Alexandre Gramfort
MIND - INRIA
EEG Foundation Challenge: From Cross-Task to Cross-Subject EEG Decoding
Jun 23, 2025Abstract:Current electroencephalogram (EEG) decoding models are typically trained on small numbers of subjects performing a single task. Here, we introduce a large-scale, code-submission-based competition comprising two challenges. First, the Transfer Challenge asks participants to build and test a model that can zero-shot decode new tasks and new subjects from their EEG data. Second, the Psychopathology factor prediction Challenge asks participants to infer subject measures of mental health from EEG data. For this, we use an unprecedented, multi-terabyte dataset of high-density EEG signals (128 channels) recorded from over 3,000 child to young adult subjects engaged in multiple active and passive tasks. We provide several tunable neural network baselines for each of these two challenges, including a simple network and demographic-based regression models. Developing models that generalise across tasks and individuals will pave the way for ML network architectures capable of adapting to EEG data collected from diverse tasks and individuals. Similarly, predicting mental health-relevant personality trait values from EEG might identify objective biomarkers useful for clinical diagnosis and design of personalised treatment for psychological conditions. Ultimately, the advances spurred by this challenge could contribute to the development of computational psychiatry and useful neurotechnology, and contribute to breakthroughs in both fundamental neuroscience and applied clinical research.
* Approved at Neurips Competition track. webpage: https://eeg2025.github.io/
PSDNorm: Test-Time Temporal Normalization for Deep Learning on EEG Signals
Mar 06, 2025



Abstract:Distribution shift poses a significant challenge in machine learning, particularly in biomedical applications such as EEG signals collected across different subjects, institutions, and recording devices. While existing normalization layers, Batch-Norm, LayerNorm and InstanceNorm, help address distribution shifts, they fail to capture the temporal dependencies inherent in temporal signals. In this paper, we propose PSDNorm, a layer that leverages Monge mapping and temporal context to normalize feature maps in deep learning models. Notably, the proposed method operates as a test-time domain adaptation technique, addressing distribution shifts without additional training. Evaluations on 10 sleep staging datasets using the U-Time model demonstrate that PSDNorm achieves state-of-the-art performance at test time on datasets not seen during training while being 4x more data-efficient than the best baseline. Additionally, PSDNorm provides a significant improvement in robustness, achieving markedly higher F1 scores for the 20% hardest subjects.
Identifiable Multi-View Causal Discovery Without Non-Gaussianity
Feb 28, 2025
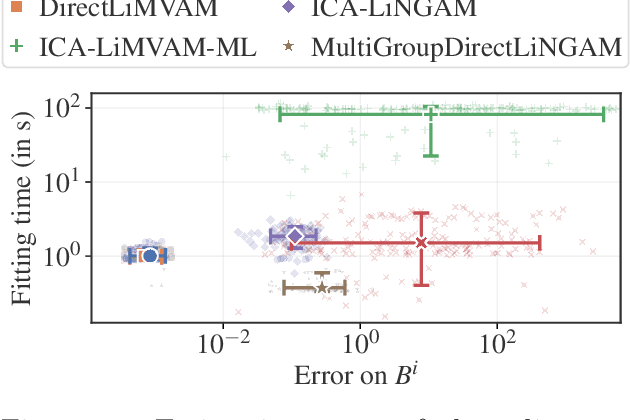


Abstract:We propose a novel approach to linear causal discovery in the framework of multi-view Structural Equation Models (SEM). Our proposed model relaxes the well-known assumption of non-Gaussian disturbances by alternatively assuming diversity of variances over views, making it more broadly applicable. We prove the identifiability of all the parameters of the model without any further assumptions on the structure of the SEM other than it being acyclic. We further propose an estimation algorithm based on recent advances in multi-view Independent Component Analysis (ICA). The proposed methodology is validated through simulations and application on real neuroimaging data, where it enables the estimation of causal graphs between brain regions.
MVICAD2: Multi-View Independent Component Analysis with Delays and Dilations
Jan 13, 2025



Abstract:Machine learning techniques in multi-view settings face significant challenges, particularly when integrating heterogeneous data, aligning feature spaces, and managing view-specific biases. These issues are prominent in neuroscience, where data from multiple subjects exposed to the same stimuli are analyzed to uncover brain activity dynamics. In magnetoencephalography (MEG), where signals are captured at the scalp level, estimating the brain's underlying sources is crucial, especially in group studies where sources are assumed to be similar for all subjects. Common methods, such as Multi-View Independent Component Analysis (MVICA), assume identical sources across subjects, but this assumption is often too restrictive due to individual variability and age-related changes. Multi-View Independent Component Analysis with Delays (MVICAD) addresses this by allowing sources to differ up to a temporal delay. However, temporal dilation effects, particularly in auditory stimuli, are common in brain dynamics, making the estimation of time delays alone insufficient. To address this, we propose Multi-View Independent Component Analysis with Delays and Dilations (MVICAD2), which allows sources to differ across subjects in both temporal delays and dilations. We present a model with identifiable sources, derive an approximation of its likelihood in closed form, and use regularization and optimization techniques to enhance performance. Through simulations, we demonstrate that MVICAD2 outperforms existing multi-view ICA methods. We further validate its effectiveness using the Cam-CAN dataset, and showing how delays and dilations are related to aging.
emg2qwerty: A Large Dataset with Baselines for Touch Typing using Surface Electromyography
Oct 26, 2024



Abstract:Surface electromyography (sEMG) non-invasively measures signals generated by muscle activity with sufficient sensitivity to detect individual spinal neurons and richness to identify dozens of gestures and their nuances. Wearable wrist-based sEMG sensors have the potential to offer low friction, subtle, information rich, always available human-computer inputs. To this end, we introduce emg2qwerty, a large-scale dataset of non-invasive electromyographic signals recorded at the wrists while touch typing on a QWERTY keyboard, together with ground-truth annotations and reproducible baselines. With 1,135 sessions spanning 108 users and 346 hours of recording, this is the largest such public dataset to date. These data demonstrate non-trivial, but well defined hierarchical relationships both in terms of the generative process, from neurons to muscles and muscle combinations, as well as in terms of domain shift across users and user sessions. Applying standard modeling techniques from the closely related field of Automatic Speech Recognition (ASR), we show strong baseline performance on predicting key-presses using sEMG signals alone. We believe the richness of this task and dataset will facilitate progress in several problems of interest to both the machine learning and neuroscientific communities. Dataset and code can be accessed at https://github.com/facebookresearch/emg2qwerty.
Multi-Source and Test-Time Domain Adaptation on Multivariate Signals using Spatio-Temporal Monge Alignment
Jul 19, 2024Abstract:Machine learning applications on signals such as computer vision or biomedical data often face significant challenges due to the variability that exists across hardware devices or session recordings. This variability poses a Domain Adaptation (DA) problem, as training and testing data distributions often differ. In this work, we propose Spatio-Temporal Monge Alignment (STMA) to mitigate these variabilities. This Optimal Transport (OT) based method adapts the cross-power spectrum density (cross-PSD) of multivariate signals by mapping them to the Wasserstein barycenter of source domains (multi-source DA). Predictions for new domains can be done with a filtering without the need for retraining a model with source data (test-time DA). We also study and discuss two special cases of the method, Temporal Monge Alignment (TMA) and Spatial Monge Alignment (SMA). Non-asymptotic concentration bounds are derived for the mappings estimation, which reveals a bias-plus-variance error structure with a variance decay rate of $\mathcal{O}(n_\ell^{-1/2})$ with $n_\ell$ the signal length. This theoretical guarantee demonstrates the efficiency of the proposed computational schema. Numerical experiments on multivariate biosignals and image data show that STMA leads to significant and consistent performance gains between datasets acquired with very different settings. Notably, STMA is a pre-processing step complementary to state-of-the-art deep learning methods.
SKADA-Bench: Benchmarking Unsupervised Domain Adaptation Methods with Realistic Validation
Jul 16, 2024



Abstract:Unsupervised Domain Adaptation (DA) consists of adapting a model trained on a labeled source domain to perform well on an unlabeled target domain with some data distribution shift. While many methods have been proposed in the literature, fair and realistic evaluation remains an open question, particularly due to methodological difficulties in selecting hyperparameters in the unsupervised setting. With SKADA-Bench, we propose a framework to evaluate DA methods and present a fair evaluation of existing shallow algorithms, including reweighting, mapping, and subspace alignment. Realistic hyperparameter selection is performed with nested cross-validation and various unsupervised model selection scores, on both simulated datasets with controlled shifts and real-world datasets across diverse modalities, such as images, text, biomedical, and tabular data with specific feature extraction. Our benchmark highlights the importance of realistic validation and provides practical guidance for real-life applications, with key insights into the choice and impact of model selection approaches. SKADA-Bench is open-source, reproducible, and can be easily extended with novel DA methods, datasets, and model selection criteria without requiring re-evaluating competitors. SKADA-Bench is available on GitHub at https://github.com/scikit-adaptation/skada-bench.
Geodesic Optimization for Predictive Shift Adaptation on EEG data
Jul 04, 2024



Abstract:Electroencephalography (EEG) data is often collected from diverse contexts involving different populations and EEG devices. This variability can induce distribution shifts in the data $X$ and in the biomedical variables of interest $y$, thus limiting the application of supervised machine learning (ML) algorithms. While domain adaptation (DA) methods have been developed to mitigate the impact of these shifts, such methods struggle when distribution shifts occur simultaneously in $X$ and $y$. As state-of-the-art ML models for EEG represent the data by spatial covariance matrices, which lie on the Riemannian manifold of Symmetric Positive Definite (SPD) matrices, it is appealing to study DA techniques operating on the SPD manifold. This paper proposes a novel method termed Geodesic Optimization for Predictive Shift Adaptation (GOPSA) to address test-time multi-source DA for situations in which source domains have distinct $y$ distributions. GOPSA exploits the geodesic structure of the Riemannian manifold to jointly learn a domain-specific re-centering operator representing site-specific intercepts and the regression model. We performed empirical benchmarks on the cross-site generalization of age-prediction models with resting-state EEG data from a large multi-national dataset (HarMNqEEG), which included $14$ recording sites and more than $1500$ human participants. Compared to state-of-the-art methods, our results showed that GOPSA achieved significantly higher performance on three regression metrics ($R^2$, MAE, and Spearman's $\rho$) for several source-target site combinations, highlighting its effectiveness in tackling multi-source DA with predictive shifts in EEG data analysis. Our method has the potential to combine the advantages of mixed-effects modeling with machine learning for biomedical applications of EEG, such as multicenter clinical trials.
Diffusion posterior sampling for simulation-based inference in tall data settings
Apr 11, 2024Abstract:Determining which parameters of a non-linear model could best describe a set of experimental data is a fundamental problem in science and it has gained much traction lately with the rise of complex large-scale simulators (a.k.a. black-box simulators). The likelihood of such models is typically intractable, which is why classical MCMC methods can not be used. Simulation-based inference (SBI) stands out in this context by only requiring a dataset of simulations to train deep generative models capable of approximating the posterior distribution that relates input parameters to a given observation. In this work, we consider a tall data extension in which multiple observations are available and one wishes to leverage their shared information to better infer the parameters of the model. The method we propose is built upon recent developments from the flourishing score-based diffusion literature and allows us to estimate the tall data posterior distribution simply using information from the score network trained on individual observations. We compare our method to recently proposed competing approaches on various numerical experiments and demonstrate its superiority in terms of numerical stability and computational cost.
Weakly supervised covariance matrices alignment through Stiefel matrices estimation for MEG applications
Jan 24, 2024Abstract:This paper introduces a novel domain adaptation technique for time series data, called Mixing model Stiefel Adaptation (MSA), specifically addressing the challenge of limited labeled signals in the target dataset. Leveraging a domain-dependent mixing model and the optimal transport domain adaptation assumption, we exploit abundant unlabeled data in the target domain to ensure effective prediction by establishing pairwise correspondence with equivalent signal variances between domains. Theoretical foundations are laid for identifying crucial Stiefel matrices, essential for recovering underlying signal variances from a Riemannian representation of observed signal covariances. We propose an integrated cost function that simultaneously learns these matrices, pairwise domain relationships, and a predictor, classifier, or regressor, depending on the task. Applied to neuroscience problems, MSA outperforms recent methods in brain-age regression with task variations using magnetoencephalography (MEG) signals from the Cam-CAN dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge