Thomas Moreau
PARIETAL
RoseCDL: Robust and Scalable Convolutional Dictionary Learning for Rare-event Detection
Sep 10, 2025Abstract:Identifying recurring patterns and rare events in large-scale signals is a fundamental challenge in fields such as astronomy, physical simulations, and biomedical science. Convolutional Dictionary Learning (CDL) offers a powerful framework for modeling local structures in signals, but its use for detecting rare or anomalous events remains largely unexplored. In particular, CDL faces two key challenges in this setting: high computational cost and sensitivity to artifacts and outliers. In this paper, we introduce RoseCDL, a scalable and robust CDL algorithm designed for unsupervised rare event detection in long signals. RoseCDL combines stochastic windowing for efficient training on large datasets with inline outlier detection to enhance robustness and isolate anomalous patterns. This reframes CDL as a practical tool for event discovery and characterization in real-world signals, extending its role beyond traditional tasks like compression or denoising.
DeepInverse: A Python package for solving imaging inverse problems with deep learning
May 26, 2025Abstract:DeepInverse is an open-source PyTorch-based library for solving imaging inverse problems. The library covers all crucial steps in image reconstruction from the efficient implementation of forward operators (e.g., optics, MRI, tomography), to the definition and resolution of variational problems and the design and training of advanced neural network architectures. In this paper, we describe the main functionality of the library and discuss the main design choices.
From Denoising Score Matching to Langevin Sampling: A Fine-Grained Error Analysis in the Gaussian Setting
Mar 14, 2025Abstract:Sampling from an unknown distribution, accessible only through discrete samples, is a fundamental problem at the core of generative AI. The current state-of-the-art methods follow a two-step process: first estimating the score function (the gradient of a smoothed log-distribution) and then applying a gradient-based sampling algorithm. The resulting distribution's correctness can be impacted by several factors: the generalization error due to a finite number of initial samples, the error in score matching, and the diffusion error introduced by the sampling algorithm. In this paper, we analyze the sampling process in a simple yet representative setting-sampling from Gaussian distributions using a Langevin diffusion sampler. We provide a sharp analysis of the Wasserstein sampling error that arises from the multiple sources of error throughout the pipeline. This allows us to rigorously track how the anisotropy of the data distribution (encoded by its power spectrum) interacts with key parameters of the end-to-end sampling method, including the noise amplitude, the step sizes in both score matching and diffusion, and the number of initial samples. Notably, we show that the Wasserstein sampling error can be expressed as a kernel-type norm of the data power spectrum, where the specific kernel depends on the method parameters. This result provides a foundation for further analysis of the tradeoffs involved in optimizing sampling accuracy, such as adapting the noise amplitude to the choice of step sizes.
FiRe: Fixed-points of Restoration Priors for Solving Inverse Problems
Nov 28, 2024Abstract:Selecting an appropriate prior to compensate for information loss due to the measurement operator is a fundamental challenge in imaging inverse problems. Implicit priors based on denoising neural networks have become central to widely-used frameworks such as Plug-and-Play (PnP) algorithms. In this work, we introduce Fixed-points of Restoration (FiRe) priors as a new framework for expanding the notion of priors in PnP to general restoration models beyond traditional denoising models. The key insight behind FiRe is that natural images emerge as fixed points of the composition of a degradation operator with the corresponding restoration model. This enables us to derive an explicit formula for our implicit prior by quantifying invariance of images under this composite operation. Adopting this fixed-point perspective, we show how various restoration networks can effectively serve as priors for solving inverse problems. The FiRe framework further enables ensemble-like combinations of multiple restoration models as well as acquisition-informed restoration networks, all within a unified optimization approach. Experimental results validate the effectiveness of FiRe across various inverse problems, establishing a new paradigm for incorporating pretrained restoration models into PnP-like algorithms.
sbi reloaded: a toolkit for simulation-based inference workflows
Nov 26, 2024
Abstract:Scientists and engineers use simulators to model empirically observed phenomena. However, tuning the parameters of a simulator to ensure its outputs match observed data presents a significant challenge. Simulation-based inference (SBI) addresses this by enabling Bayesian inference for simulators, identifying parameters that match observed data and align with prior knowledge. Unlike traditional Bayesian inference, SBI only needs access to simulations from the model and does not require evaluations of the likelihood-function. In addition, SBI algorithms do not require gradients through the simulator, allow for massive parallelization of simulations, and can perform inference for different observations without further simulations or training, thereby amortizing inference. Over the past years, we have developed, maintained, and extended $\texttt{sbi}$, a PyTorch-based package that implements Bayesian SBI algorithms based on neural networks. The $\texttt{sbi}$ toolkit implements a wide range of inference methods, neural network architectures, sampling methods, and diagnostic tools. In addition, it provides well-tested default settings but also offers flexibility to fully customize every step of the simulation-based inference workflow. Taken together, the $\texttt{sbi}$ toolkit enables scientists and engineers to apply state-of-the-art SBI methods to black-box simulators, opening up new possibilities for aligning simulations with empirically observed data.
Analysis and Synthesis Denoisers for Forward-Backward Plug-and-Play Algorithms
Nov 20, 2024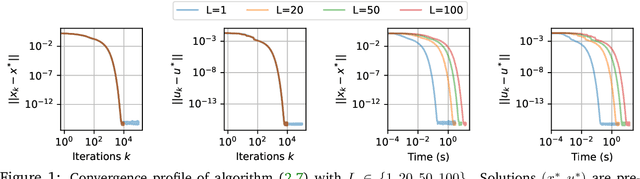
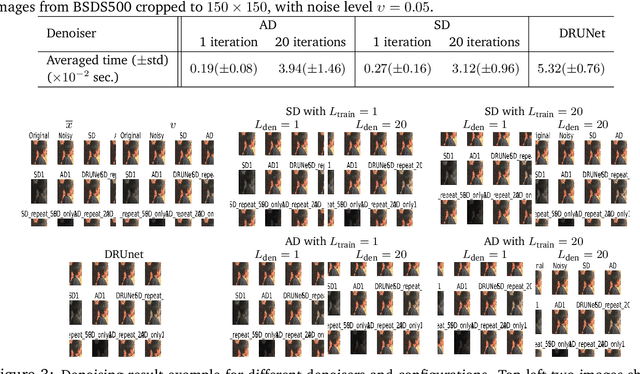
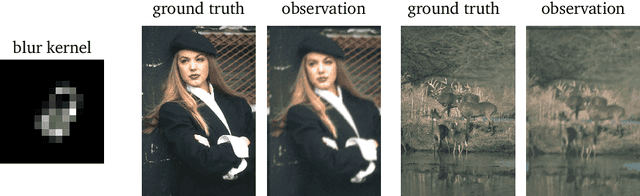

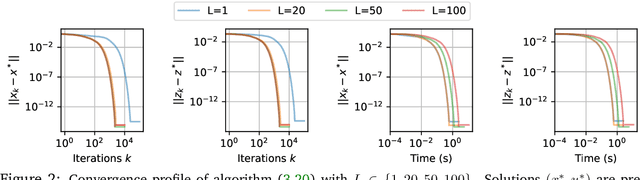

Abstract:In this work we study the behavior of the forward-backward (FB) algorithm when the proximity operator is replaced by a sub-iterative procedure to approximate a Gaussian denoiser, in a Plug-and-Play (PnP) fashion. In particular, we consider both analysis and synthesis Gaussian denoisers within a dictionary framework, obtained by unrolling dual-FB iterations or FB iterations, respectively. We analyze the associated minimization problems as well as the asymptotic behavior of the resulting FB-PnP iterations. In particular, we show that the synthesis Gaussian denoising problem can be viewed as a proximity operator. For each case, analysis and synthesis, we show that the FB-PnP algorithms solve the same problem whether we use only one or an infinite number of sub-iteration to solve the denoising problem at each iteration. To this aim, we show that each "one sub-iteration" strategy within the FB-PnP can be interpreted as a primal-dual algorithm when a warm-restart strategy is used. We further present similar results when using a Moreau-Yosida smoothing of the global problem, for an arbitrary number of sub-iterations. Finally, we provide numerical simulations to illustrate our theoretical results. In particular we first consider a toy compressive sensing example, as well as an image restoration problem in a deep dictionary framework.
SKADA-Bench: Benchmarking Unsupervised Domain Adaptation Methods with Realistic Validation
Jul 16, 2024



Abstract:Unsupervised Domain Adaptation (DA) consists of adapting a model trained on a labeled source domain to perform well on an unlabeled target domain with some data distribution shift. While many methods have been proposed in the literature, fair and realistic evaluation remains an open question, particularly due to methodological difficulties in selecting hyperparameters in the unsupervised setting. With SKADA-Bench, we propose a framework to evaluate DA methods and present a fair evaluation of existing shallow algorithms, including reweighting, mapping, and subspace alignment. Realistic hyperparameter selection is performed with nested cross-validation and various unsupervised model selection scores, on both simulated datasets with controlled shifts and real-world datasets across diverse modalities, such as images, text, biomedical, and tabular data with specific feature extraction. Our benchmark highlights the importance of realistic validation and provides practical guidance for real-life applications, with key insights into the choice and impact of model selection approaches. SKADA-Bench is open-source, reproducible, and can be easily extended with novel DA methods, datasets, and model selection criteria without requiring re-evaluating competitors. SKADA-Bench is available on GitHub at https://github.com/scikit-adaptation/skada-bench.
Unmixing Noise from Hawkes Process to Model Learned Physiological Events
Jun 17, 2024



Abstract:Physiological signal analysis often involves identifying events crucial to understanding biological dynamics. Traditional methods rely on handcrafted procedures or supervised learning, presenting challenges such as expert dependence, lack of robustness, and the need for extensive labeled data. Data-driven methods like Convolutional Dictionary Learning (CDL) offer an alternative but tend to produce spurious detections. This work introduces UNHaP (Unmix Noise from Hawkes Processes), a novel approach addressing the joint learning of temporal structures in events and the removal of spurious detections. Leveraging marked Hawkes processes, UNHaP distinguishes between events of interest and spurious ones. By treating the event detection output as a mixture of structured and unstructured events, UNHaP efficiently unmixes these processes and estimates their parameters. This approach significantly enhances the understanding of event distributions while minimizing false detection rates.
Flexible Parametric Inference for Space-Time Hawkes Processes
Jun 10, 2024



Abstract:Many modern spatio-temporal data sets, in sociology, epidemiology or seismology, for example, exhibit self-exciting characteristics, triggering and clustering behaviors both at the same time, that a suitable Hawkes space-time process can accurately capture. This paper aims to develop a fast and flexible parametric inference technique to recover the parameters of the kernel functions involved in the intensity function of a space-time Hawkes process based on such data. Our statistical approach combines three key ingredients: 1) kernels with finite support are considered, 2) the space-time domain is appropriately discretized, and 3) (approximate) precomputations are used. The inference technique we propose then consists of a $\ell_2$ gradient-based solver that is fast and statistically accurate. In addition to describing the algorithmic aspects, numerical experiments have been carried out on synthetic and real spatio-temporal data, providing solid empirical evidence of the relevance of the proposed methodology.
S-JEPA: towards seamless cross-dataset transfer through dynamic spatial attention
Mar 18, 2024



Abstract:Motivated by the challenge of seamless cross-dataset transfer in EEG signal processing, this article presents an exploratory study on the use of Joint Embedding Predictive Architectures (JEPAs). In recent years, self-supervised learning has emerged as a promising approach for transfer learning in various domains. However, its application to EEG signals remains largely unexplored. In this article, we introduce Signal-JEPA for representing EEG recordings which includes a novel domain-specific spatial block masking strategy and three novel architectures for downstream classification. The study is conducted on a 54~subjects dataset and the downstream performance of the models is evaluated on three different BCI paradigms: motor imagery, ERP and SSVEP. Our study provides preliminary evidence for the potential of JEPAs in EEG signal encoding. Notably, our results highlight the importance of spatial filtering for accurate downstream classification and reveal an influence of the length of the pre-training examples but not of the mask size on the downstream performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge