analysis of the heart. Inspired by how experienced clinicians assess the cardiac morphology and function across multiple standard views (i.e. long- and short-axis views), we propose a novel approach which learns anatomical shape priors across different 2D standard views and leverages these priors to segment the left ventricular (LV) myocardium from short-axis MR image stacks. The proposed segmentation method has the advantage of being a 2D network but at the same time incorporates spatial context from multiple, complementary views that span a 3D space. Our method achieves accurate and robust segmentation of the myocardium across different short-axis slices (from apex to base), outperforming baseline models (e.g. 2D U-Net, 3D U-Net) while achieving higher data efficiency. Compared to the 2D U-Net, the proposed method reduces the mean Hausdorff distance (mm) from 3.24 to 2.49 on the apical slices, from 2.34 to 2.09 on the middle slices and from 3.62 to 2.76 on the basal slices on the test set, when only 10% of the training data was used.
Deeply-Supervised CNN for Prostate Segmentation
Mar 28, 2017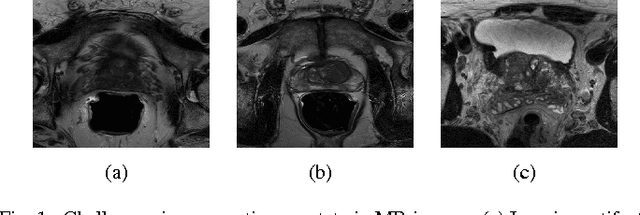
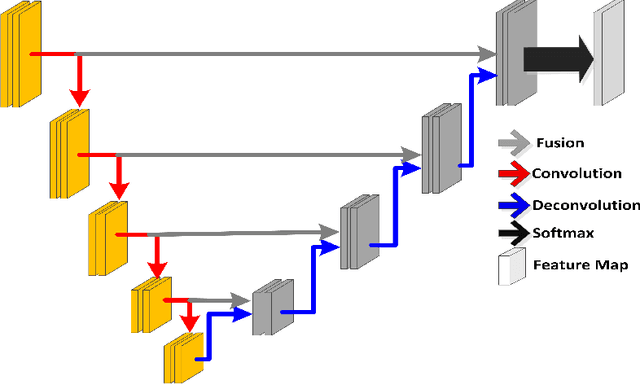
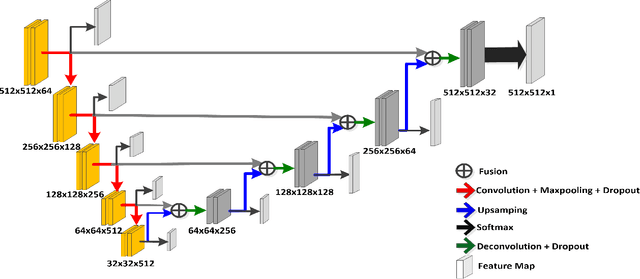
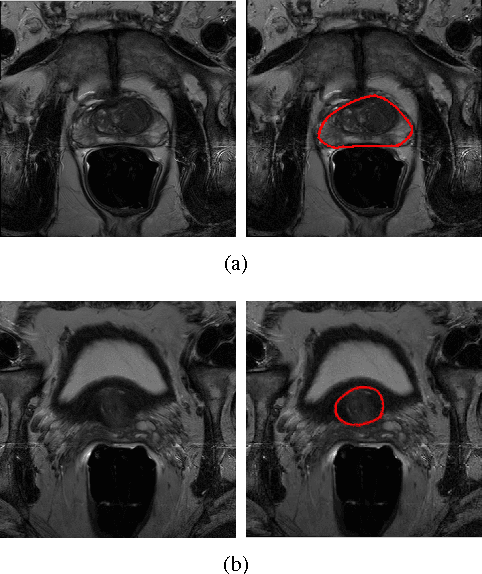
Prostate segmentation from Magnetic Resonance (MR) images plays an important role in image guided interven- tion. However, the lack of clear boundary specifically at the apex and base, and huge variation of shape and texture between the images from different patients make the task very challenging. To overcome these problems, in this paper, we propose a deeply supervised convolutional neural network (CNN) utilizing the convolutional information to accurately segment the prostate from MR images. The proposed model can effectively detect the prostate region with additional deeply supervised layers compared with other approaches. Since some information will be abandoned after convolution, it is necessary to pass the features extracted from early stages to later stages. The experimental results show that significant segmentation accuracy improvement has been achieved by our proposed method compared to other reported approaches.
Recurrent Fully Convolutional Neural Networks for Multi-slice MRI Cardiac Segmentation
Aug 13, 2016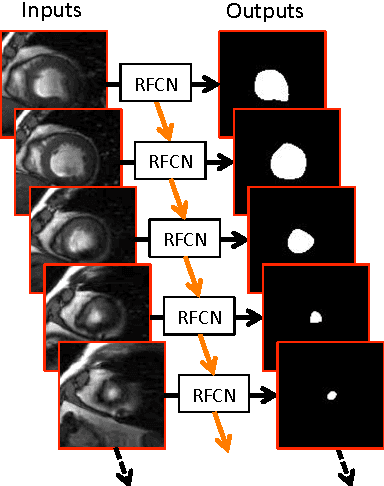
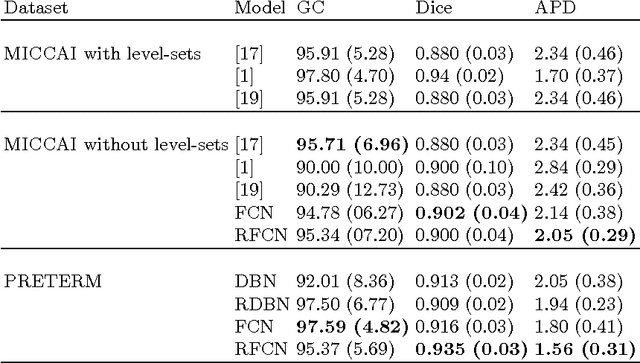
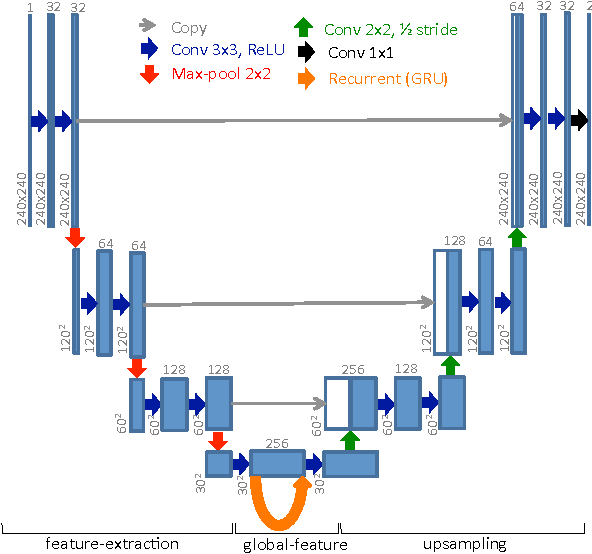
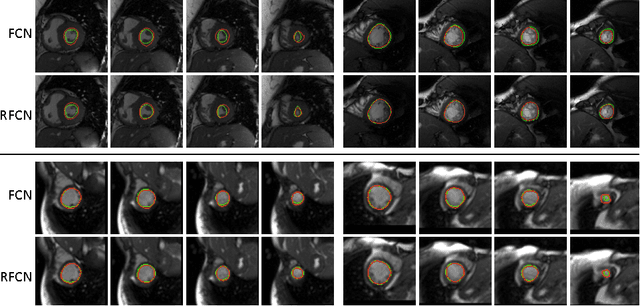
In cardiac magnetic resonance imaging, fully-automatic segmentation of the heart enables precise structural and functional measurements to be taken, e.g. from short-axis MR images of the left-ventricle. In this work we propose a recurrent fully-convolutional network (RFCN) that learns image representations from the full stack of 2D slices and has the ability to leverage inter-slice spatial dependences through internal memory units. RFCN combines anatomical detection and segmentation into a single architecture that is trained end-to-end thus significantly reducing computational time, simplifying the segmentation pipeline, and potentially enabling real-time applications. We report on an investigation of RFCN using two datasets, including the publicly available MICCAI 2009 Challenge dataset. Comparisons have been carried out between fully convolutional networks and deep restricted Boltzmann machines, including a recurrent version that leverages inter-slice spatial correlation. Our studies suggest that RFCN produces state-of-the-art results and can substantially improve the delineation of contours near the apex of the heart.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge