Sotirios A. Tsaftaris
Metrics reloaded: Pitfalls and recommendations for image analysis validation
Jun 03, 2022
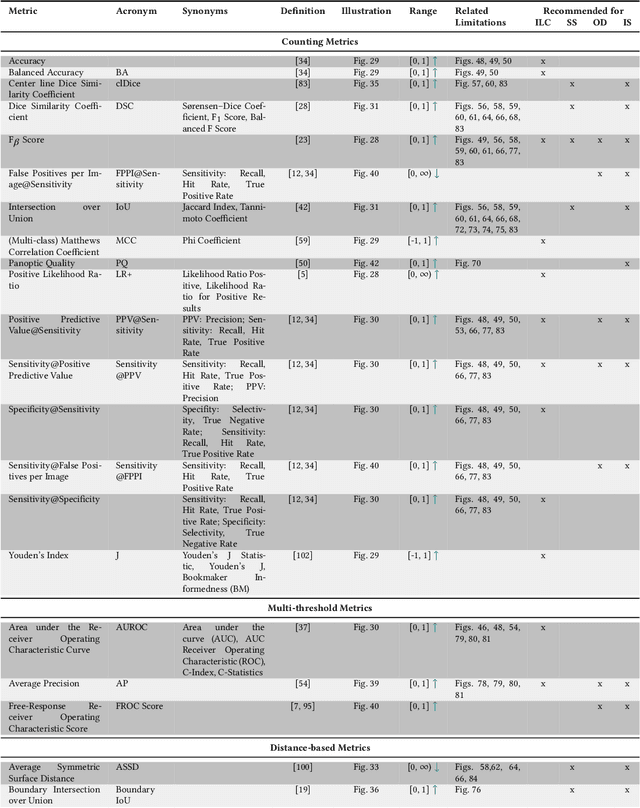
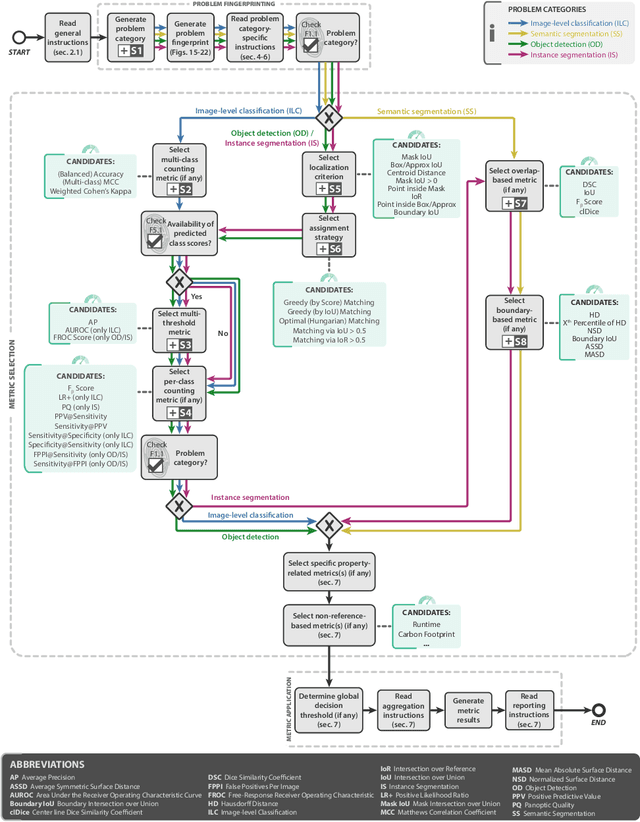
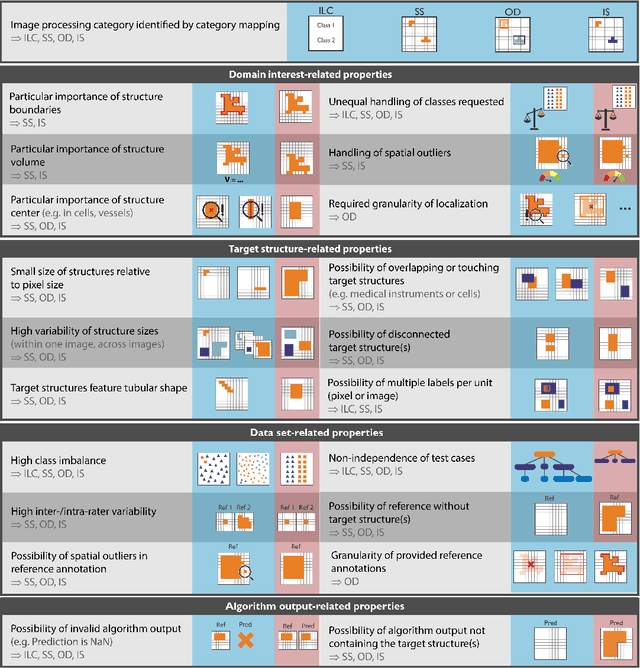
Abstract:The field of automatic biomedical image analysis crucially depends on robust and meaningful performance metrics for algorithm validation. Current metric usage, however, is often ill-informed and does not reflect the underlying domain interest. Here, we present a comprehensive framework that guides researchers towards choosing performance metrics in a problem-aware manner. Specifically, we focus on biomedical image analysis problems that can be interpreted as a classification task at image, object or pixel level. The framework first compiles domain interest-, target structure-, data set- and algorithm output-related properties of a given problem into a problem fingerprint, while also mapping it to the appropriate problem category, namely image-level classification, semantic segmentation, instance segmentation, or object detection. It then guides users through the process of selecting and applying a set of appropriate validation metrics while making them aware of potential pitfalls related to individual choices. In this paper, we describe the current status of the Metrics Reloaded recommendation framework, with the goal of obtaining constructive feedback from the image analysis community. The current version has been developed within an international consortium of more than 60 image analysis experts and will be made openly available as a user-friendly toolkit after community-driven optimization.
Causal Machine Learning for Healthcare and Precision Medicine
May 31, 2022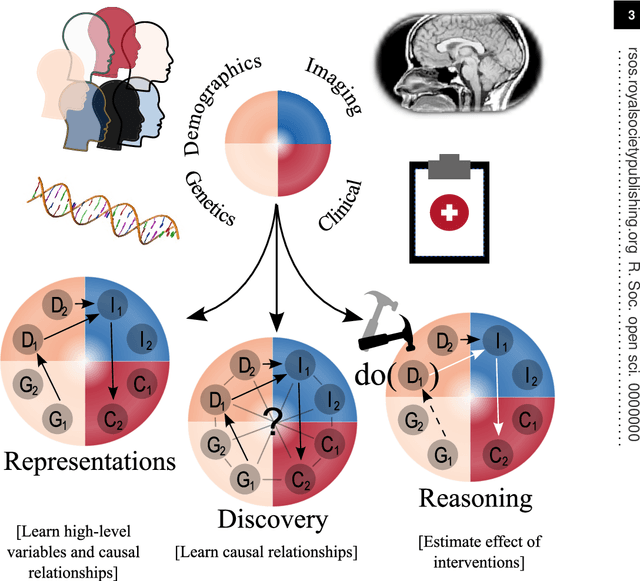
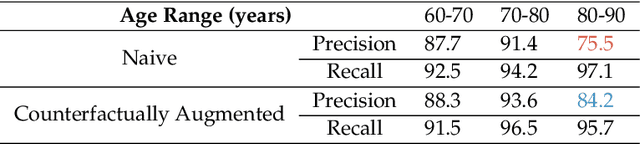
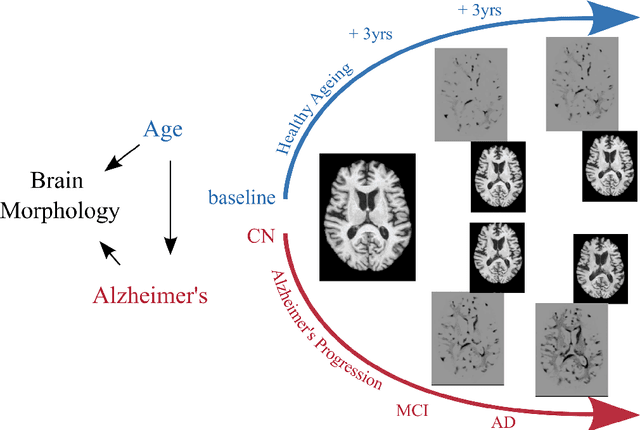
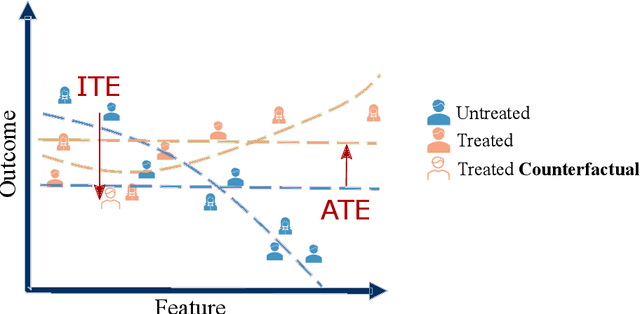
Abstract:Causal machine learning (CML) has experienced increasing popularity in healthcare. Beyond the inherent capabilities of adding domain knowledge into learning systems, CML provides a complete toolset for investigating how a system would react to an intervention (e.g.\ outcome given a treatment). Quantifying effects of interventions allows actionable decisions to be made whilst maintaining robustness in the presence of confounders. Here, we explore how causal inference can be incorporated into different aspects of clinical decision support (CDS) systems by using recent advances in machine learning. Throughout this paper, we use Alzheimer's disease (AD) to create examples for illustrating how CML can be advantageous in clinical scenarios. Furthermore, we discuss important challenges present in healthcare applications such as processing high-dimensional and unstructured data, generalisation to out-of-distribution samples, and temporal relationships, that despite the great effort from the research community remain to be solved. Finally, we review lines of research within causal representation learning, causal discovery and causal reasoning which offer the potential towards addressing the aforementioned challenges.
Unintended memorisation of unique features in neural networks
May 20, 2022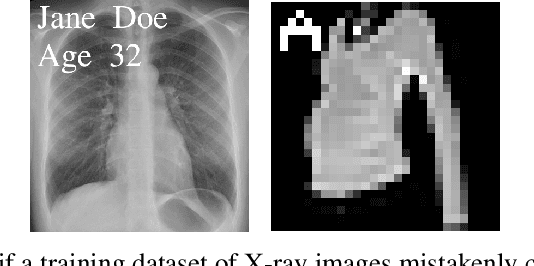
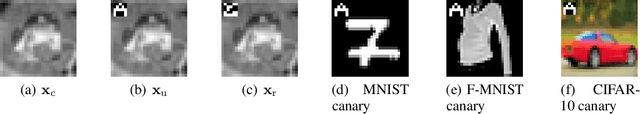
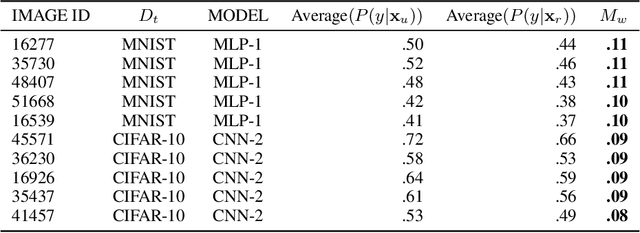

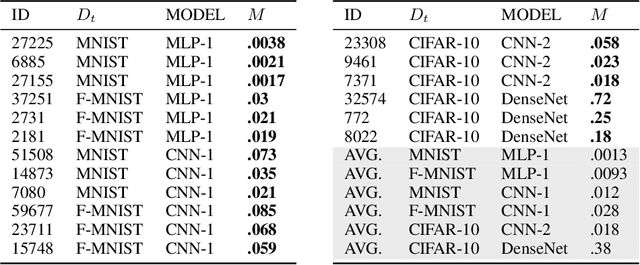
Abstract:Neural networks pose a privacy risk due to their propensity to memorise and leak training data. We show that unique features occurring only once in training data are memorised by discriminative multi-layer perceptrons and convolutional neural networks trained on benchmark imaging datasets. We design our method for settings where sensitive training data is not available, for example medical imaging. Our setting knows the unique feature, but not the training data, model weights or the unique feature's label. We develop a score estimating a model's sensitivity to a unique feature by comparing the KL divergences of the model's output distributions given modified out-of-distribution images. We find that typical strategies to prevent overfitting do not prevent unique feature memorisation. And that images containing a unique feature are highly influential, regardless of the influence the images's other features. We also find a significant variation in memorisation with training seed. These results imply that neural networks pose a privacy risk to rarely occurring private information. This risk is more pronounced in healthcare applications since sensitive patient information can be memorised when it remains in training data due to an imperfect data sanitisation process.
Adversarial Counterfactual Augmentation: Application in Alzheimer's Disease Classification
Mar 15, 2022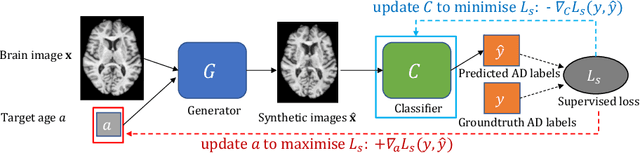
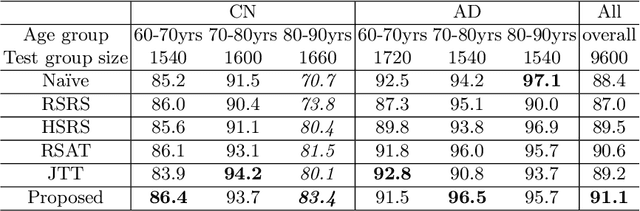
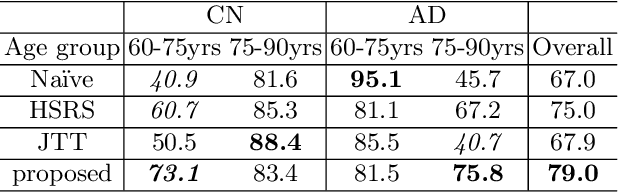
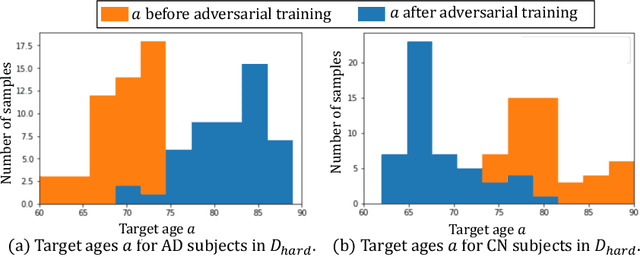
Abstract:Data augmentation has been widely used in deep learning to reduce over-fitting and improve the robustness of models. However, traditional data augmentation techniques, e.g., rotation, cropping, flipping, etc., do not consider \textit{semantic} transformations, e.g., changing the age of a brain image. Previous works tried to achieve semantic augmentation by generating \textit{counterfactuals}, but they focused on how to train deep generative models and randomly created counterfactuals with the generative models without considering which counterfactuals are most \textit{effective} for improving downstream training. Different from these approaches, in this work, we propose a novel adversarial counterfactual augmentation scheme that aims to find the most \textit{effective} counterfactuals to improve downstream tasks with a pre-trained generative model. Specifically, we construct an adversarial game where we update the input \textit{conditional factor} of the generator and the downstream \textit{classifier} with gradient backpropagation alternatively and iteratively. The key idea is to find conditional factors that can result in \textit{hard} counterfactuals for the classifier. This can be viewed as finding the `\textit{weakness}' of the classifier and purposely forcing it to \textit{overcome} its weakness via the generative model. To demonstrate the effectiveness of the proposed approach, we validate the method with the classification of Alzheimer's Disease (AD) as the downstream task based on a pre-trained brain ageing synthesis model. We show the proposed approach improves test accuracy and can alleviate spurious correlations. Code will be released upon acceptance.
Diffusion Causal Models for Counterfactual Estimation
Feb 21, 2022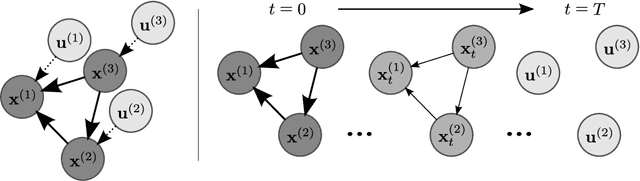
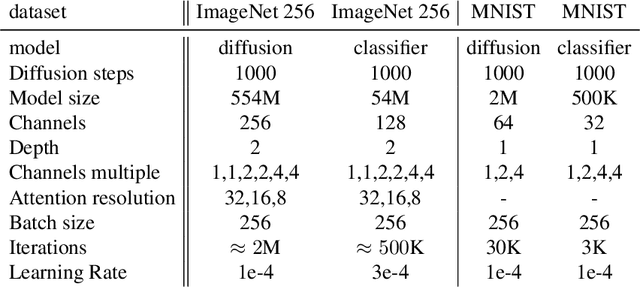
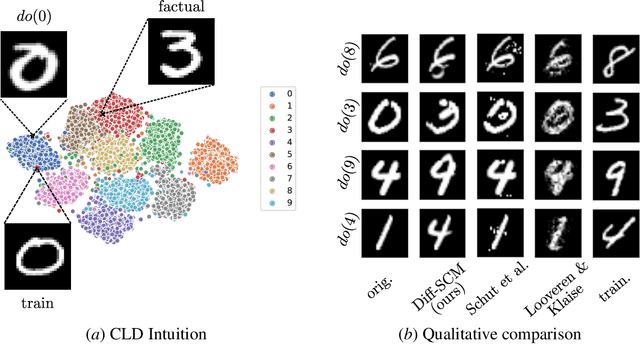
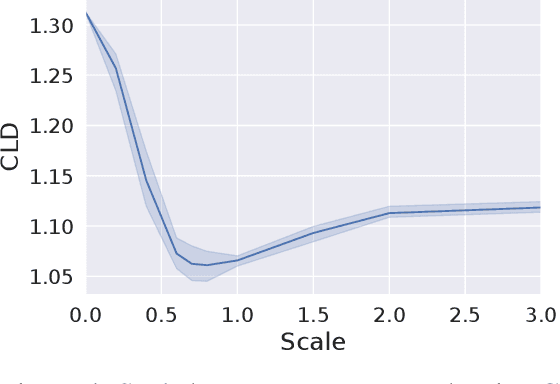
Abstract:We consider the task of counterfactual estimation from observational imaging data given a known causal structure. In particular, quantifying the causal effect of interventions for high-dimensional data with neural networks remains an open challenge. Herein we propose Diff-SCM, a deep structural causal model that builds on recent advances of generative energy-based models. In our setting, inference is performed by iteratively sampling gradients of the marginal and conditional distributions entailed by the causal model. Counterfactual estimation is achieved by firstly inferring latent variables with deterministic forward diffusion, then intervening on a reverse diffusion process using the gradients of an anti-causal predictor w.r.t the input. Furthermore, we propose a metric for evaluating the generated counterfactuals. We find that Diff-SCM produces more realistic and minimal counterfactuals than baselines on MNIST data and can also be applied to ImageNet data. Code is available https://github.com/vios-s/Diff-SCM.
Measuring Unintended Memorisation of Unique Private Features in Neural Networks
Feb 16, 2022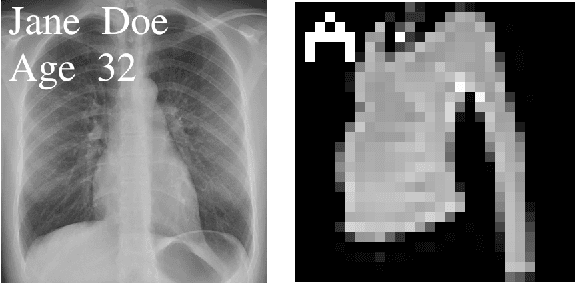
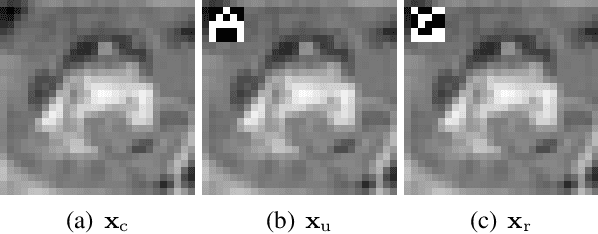
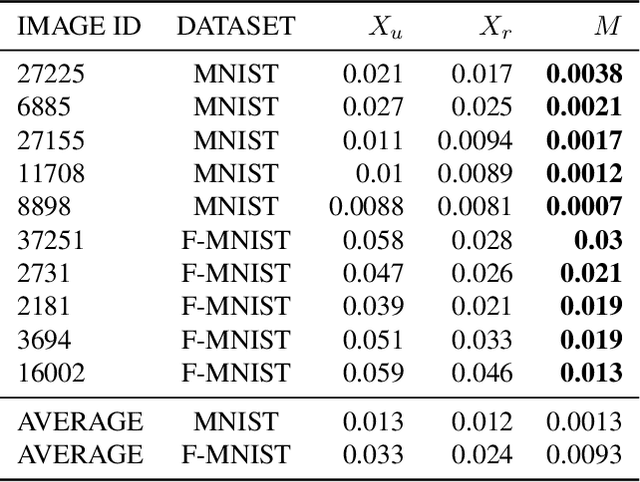

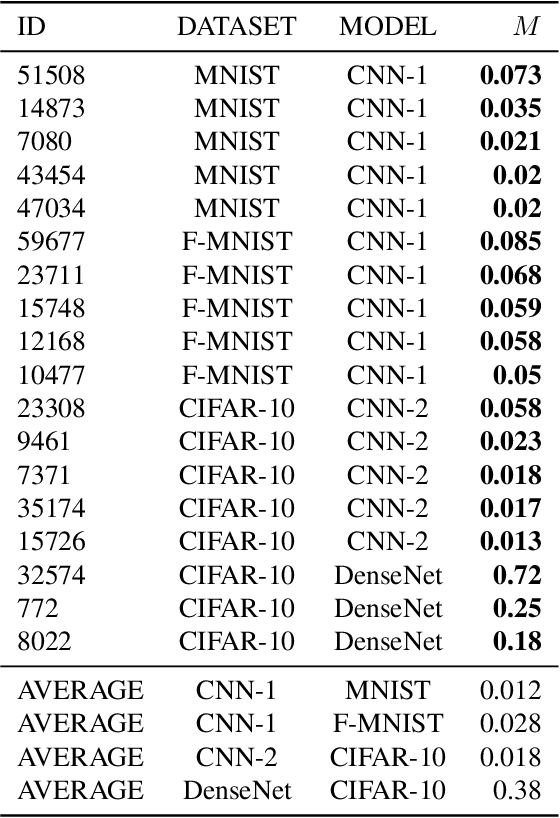
Abstract:Neural networks pose a privacy risk to training data due to their propensity to memorise and leak information. Focusing on image classification, we show that neural networks also unintentionally memorise unique features even when they occur only once in training data. An example of a unique feature is a person's name that is accidentally present on a training image. Assuming access to the inputs and outputs of a trained model, the domain of the training data, and knowledge of unique features, we develop a score estimating the model's sensitivity to a unique feature by comparing the KL divergences of the model's output distributions given modified out-of-distribution images. Our results suggest that unique features are memorised by multi-layer perceptrons and convolutional neural networks trained on benchmark datasets, such as MNIST, Fashion-MNIST and CIFAR-10. We find that strategies to prevent overfitting (e.g.\ early stopping, regularisation, batch normalisation) do not prevent memorisation of unique features. These results imply that neural networks pose a privacy risk to rarely occurring private information. These risks can be more pronounced in healthcare applications if patient information is present in the training data.
Indication as Prior Knowledge for Multimodal Disease Classification in Chest Radiographs with Transformers
Feb 12, 2022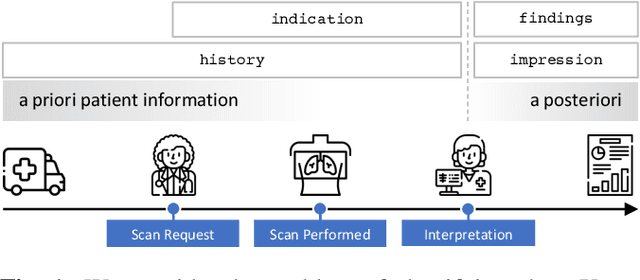
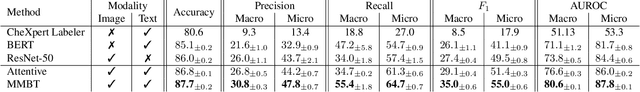
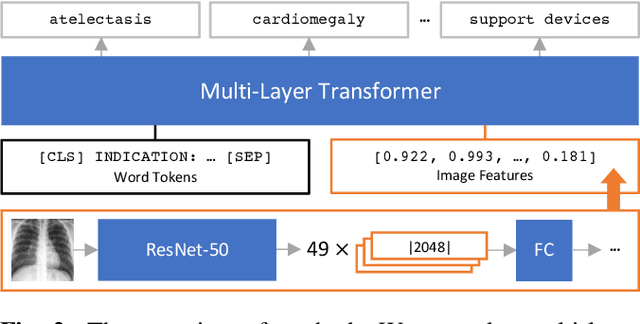
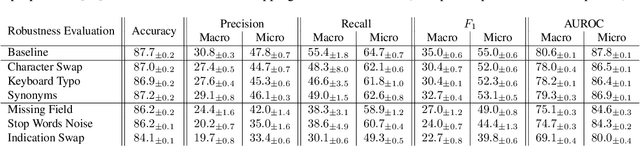
Abstract:When a clinician refers a patient for an imaging exam, they include the reason (e.g. relevant patient history, suspected disease) in the scan request; this appears as the indication field in the radiology report. The interpretation and reporting of the image are substantially influenced by this request text, steering the radiologist to focus on particular aspects of the image. We use the indication field to drive better image classification, by taking a transformer network which is unimodally pre-trained on text (BERT) and fine-tuning it for multimodal classification of a dual image-text input. We evaluate the method on the MIMIC-CXR dataset, and present ablation studies to investigate the effect of the indication field on the classification performance. The experimental results show our approach achieves 87.8 average micro AUROC, outperforming the state-of-the-art methods for unimodal (84.4) and multimodal (86.0) classification. Our code is available at https://github.com/jacenkow/mmbt.
MyoPS: A Benchmark of Myocardial Pathology Segmentation Combining Three-Sequence Cardiac Magnetic Resonance Images
Jan 10, 2022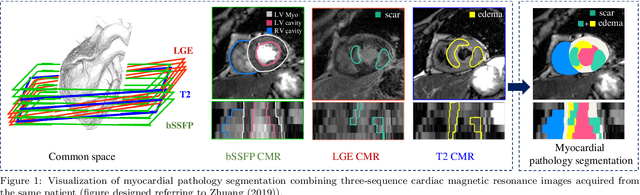
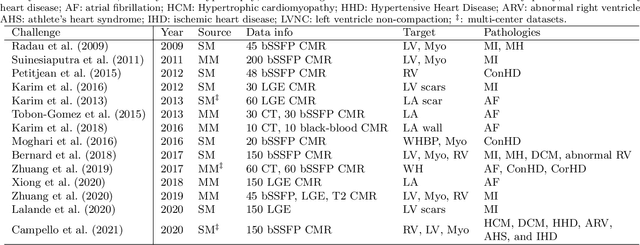
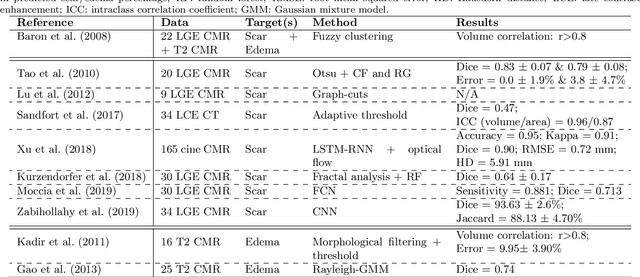
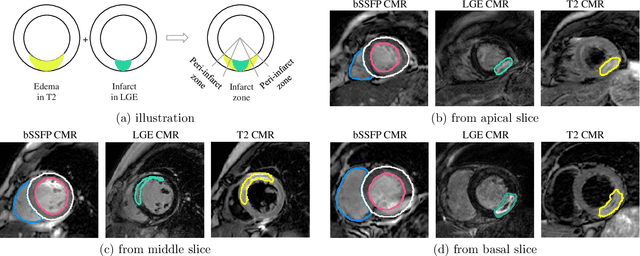
Abstract:Assessment of myocardial viability is essential in diagnosis and treatment management of patients suffering from myocardial infarction, and classification of pathology on myocardium is the key to this assessment. This work defines a new task of medical image analysis, i.e., to perform myocardial pathology segmentation (MyoPS) combining three-sequence cardiac magnetic resonance (CMR) images, which was first proposed in the MyoPS challenge, in conjunction with MICCAI 2020. The challenge provided 45 paired and pre-aligned CMR images, allowing algorithms to combine the complementary information from the three CMR sequences for pathology segmentation. In this article, we provide details of the challenge, survey the works from fifteen participants and interpret their methods according to five aspects, i.e., preprocessing, data augmentation, learning strategy, model architecture and post-processing. In addition, we analyze the results with respect to different factors, in order to examine the key obstacles and explore potential of solutions, as well as to provide a benchmark for future research. We conclude that while promising results have been reported, the research is still in the early stage, and more in-depth exploration is needed before a successful application to the clinics. Note that MyoPS data and evaluation tool continue to be publicly available upon registration via its homepage (www.sdspeople.fudan.edu.cn/zhuangxiahai/0/myops20/).
A Tutorial on Learning Disentangled Representations in the Imaging Domain
Aug 26, 2021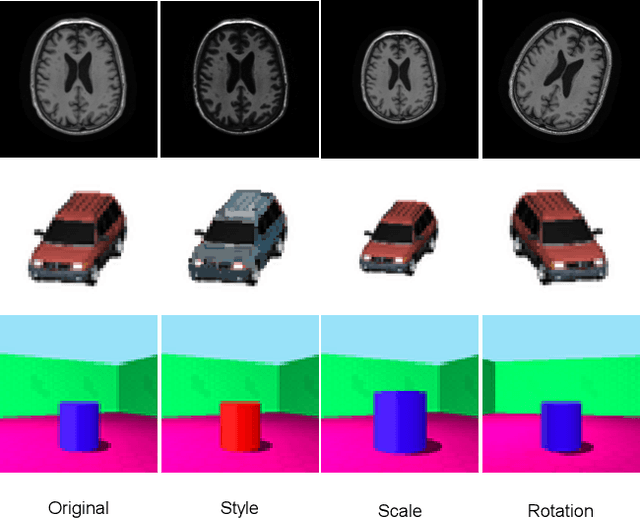
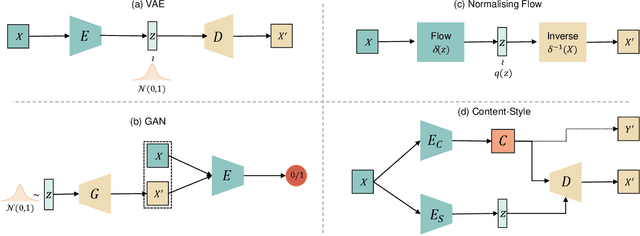
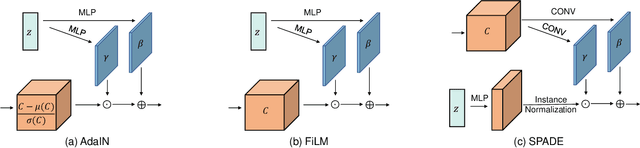
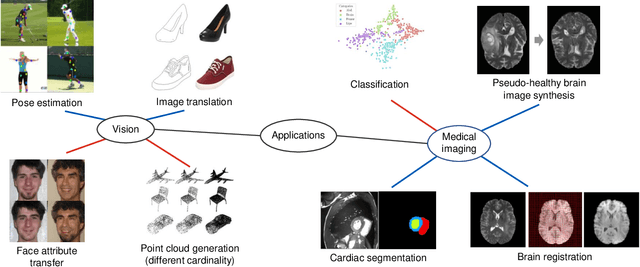
Abstract:Disentangled representation learning has been proposed as an approach to learning general representations. This can be done in the absence of, or with limited, annotations. A good general representation can be readily fine-tuned for new target tasks using modest amounts of data, or even be used directly in unseen domains achieving remarkable performance in the corresponding task. This alleviation of the data and annotation requirements offers tantalising prospects for tractable and affordable applications in computer vision and healthcare. Finally, disentangled representations can offer model explainability and can help us understand the underlying causal relations of the factors of variation, increasing their suitability for real-world deployment. In this tutorial paper, we will offer an overview of the disentangled representation learning, its building blocks and criteria, and discuss applications in computer vision and medical imaging. We conclude our tutorial by presenting the identified opportunities for the integration of recent machine learning advances into disentanglement, as well as the remaining challenges.
Re-using Adversarial Mask Discriminators for Test-time Training under Distribution Shifts
Aug 26, 2021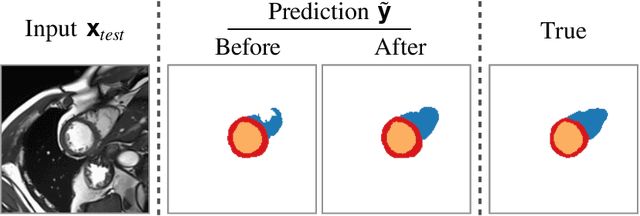

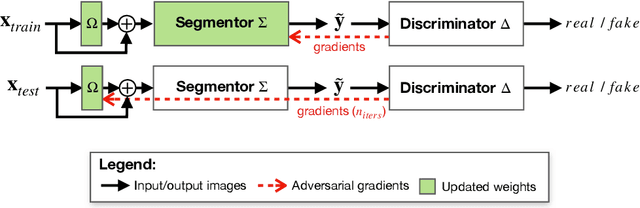

Abstract:Thanks to their ability to learn flexible data-driven losses, Generative Adversarial Networks (GANs) are an integral part of many semi- and weakly-supervised methods for medical image segmentation. GANs jointly optimise a generator and an adversarial discriminator on a set of training data. After training has completed, the discriminator is usually discarded and only the generator is used for inference. But should we discard discriminators? In this work, we argue that training stable discriminators produces expressive loss functions that we can re-use at inference to detect and correct segmentation mistakes. First, we identify key challenges and suggest possible solutions to make discriminators re-usable at inference. Then, we show that we can combine discriminators with image reconstruction costs (via decoders) to further improve the model. Our method is simple and improves the test-time performance of pre-trained GANs. Moreover, we show that it is compatible with standard post-processing techniques and it has potentials to be used for Online Continual Learning. With our work, we open new research avenues for re-using adversarial discriminators at inference.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge