Yuanqi Du
AlphaNet: Scaling Up Local Frame-based Atomistic Foundation Model
Jan 13, 2025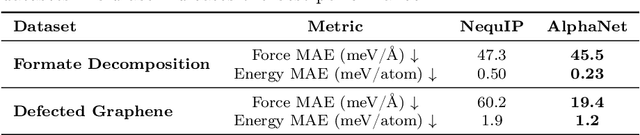
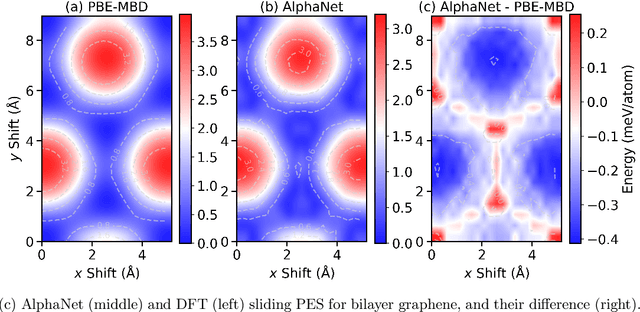
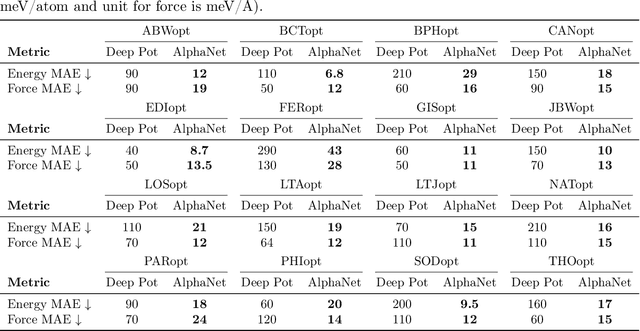
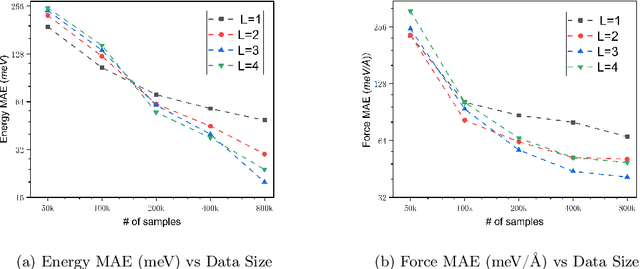
Abstract:We present AlphaNet, a local frame-based equivariant model designed to achieve both accurate and efficient simulations for atomistic systems. Recently, machine learning force fields (MLFFs) have gained prominence in molecular dynamics simulations due to their advantageous efficiency-accuracy balance compared to classical force fields and quantum mechanical calculations, alongside their transferability across various systems. Despite the advancements in improving model accuracy, the efficiency and scalability of MLFFs remain significant obstacles in practical applications. AlphaNet enhances computational efficiency and accuracy by leveraging the local geometric structures of atomic environments through the construction of equivariant local frames and learnable frame transitions. We substantiate the efficacy of AlphaNet across diverse datasets, including defected graphene, formate decomposition, zeolites, and surface reactions. AlphaNet consistently surpasses well-established models, such as NequIP and DeepPot, in terms of both energy and force prediction accuracy. Notably, AlphaNet offers one of the best trade-offs between computational efficiency and accuracy among existing models. Moreover, AlphaNet exhibits scalability across a broad spectrum of system and dataset sizes, affirming its versatility.
Graph Generative Pre-trained Transformer
Jan 02, 2025



Abstract:Graph generation is a critical task in numerous domains, including molecular design and social network analysis, due to its ability to model complex relationships and structured data. While most modern graph generative models utilize adjacency matrix representations, this work revisits an alternative approach that represents graphs as sequences of node set and edge set. We advocate for this approach due to its efficient encoding of graphs and propose a novel representation. Based on this representation, we introduce the Graph Generative Pre-trained Transformer (G2PT), an auto-regressive model that learns graph structures via next-token prediction. To further exploit G2PT's capabilities as a general-purpose foundation model, we explore fine-tuning strategies for two downstream applications: goal-oriented generation and graph property prediction. We conduct extensive experiments across multiple datasets. Results indicate that G2PT achieves superior generative performance on both generic graph and molecule datasets. Furthermore, G2PT exhibits strong adaptability and versatility in downstream tasks from molecular design to property prediction.
Generative Design of Functional Metal Complexes Utilizing the Internal Knowledge of Large Language Models
Oct 21, 2024



Abstract:Designing functional transition metal complexes (TMCs) faces challenges due to the vast search space of metals and ligands, requiring efficient optimization strategies. Traditional genetic algorithms (GAs) are commonly used, employing random mutations and crossovers driven by explicit mathematical objectives to explore this space. Transferring knowledge between different GA tasks, however, is difficult. We integrate large language models (LLMs) into the evolutionary optimization framework (LLM-EO) and apply it in both single- and multi-objective optimization for TMCs. We find that LLM-EO surpasses traditional GAs by leveraging the chemical knowledge of LLMs gained during their extensive pretraining. Remarkably, without supervised fine-tuning, LLMs utilize the full historical data from optimization processes, outperforming those focusing only on top-performing TMCs. LLM-EO successfully identifies eight of the top-20 TMCs with the largest HOMO-LUMO gaps by proposing only 200 candidates out of a 1.37 million TMCs space. Through prompt engineering using natural language, LLM-EO introduces unparalleled flexibility into multi-objective optimizations, thereby circumventing the necessity for intricate mathematical formulations. As generative models, LLMs can suggest new ligands and TMCs with unique properties by merging both internal knowledge and external chemistry data, thus combining the benefits of efficient optimization and molecular generation. With increasing potential of LLMs as pretrained foundational models and new post-training inference strategies, we foresee broad applications of LLM-based evolutionary optimization in chemistry and materials design.
Generalized Flow Matching for Transition Dynamics Modeling
Oct 19, 2024



Abstract:Simulating transition dynamics between metastable states is a fundamental challenge in dynamical systems and stochastic processes with wide real-world applications in understanding protein folding, chemical reactions and neural activities. However, the computational challenge often lies on sampling exponentially many paths in which only a small fraction ends in the target metastable state due to existence of high energy barriers. To amortize the cost, we propose a data-driven approach to warm-up the simulation by learning nonlinear interpolations from local dynamics. Specifically, we infer a potential energy function from local dynamics data. To find plausible paths between two metastable states, we formulate a generalized flow matching framework that learns a vector field to sample propable paths between the two marginal densities under the learned energy function. Furthermore, we iteratively refine the model by assigning importance weights to the sampled paths and buffering more likely paths for training. We validate the effectiveness of the proposed method to sample probable paths on both synthetic and real-world molecular systems.
Doob's Lagrangian: A Sample-Efficient Variational Approach to Transition Path Sampling
Oct 10, 2024



Abstract:Rare event sampling in dynamical systems is a fundamental problem arising in the natural sciences, which poses significant computational challenges due to an exponentially large space of trajectories. For settings where the dynamical system of interest follows a Brownian motion with known drift, the question of conditioning the process to reach a given endpoint or desired rare event is definitively answered by Doob's h-transform. However, the naive estimation of this transform is infeasible, as it requires simulating sufficiently many forward trajectories to estimate rare event probabilities. In this work, we propose a variational formulation of Doob's $h$-transform as an optimization problem over trajectories between a given initial point and the desired ending point. To solve this optimization, we propose a simulation-free training objective with a model parameterization that imposes the desired boundary conditions by design. Our approach significantly reduces the search space over trajectories and avoids expensive trajectory simulation and inefficient importance sampling estimators which are required in existing methods. We demonstrate the ability of our method to find feasible transition paths on real-world molecular simulation and protein folding tasks.
Efficient Evolutionary Search Over Chemical Space with Large Language Models
Jun 23, 2024



Abstract:Molecular discovery, when formulated as an optimization problem, presents significant computational challenges because optimization objectives can be non-differentiable. Evolutionary Algorithms (EAs), often used to optimize black-box objectives in molecular discovery, traverse chemical space by performing random mutations and crossovers, leading to a large number of expensive objective evaluations. In this work, we ameliorate this shortcoming by incorporating chemistry-aware Large Language Models (LLMs) into EAs. Namely, we redesign crossover and mutation operations in EAs using LLMs trained on large corpora of chemical information. We perform extensive empirical studies on both commercial and open-source models on multiple tasks involving property optimization, molecular rediscovery, and structure-based drug design, demonstrating that the joint usage of LLMs with EAs yields superior performance over all baseline models across single- and multi-objective settings. We demonstrate that our algorithm improves both the quality of the final solution and convergence speed, thereby reducing the number of required objective evaluations. Our code is available at http://github.com/zoom-wang112358/MOLLEO
Navigating Chemical Space with Latent Flows
May 08, 2024



Abstract:Recent progress of deep generative models in the vision and language domain has stimulated significant interest in more structured data generation such as molecules. However, beyond generating new random molecules, efficient exploration and a comprehensive understanding of the vast chemical space are of great importance to molecular science and applications in drug design and materials discovery. In this paper, we propose a new framework, ChemFlow, to traverse chemical space through navigating the latent space learned by molecule generative models through flows. We introduce a dynamical system perspective that formulates the problem as learning a vector field that transports the mass of the molecular distribution to the region with desired molecular properties or structure diversity. Under this framework, we unify previous approaches on molecule latent space traversal and optimization and propose alternative competing methods incorporating different physical priors. We validate the efficacy of ChemFlow on molecule manipulation and single- and multi-objective molecule optimization tasks under both supervised and unsupervised molecular discovery settings. Codes and demos are publicly available on GitHub at https://github.com/garywei944/ChemFlow.
React-OT: Optimal Transport for Generating Transition State in Chemical Reactions
Apr 20, 2024



Abstract:Transition states (TSs) are transient structures that are key in understanding reaction mechanisms and designing catalysts but challenging to be captured in experiments. Alternatively, many optimization algorithms have been developed to search for TSs computationally. Yet the cost of these algorithms driven by quantum chemistry methods (usually density functional theory) is still high, posing challenges for their applications in building large reaction networks for reaction exploration. Here we developed React-OT, an optimal transport approach for generating unique TS structures from reactants and products. React-OT generates highly accurate TS structures with a median structural root mean square deviation (RMSD) of 0.053{\AA} and median barrier height error of 1.06 kcal/mol requiring only 0.4 second per reaction. The RMSD and barrier height error is further improved by roughly 25% through pretraining React-OT on a large reaction dataset obtained with a lower level of theory, GFN2-xTB. We envision the great accuracy and fast inference of React-OT useful in targeting TSs when exploring chemical reactions with unknown mechanisms.
Diffusion Models as Constrained Samplers for Optimization with Unknown Constraints
Feb 28, 2024



Abstract:Addressing real-world optimization problems becomes particularly challenging when analytic objective functions or constraints are unavailable. While numerous studies have addressed the issue of unknown objectives, limited research has focused on scenarios where feasibility constraints are not given explicitly. Overlooking these constraints can lead to spurious solutions that are unrealistic in practice. To deal with such unknown constraints, we propose to perform optimization within the data manifold using diffusion models. To constrain the optimization process to the data manifold, we reformulate the original optimization problem as a sampling problem from the product of the Boltzmann distribution defined by the objective function and the data distribution learned by the diffusion model. To enhance sampling efficiency, we propose a two-stage framework that begins with a guided diffusion process for warm-up, followed by a Langevin dynamics stage for further correction. Theoretical analysis shows that the initial stage results in a distribution focused on feasible solutions, thereby providing a better initialization for the later stage. Comprehensive experiments on a synthetic dataset, six real-world black-box optimization datasets, and a multi-objective optimization dataset show that our method achieves better or comparable performance with previous state-of-the-art baselines.
Learning Over Molecular Conformer Ensembles: Datasets and Benchmarks
Sep 29, 2023



Abstract:Molecular Representation Learning (MRL) has proven impactful in numerous biochemical applications such as drug discovery and enzyme design. While Graph Neural Networks (GNNs) are effective at learning molecular representations from a 2D molecular graph or a single 3D structure, existing works often overlook the flexible nature of molecules, which continuously interconvert across conformations via chemical bond rotations and minor vibrational perturbations. To better account for molecular flexibility, some recent works formulate MRL as an ensemble learning problem, focusing on explicitly learning from a set of conformer structures. However, most of these studies have limited datasets, tasks, and models. In this work, we introduce the first MoleculAR Conformer Ensemble Learning (MARCEL) benchmark to thoroughly evaluate the potential of learning on conformer ensembles and suggest promising research directions. MARCEL includes four datasets covering diverse molecule- and reaction-level properties of chemically diverse molecules including organocatalysts and transition-metal catalysts, extending beyond the scope of common GNN benchmarks that are confined to drug-like molecules. In addition, we conduct a comprehensive empirical study, which benchmarks representative 1D, 2D, and 3D molecular representation learning models, along with two strategies that explicitly incorporate conformer ensembles into 3D MRL models. Our findings reveal that direct learning from an accessible conformer space can improve performance on a variety of tasks and models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge