Mercy Ranjit
RadPhi-3: Small Language Models for Radiology
Nov 19, 2024



Abstract:LLM based copilot assistants are useful in everyday tasks. There is a proliferation in the exploration of AI assistant use cases to support radiology workflows in a reliable manner. In this work, we present RadPhi-3, a Small Language Model instruction tuned from Phi-3-mini-4k-instruct with 3.8B parameters to assist with various tasks in radiology workflows. While impression summary generation has been the primary task which has been explored in prior works w.r.t radiology reports of Chest X-rays, we also explore other useful tasks like change summary generation comparing the current radiology report and its prior report, section extraction from radiology reports, tagging the reports with various pathologies and tubes, lines or devices present in them etc. In-addition, instruction tuning RadPhi-3 involved learning from a credible knowledge source used by radiologists, Radiopaedia.org. RadPhi-3 can be used both to give reliable answers for radiology related queries as well as perform useful tasks related to radiology reports. RadPhi-3 achieves SOTA results on the RaLEs radiology report generation benchmark.
MAIRA-2: Grounded Radiology Report Generation
Jun 06, 2024



Abstract:Radiology reporting is a complex task that requires detailed image understanding, integration of multiple inputs, including comparison with prior imaging, and precise language generation. This makes it ideal for the development and use of generative multimodal models. Here, we extend report generation to include the localisation of individual findings on the image - a task we call grounded report generation. Prior work indicates that grounding is important for clarifying image understanding and interpreting AI-generated text. Therefore, grounded reporting stands to improve the utility and transparency of automated report drafting. To enable evaluation of grounded reporting, we propose a novel evaluation framework - RadFact - leveraging the reasoning capabilities of large language models (LLMs). RadFact assesses the factuality of individual generated sentences, as well as correctness of generated spatial localisations when present. We introduce MAIRA-2, a large multimodal model combining a radiology-specific image encoder with a LLM, and trained for the new task of grounded report generation on chest X-rays. MAIRA-2 uses more comprehensive inputs than explored previously: the current frontal image, the current lateral image, the prior frontal image and prior report, as well as the Indication, Technique and Comparison sections of the current report. We demonstrate that these additions significantly improve report quality and reduce hallucinations, establishing a new state of the art on findings generation (without grounding) on MIMIC-CXR while demonstrating the feasibility of grounded reporting as a novel and richer task.
Bridging the Gap: Dynamic Learning Strategies for Improving Multilingual Performance in LLMs
May 28, 2024Abstract:Large language models (LLMs) are at the forefront of transforming numerous domains globally. However, their inclusivity and effectiveness remain limited for non-Latin scripts and low-resource languages. This paper tackles the imperative challenge of enhancing the multilingual performance of LLMs without extensive training or fine-tuning. Through systematic investigation and evaluation of diverse languages using popular question-answering (QA) datasets, we present novel techniques that unlock the true potential of LLMs in a polyglot landscape. Our approach encompasses three key strategies that yield significant improvements in multilingual proficiency. First, by meticulously optimizing prompts tailored for polyglot LLMs, we unlock their latent capabilities, resulting in substantial performance boosts across languages. Second, we introduce a new hybrid approach that synergizes LLM Retrieval Augmented Generation (RAG) with multilingual embeddings and achieves improved multilingual task performance. Finally, we introduce a novel learning approach that dynamically selects the optimal prompt strategy, LLM model, and embedding model per query at run-time. This dynamic adaptation maximizes the efficacy of LLMs across languages, outperforming best static and random strategies. Additionally, our approach adapts configurations in both offline and online settings, and can seamlessly adapt to new languages and datasets, leading to substantial advancements in multilingual understanding and generation across diverse languages.
Challenges for Responsible AI Design and Workflow Integration in Healthcare: A Case Study of Automatic Feeding Tube Qualification in Radiology
May 08, 2024



Abstract:Nasogastric tubes (NGTs) are feeding tubes that are inserted through the nose into the stomach to deliver nutrition or medication. If not placed correctly, they can cause serious harm, even death to patients. Recent AI developments demonstrate the feasibility of robustly detecting NGT placement from Chest X-ray images to reduce risks of sub-optimally or critically placed NGTs being missed or delayed in their detection, but gaps remain in clinical practice integration. In this study, we present a human-centered approach to the problem and describe insights derived following contextual inquiry and in-depth interviews with 15 clinical stakeholders. The interviews helped understand challenges in existing workflows, and how best to align technical capabilities with user needs and expectations. We discovered the trade-offs and complexities that need consideration when choosing suitable workflow stages, target users, and design configurations for different AI proposals. We explored how to balance AI benefits and risks for healthcare staff and patients within broader organizational and medical-legal constraints. We also identified data issues related to edge cases and data biases that affect model training and evaluation; how data documentation practices influence data preparation and labelling; and how to measure relevant AI outcomes reliably in future evaluations. We discuss how our work informs design and development of AI applications that are clinically useful, ethical, and acceptable in real-world healthcare services.
RAD-PHI2: Instruction Tuning PHI-2 for Radiology
Mar 12, 2024



Abstract:Small Language Models (SLMs) have shown remarkable performance in general domain language understanding, reasoning and coding tasks, but their capabilities in the medical domain, particularly concerning radiology text, is less explored. In this study, we investigate the application of SLMs for general radiology knowledge specifically question answering related to understanding of symptoms, radiological appearances of findings, differential diagnosis, assessing prognosis, and suggesting treatments w.r.t diseases pertaining to different organ systems. Additionally, we explore the utility of SLMs in handling text-related tasks with respect to radiology reports within AI-driven radiology workflows. We fine-tune Phi-2, a SLM with 2.7 billion parameters using high-quality educational content from Radiopaedia, a collaborative online radiology resource. The resulting language model, RadPhi-2-Base, exhibits the ability to address general radiology queries across various systems (e.g., chest, cardiac). Furthermore, we investigate Phi-2 for instruction tuning, enabling it to perform specific tasks. By fine-tuning Phi-2 on both general domain tasks and radiology-specific tasks related to chest X-ray reports, we create Rad-Phi2. Our empirical results reveal that Rad-Phi2 Base and Rad-Phi2 perform comparably or even outperform larger models such as Mistral-7B-Instruct-v0.2 and GPT-4 providing concise and precise answers. In summary, our work demonstrates the feasibility and effectiveness of utilizing SLMs in radiology workflows both for knowledge related queries as well as for performing specific tasks related to radiology reports thereby opening up new avenues for enhancing the quality and efficiency of radiology practice.
MAIRA-1: A specialised large multimodal model for radiology report generation
Nov 22, 2023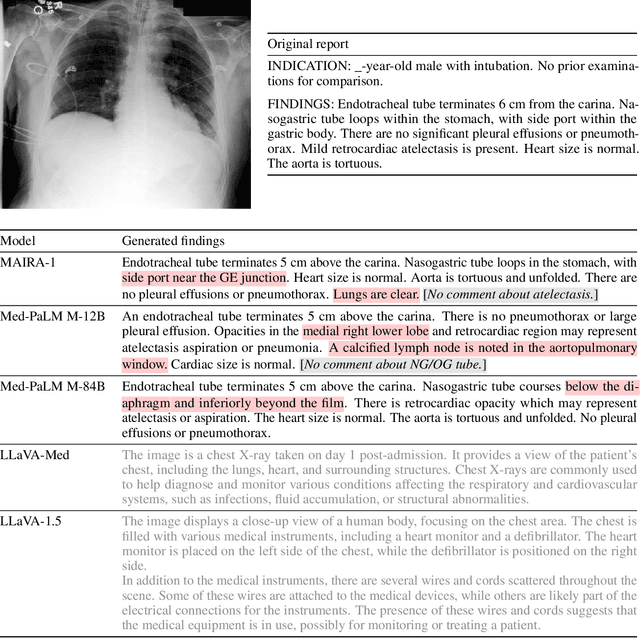
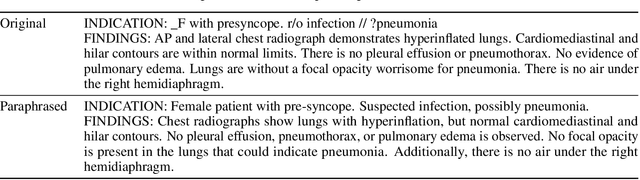
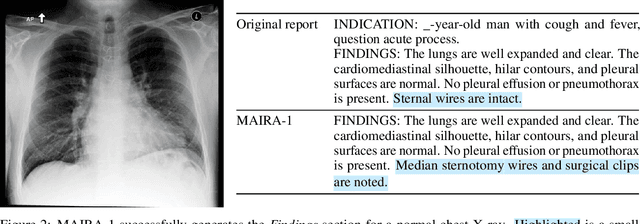
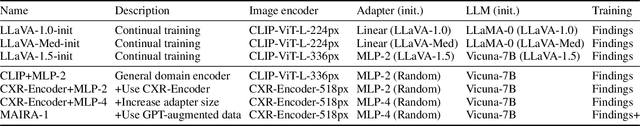



Abstract:We present a radiology-specific multimodal model for the task for generating radiological reports from chest X-rays (CXRs). Our work builds on the idea that large language model(s) can be equipped with multimodal capabilities through alignment with pre-trained vision encoders. On natural images, this has been shown to allow multimodal models to gain image understanding and description capabilities. Our proposed model (MAIRA-1) leverages a CXR-specific image encoder in conjunction with a fine-tuned large language model based on Vicuna-7B, and text-based data augmentation, to produce reports with state-of-the-art quality. In particular, MAIRA-1 significantly improves on the radiologist-aligned RadCliQ metric and across all lexical metrics considered. Manual review of model outputs demonstrates promising fluency and accuracy of generated reports while uncovering failure modes not captured by existing evaluation practices. More information and resources can be found on the project website: https://aka.ms/maira.
Breaking Language Barriers with a LEAP: Learning Strategies for Polyglot LLMs
May 28, 2023



Abstract:Large language models (LLMs) are at the forefront of transforming numerous domains globally. However, their inclusivity and effectiveness remain limited for non-Latin scripts and low-resource languages. This paper tackles the imperative challenge of enhancing the multilingual performance of LLMs, specifically focusing on Generative models. Through systematic investigation and evaluation of diverse languages using popular question-answering (QA) datasets, we present novel techniques that unlock the true potential of LLMs in a polyglot landscape. Our approach encompasses three key strategies that yield remarkable improvements in multilingual proficiency. First, by meticulously optimizing prompts tailored for polyglot LLMs, we unlock their latent capabilities, resulting in substantial performance boosts across languages. Second, we introduce a new hybrid approach that synergizes GPT generation with multilingual embeddings and achieves significant multilingual performance improvement on critical tasks like QA and retrieval. Finally, to further propel the performance of polyglot LLMs, we introduce a novel learning algorithm that dynamically selects the optimal prompt strategy, LLM model, and embeddings per query. This dynamic adaptation maximizes the efficacy of LLMs across languages, outperforming best static and random strategies. Our results show substantial advancements in multilingual understanding and generation across a diverse range of languages.
Retrieval Augmented Chest X-Ray Report Generation using OpenAI GPT models
May 05, 2023



Abstract:We propose Retrieval Augmented Generation (RAG) as an approach for automated radiology report writing that leverages multimodally aligned embeddings from a contrastively pretrained vision language model for retrieval of relevant candidate radiology text for an input radiology image and a general domain generative model like OpenAI text-davinci-003, gpt-3.5-turbo and gpt-4 for report generation using the relevant radiology text retrieved. This approach keeps hallucinated generations under check and provides capabilities to generate report content in the format we desire leveraging the instruction following capabilities of these generative models. Our approach achieves better clinical metrics with a BERTScore of 0.2865 ({\Delta}+ 25.88%) and Semb score of 0.4026 ({\Delta}+ 6.31%). Our approach can be broadly relevant for different clinical settings as it allows to augment the automated radiology report generation process with content relevant for that setting while also having the ability to inject user intents and requirements in the prompts as part of the report generation process to modulate the content and format of the generated reports as applicable for that clinical setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge