Trinh Thi Le Vuong
FALFormer: Feature-aware Landmarks self-attention for Whole-slide Image Classification
Jul 11, 2024Abstract:Slide-level classification for whole-slide images (WSIs) has been widely recognized as a crucial problem in digital and computational pathology. Current approaches commonly consider WSIs as a bag of cropped patches and process them via multiple instance learning due to the large number of patches, which cannot fully explore the relationship among patches; in other words, the global information cannot be fully incorporated into decision making. Herein, we propose an efficient and effective slide-level classification model, named as FALFormer, that can process a WSI as a whole so as to fully exploit the relationship among the entire patches and to improve the classification performance. FALFormer is built based upon Transformers and self-attention mechanism. To lessen the computational burden of the original self-attention mechanism and to process the entire patches together in a WSI, FALFormer employs Nystr\"om self-attention which approximates the computation by using a smaller number of tokens or landmarks. For effective learning, FALFormer introduces feature-aware landmarks to enhance the representation power of the landmarks and the quality of the approximation. We systematically evaluate the performance of FALFormer using two public datasets, including CAMELYON16 and TCGA-BRCA. The experimental results demonstrate that FALFormer achieves superior performance on both datasets, outperforming the state-of-the-art methods for the slide-level classification. This suggests that FALFormer can facilitate an accurate and precise analysis of WSIs, potentially leading to improved diagnosis and prognosis on WSIs.
Towards a text-based quantitative and explainable histopathology image analysis
Jul 10, 2024Abstract:Recently, vision-language pre-trained models have emerged in computational pathology. Previous works generally focused on the alignment of image-text pairs via the contrastive pre-training paradigm. Such pre-trained models have been applied to pathology image classification in zero-shot learning or transfer learning fashion. Herein, we hypothesize that the pre-trained vision-language models can be utilized for quantitative histopathology image analysis through a simple image-to-text retrieval. To this end, we propose a Text-based Quantitative and Explainable histopathology image analysis, which we call TQx. Given a set of histopathology images, we adopt a pre-trained vision-language model to retrieve a word-of-interest pool. The retrieved words are then used to quantify the histopathology images and generate understandable feature embeddings due to the direct mapping to the text description. To evaluate the proposed method, the text-based embeddings of four histopathology image datasets are utilized to perform clustering and classification tasks. The results demonstrate that TQx is able to quantify and analyze histopathology images that are comparable to the prevalent visual models in computational pathology.
MoMA: Momentum Contrastive Learning with Multi-head Attention-based Knowledge Distillation for Histopathology Image Analysis
Aug 31, 2023Abstract:There is no doubt that advanced artificial intelligence models and high quality data are the keys to success in developing computational pathology tools. Although the overall volume of pathology data keeps increasing, a lack of quality data is a common issue when it comes to a specific task due to several reasons including privacy and ethical issues with patient data. In this work, we propose to exploit knowledge distillation, i.e., utilize the existing model to learn a new, target model, to overcome such issues in computational pathology. Specifically, we employ a student-teacher framework to learn a target model from a pre-trained, teacher model without direct access to source data and distill relevant knowledge via momentum contrastive learning with multi-head attention mechanism, which provides consistent and context-aware feature representations. This enables the target model to assimilate informative representations of the teacher model while seamlessly adapting to the unique nuances of the target data. The proposed method is rigorously evaluated across different scenarios where the teacher model was trained on the same, relevant, and irrelevant classification tasks with the target model. Experimental results demonstrate the accuracy and robustness of our approach in transferring knowledge to different domains and tasks, outperforming other related methods. Moreover, the results provide a guideline on the learning strategy for different types of tasks and scenarios in computational pathology. Code is available at: \url{https://github.com/trinhvg/MoMA}.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
IMPaSh: A Novel Domain-shift Resistant Representation for Colorectal Cancer Tissue Classification
Aug 23, 2022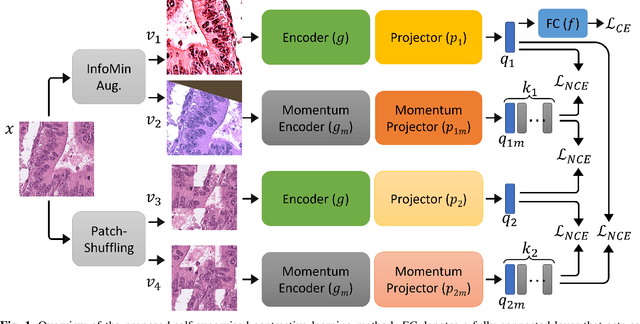
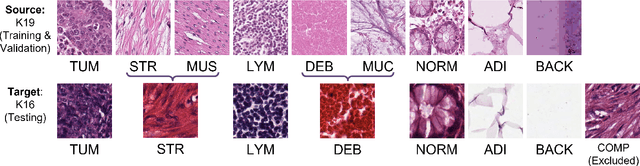
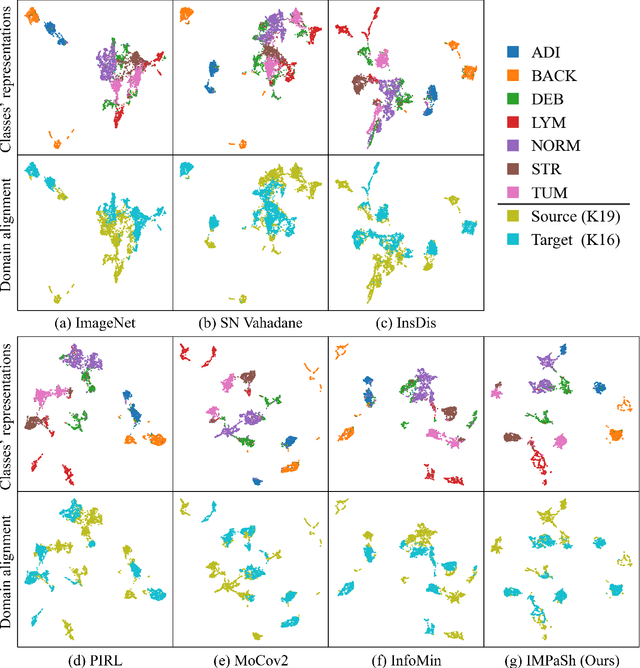
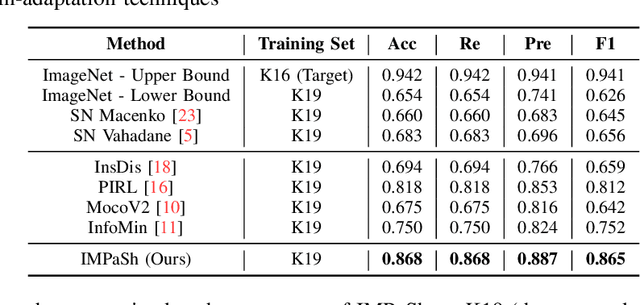
Abstract:The appearance of histopathology images depends on tissue type, staining and digitization procedure. These vary from source to source and are the potential causes for domain-shift problems. Owing to this problem, despite the great success of deep learning models in computational pathology, a model trained on a specific domain may still perform sub-optimally when we apply them to another domain. To overcome this, we propose a new augmentation called PatchShuffling and a novel self-supervised contrastive learning framework named IMPaSh for pre-training deep learning models. Using these, we obtained a ResNet50 encoder that can extract image representation resistant to domain-shift. We compared our derived representation against those acquired based on other domain-generalization techniques by using them for the cross-domain classification of colorectal tissue images. We show that the proposed method outperforms other traditional histology domain-adaptation and state-of-the-art self-supervised learning methods. Code is available at: https://github.com/trinhvg/IMPash .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge