Yueyang Huang
HelixDesign-Antibody: A Scalable Production-Grade Platform for Antibody Design Built on HelixFold3
Jul 03, 2025Abstract:Antibody engineering is essential for developing therapeutics and advancing biomedical research. Traditional discovery methods often rely on time-consuming and resource-intensive experimental screening. To enhance and streamline this process, we introduce a production-grade, high-throughput platform built on HelixFold3, HelixDesign-Antibody, which utilizes the high-accuracy structure prediction model, HelixFold3. The platform facilitates the large-scale generation of antibody candidate sequences and evaluates their interaction with antigens. Integrated high-performance computing (HPC) support enables high-throughput screening, addressing challenges such as fragmented toolchains and high computational demands. Validation on multiple antigens showcases the platform's ability to generate diverse and high-quality antibodies, confirming a scaling law where exploring larger sequence spaces increases the likelihood of identifying optimal binders. This platform provides a seamless, accessible solution for large-scale antibody design and is available via the antibody design page of PaddleHelix platform.
HelixDesign-Binder: A Scalable Production-Grade Platform for Binder Design Built on HelixFold3
May 28, 2025Abstract:Protein binder design is central to therapeutics, diagnostics, and synthetic biology, yet practical deployment remains challenging due to fragmented workflows, high computational costs, and complex tool integration. We present HelixDesign-Binder, a production-grade, high-throughput platform built on HelixFold3 that automates the full binder design pipeline, from backbone generation and sequence design to structural evaluation and multi-dimensional scoring. By unifying these stages into a scalable and user-friendly system, HelixDesign-Binder enables efficient exploration of binder candidates with favorable structural, energetic, and physicochemical properties. The platform leverages Baidu Cloud's high-performance infrastructure to support large-scale design and incorporates advanced scoring metrics, including ipTM, predicted binding free energy, and interface hydrophobicity. Benchmarking across six protein targets demonstrates that HelixDesign-Binder reliably produces diverse and high-quality binders, some of which match or exceed validated designs in predicted binding affinity. HelixDesign-Binder is accessible via an interactive web interface in PaddleHelix platform, supporting both academic research and industrial applications in antibody and protein binder development.
HelixADMET: a robust and endpoint extensible ADMET system incorporating self-supervised knowledge transfer
May 17, 2022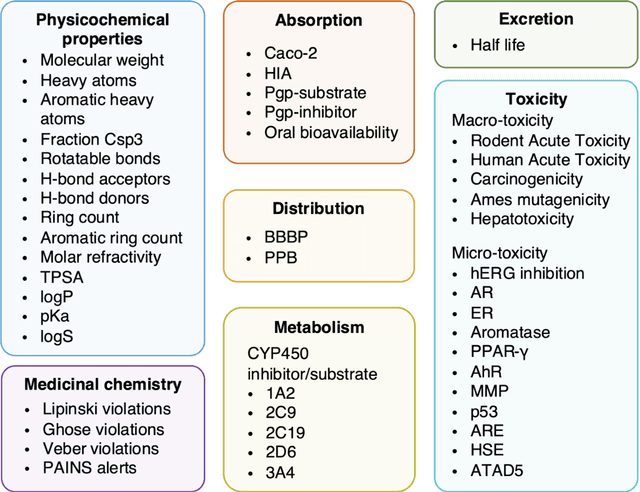
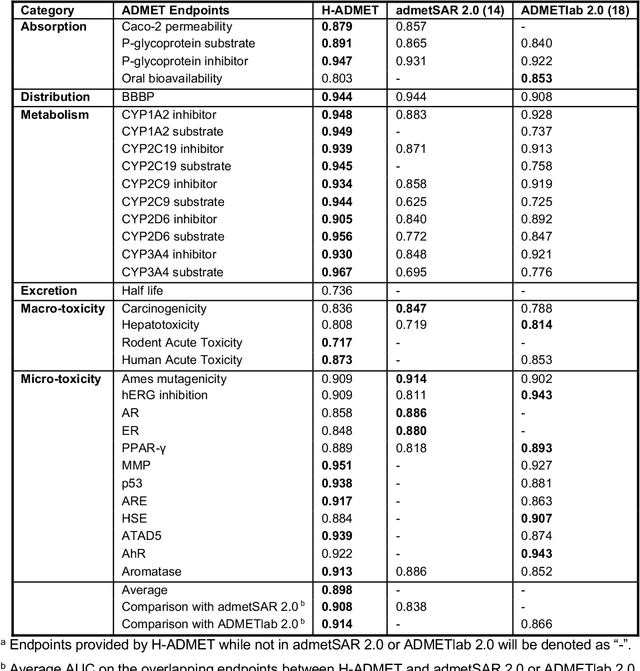
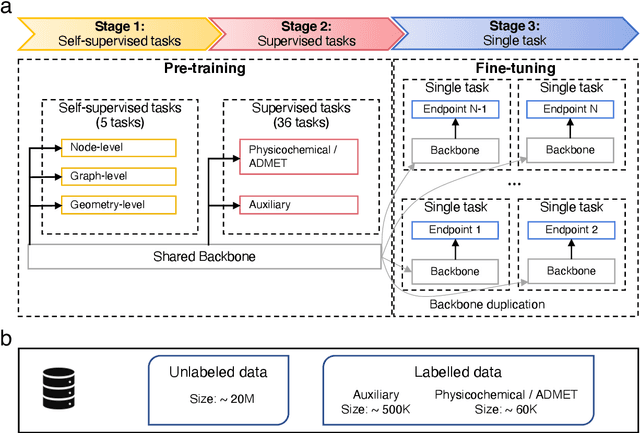
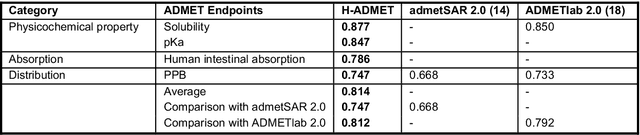
Abstract:Accurate ADMET (an abbreviation for "absorption, distribution, metabolism, excretion, and toxicity") predictions can efficiently screen out undesirable drug candidates in the early stage of drug discovery. In recent years, multiple comprehensive ADMET systems that adopt advanced machine learning models have been developed, providing services to estimate multiple endpoints. However, those ADMET systems usually suffer from weak extrapolation ability. First, due to the lack of labelled data for each endpoint, typical machine learning models perform frail for the molecules with unobserved scaffolds. Second, most systems only provide fixed built-in endpoints and cannot be customised to satisfy various research requirements. To this end, we develop a robust and endpoint extensible ADMET system, HelixADMET (H-ADMET). H-ADMET incorporates the concept of self-supervised learning to produce a robust pre-trained model. The model is then fine-tuned with a multi-task and multi-stage framework to transfer knowledge between ADMET endpoints, auxiliary tasks, and self-supervised tasks. Our results demonstrate that H-ADMET achieves an overall improvement of 4%, compared with existing ADMET systems on comparable endpoints. Additionally, the pre-trained model provided by H-ADMET can be fine-tuned to generate new and customised ADMET endpoints, meeting various demands of drug research and development requirements.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge