Stephanie L. Hyland
NOVA: An Agentic Framework for Automated Histopathology Analysis and Discovery
Nov 14, 2025



Abstract:Digitized histopathology analysis involves complex, time-intensive workflows and specialized expertise, limiting its accessibility. We introduce NOVA, an agentic framework that translates scientific queries into executable analysis pipelines by iteratively generating and running Python code. NOVA integrates 49 domain-specific tools (e.g., nuclei segmentation, whole-slide encoding) built on open-source software, and can also create new tools ad hoc. To evaluate such systems, we present SlideQuest, a 90-question benchmark -- verified by pathologists and biomedical scientists -- spanning data processing, quantitative analysis, and hypothesis testing. Unlike prior biomedical benchmarks focused on knowledge recall or diagnostic QA, SlideQuest demands multi-step reasoning, iterative coding, and computational problem solving. Quantitative evaluation shows NOVA outperforms coding-agent baselines, and a pathologist-verified case study links morphology to prognostically relevant PAM50 subtypes, demonstrating its scalable discovery potential.
MAIRA-Seg: Enhancing Radiology Report Generation with Segmentation-Aware Multimodal Large Language Models
Nov 18, 2024



Abstract:There is growing interest in applying AI to radiology report generation, particularly for chest X-rays (CXRs). This paper investigates whether incorporating pixel-level information through segmentation masks can improve fine-grained image interpretation of multimodal large language models (MLLMs) for radiology report generation. We introduce MAIRA-Seg, a segmentation-aware MLLM framework designed to utilize semantic segmentation masks alongside CXRs for generating radiology reports. We train expert segmentation models to obtain mask pseudolabels for radiology-specific structures in CXRs. Subsequently, building on the architectures of MAIRA, a CXR-specialised model for report generation, we integrate a trainable segmentation tokens extractor that leverages these mask pseudolabels, and employ mask-aware prompting to generate draft radiology reports. Our experiments on the publicly available MIMIC-CXR dataset show that MAIRA-Seg outperforms non-segmentation baselines. We also investigate set-of-marks prompting with MAIRA and find that MAIRA-Seg consistently demonstrates comparable or superior performance. The results confirm that using segmentation masks enhances the nuanced reasoning of MLLMs, potentially contributing to better clinical outcomes.
PadChest-GR: A Bilingual Chest X-ray Dataset for Grounded Radiology Report Generation
Nov 07, 2024



Abstract:Radiology report generation (RRG) aims to create free-text radiology reports from clinical imaging. Grounded radiology report generation (GRRG) extends RRG by including the localisation of individual findings on the image. Currently, there are no manually annotated chest X-ray (CXR) datasets to train GRRG models. In this work, we present a dataset called PadChest-GR (Grounded-Reporting) derived from PadChest aimed at training GRRG models for CXR images. We curate a public bi-lingual dataset of 4,555 CXR studies with grounded reports (3,099 abnormal and 1,456 normal), each containing complete lists of sentences describing individual present (positive) and absent (negative) findings in English and Spanish. In total, PadChest-GR contains 7,037 positive and 3,422 negative finding sentences. Every positive finding sentence is associated with up to two independent sets of bounding boxes labelled by different readers and has categorical labels for finding type, locations, and progression. To the best of our knowledge, PadChest-GR is the first manually curated dataset designed to train GRRG models for understanding and interpreting radiological images and generated text. By including detailed localization and comprehensive annotations of all clinically relevant findings, it provides a valuable resource for developing and evaluating GRRG models from CXR images. PadChest-GR can be downloaded under request from https://bimcv.cipf.es/bimcv-projects/padchest-gr/
MAIRA-2: Grounded Radiology Report Generation
Jun 06, 2024



Abstract:Radiology reporting is a complex task that requires detailed image understanding, integration of multiple inputs, including comparison with prior imaging, and precise language generation. This makes it ideal for the development and use of generative multimodal models. Here, we extend report generation to include the localisation of individual findings on the image - a task we call grounded report generation. Prior work indicates that grounding is important for clarifying image understanding and interpreting AI-generated text. Therefore, grounded reporting stands to improve the utility and transparency of automated report drafting. To enable evaluation of grounded reporting, we propose a novel evaluation framework - RadFact - leveraging the reasoning capabilities of large language models (LLMs). RadFact assesses the factuality of individual generated sentences, as well as correctness of generated spatial localisations when present. We introduce MAIRA-2, a large multimodal model combining a radiology-specific image encoder with a LLM, and trained for the new task of grounded report generation on chest X-rays. MAIRA-2 uses more comprehensive inputs than explored previously: the current frontal image, the current lateral image, the prior frontal image and prior report, as well as the Indication, Technique and Comparison sections of the current report. We demonstrate that these additions significantly improve report quality and reduce hallucinations, establishing a new state of the art on findings generation (without grounding) on MIMIC-CXR while demonstrating the feasibility of grounded reporting as a novel and richer task.
Challenges for Responsible AI Design and Workflow Integration in Healthcare: A Case Study of Automatic Feeding Tube Qualification in Radiology
May 08, 2024



Abstract:Nasogastric tubes (NGTs) are feeding tubes that are inserted through the nose into the stomach to deliver nutrition or medication. If not placed correctly, they can cause serious harm, even death to patients. Recent AI developments demonstrate the feasibility of robustly detecting NGT placement from Chest X-ray images to reduce risks of sub-optimally or critically placed NGTs being missed or delayed in their detection, but gaps remain in clinical practice integration. In this study, we present a human-centered approach to the problem and describe insights derived following contextual inquiry and in-depth interviews with 15 clinical stakeholders. The interviews helped understand challenges in existing workflows, and how best to align technical capabilities with user needs and expectations. We discovered the trade-offs and complexities that need consideration when choosing suitable workflow stages, target users, and design configurations for different AI proposals. We explored how to balance AI benefits and risks for healthcare staff and patients within broader organizational and medical-legal constraints. We also identified data issues related to edge cases and data biases that affect model training and evaluation; how data documentation practices influence data preparation and labelling; and how to measure relevant AI outcomes reliably in future evaluations. We discuss how our work informs design and development of AI applications that are clinically useful, ethical, and acceptable in real-world healthcare services.
RAD-DINO: Exploring Scalable Medical Image Encoders Beyond Text Supervision
Jan 19, 2024Abstract:Language-supervised pre-training has proven to be a valuable method for extracting semantically meaningful features from images, serving as a foundational element in multimodal systems within the computer vision and medical imaging domains. However, resulting features are limited by the information contained within the text. This is particularly problematic in medical imaging, where radiologists' written findings focus on specific observations; a challenge compounded by the scarcity of paired imaging-text data due to concerns over leakage of personal health information. In this work, we fundamentally challenge the prevailing reliance on language supervision for learning general purpose biomedical imaging encoders. We introduce RAD-DINO, a biomedical image encoder pre-trained solely on unimodal biomedical imaging data that obtains similar or greater performance than state-of-the-art biomedical language supervised models on a diverse range of benchmarks. Specifically, the quality of learned representations is evaluated on standard imaging tasks (classification and semantic segmentation), and a vision-language alignment task (text report generation from images). To further demonstrate the drawback of language supervision, we show that features from RAD-DINO correlate with other medical records (e.g., sex or age) better than language-supervised models, which are generally not mentioned in radiology reports. Finally, we conduct a series of ablations determining the factors in RAD-DINO's performance; notably, we observe that RAD-DINO's downstream performance scales well with the quantity and diversity of training data, demonstrating that image-only supervision is a scalable approach for training a foundational biomedical image encoder.
MAIRA-1: A specialised large multimodal model for radiology report generation
Nov 22, 2023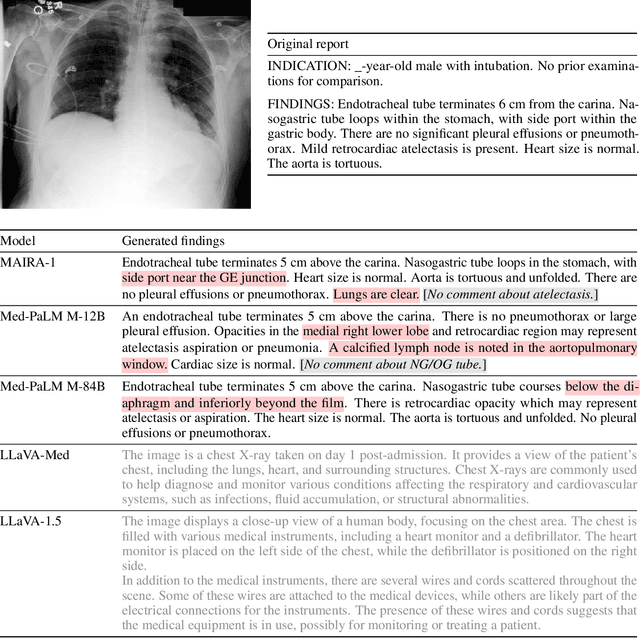
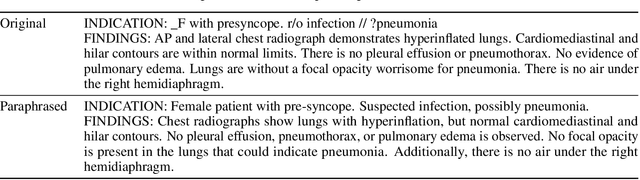
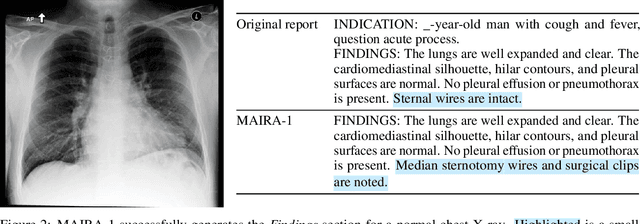
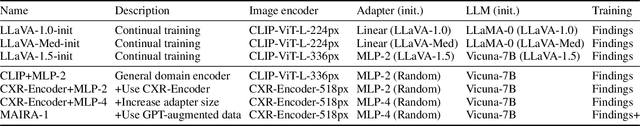



Abstract:We present a radiology-specific multimodal model for the task for generating radiological reports from chest X-rays (CXRs). Our work builds on the idea that large language model(s) can be equipped with multimodal capabilities through alignment with pre-trained vision encoders. On natural images, this has been shown to allow multimodal models to gain image understanding and description capabilities. Our proposed model (MAIRA-1) leverages a CXR-specific image encoder in conjunction with a fine-tuned large language model based on Vicuna-7B, and text-based data augmentation, to produce reports with state-of-the-art quality. In particular, MAIRA-1 significantly improves on the radiologist-aligned RadCliQ metric and across all lexical metrics considered. Manual review of model outputs demonstrates promising fluency and accuracy of generated reports while uncovering failure modes not captured by existing evaluation practices. More information and resources can be found on the project website: https://aka.ms/maira.
Compositional Zero-Shot Domain Transfer with Text-to-Text Models
Mar 23, 2023Abstract:Label scarcity is a bottleneck for improving task performance in specialised domains. We propose a novel compositional transfer learning framework (DoT5 - domain compositional zero-shot T5) for zero-shot domain transfer. Without access to in-domain labels, DoT5 jointly learns domain knowledge (from MLM of unlabelled in-domain free text) and task knowledge (from task training on more readily available general-domain data) in a multi-task manner. To improve the transferability of task training, we design a strategy named NLGU: we simultaneously train NLG for in-domain label-to-data generation which enables data augmentation for self-finetuning and NLU for label prediction. We evaluate DoT5 on the biomedical domain and the resource-lean subdomain of radiology, focusing on NLI, text summarisation and embedding learning. DoT5 demonstrates the effectiveness of compositional transfer learning through multi-task learning. In particular, DoT5 outperforms the current SOTA in zero-shot transfer by over 7 absolute points in accuracy on RadNLI. We validate DoT5 with ablations and a case study demonstrating its ability to solve challenging NLI examples requiring in-domain expertise.
Looking for Out-of-Distribution Environments in Critical Care: A case study with the eICU Database
May 26, 2022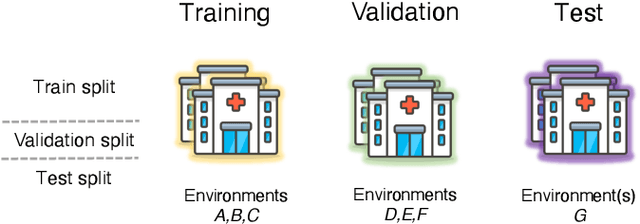
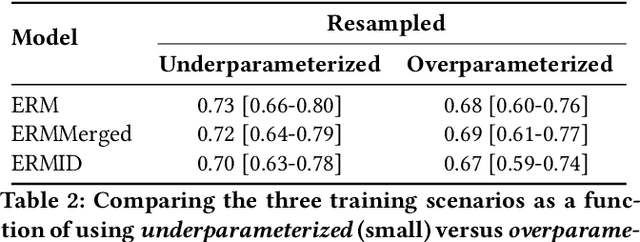
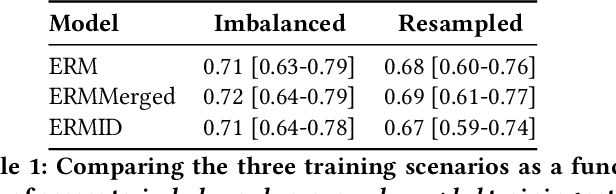
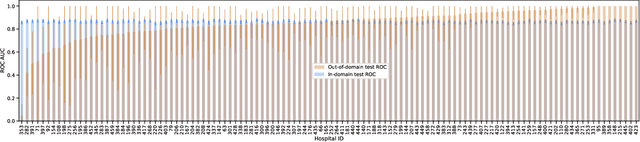

Abstract:Generalizing to new populations and domains in machine learning is still an open problem which has seen increased interest recently. In particular, clinical models show a significant performance drop when tested in settings not seen during training, e.g., new hospitals or population demographics. Recently proposed models for domain generalisation promise to alleviate this problem by learning invariant characteristics across environments, however, there is still scepticism about whether they improve over traditional training. In this work, we take a principled approach to identifying Out of Distribution (OoD) environments, motivated by the problem of cross-hospital generalization in critical care. We propose model-based and heuristic approaches to identify OoD environments and systematically compare models with different levels of held-out information. In particular, based on the assumption that models with access to OoD data should outperform other models, we train models across a range of experimental setups that include leave-one-hospital-out training and cross-sectional feature splits. We find that access to OoD data does not translate to increased performance, pointing to inherent limitations in defining potential OoD environments in the eICU Database potentially due to data harmonisation and sampling. Echoing similar results with other popular clinical benchmarks in the literature, new approaches are required to evaluate robust models in critical care.
Early prediction of respiratory failure in the intensive care unit
May 12, 2021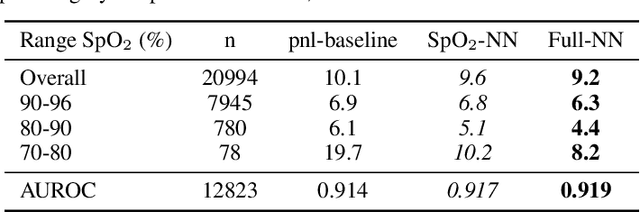
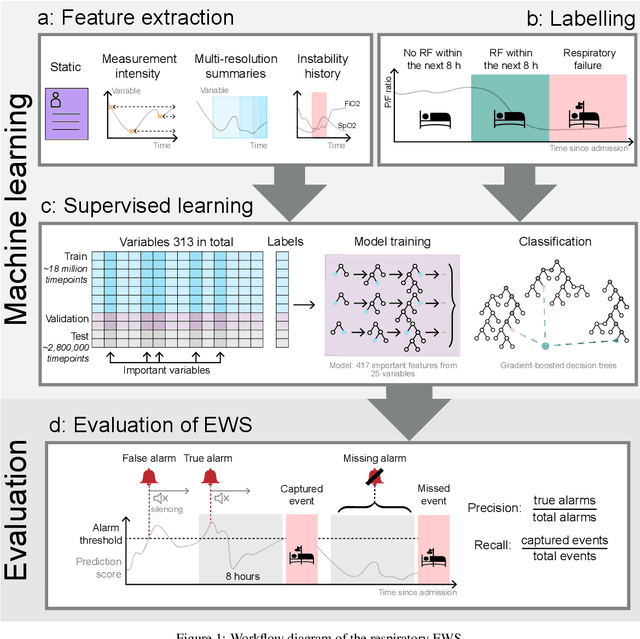
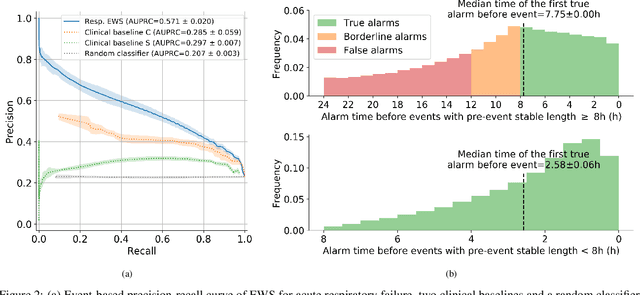
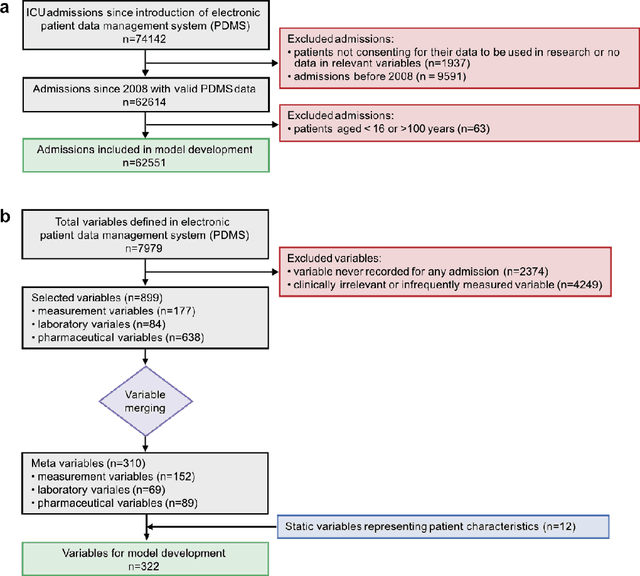
Abstract:The development of respiratory failure is common among patients in intensive care units (ICU). Large data quantities from ICU patient monitoring systems make timely and comprehensive analysis by clinicians difficult but are ideal for automatic processing by machine learning algorithms. Early prediction of respiratory system failure could alert clinicians to patients at risk of respiratory failure and allow for early patient reassessment and treatment adjustment. We propose an early warning system that predicts moderate/severe respiratory failure up to 8 hours in advance. Our system was trained on HiRID-II, a data-set containing more than 60,000 admissions to a tertiary care ICU. An alarm is typically triggered several hours before the beginning of respiratory failure. Our system outperforms a clinical baseline mimicking traditional clinical decision-making based on pulse-oximetric oxygen saturation and the fraction of inspired oxygen. To provide model introspection and diagnostics, we developed an easy-to-use web browser-based system to explore model input data and predictions visually.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge