Pengfei Guo
Over-and-Under Complete Convolutional RNN for MRI Reconstruction
Jun 25, 2021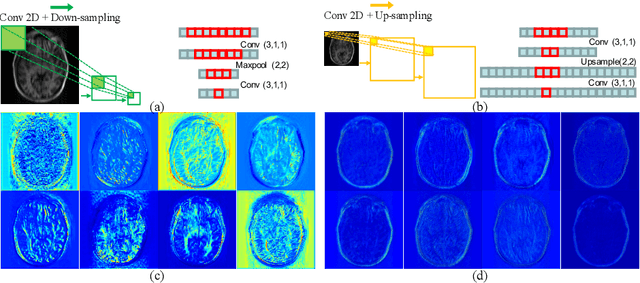
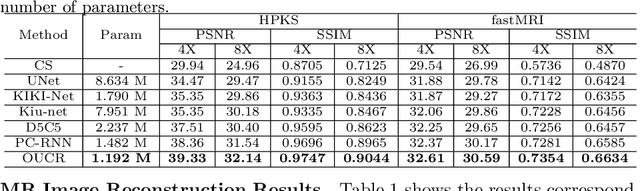
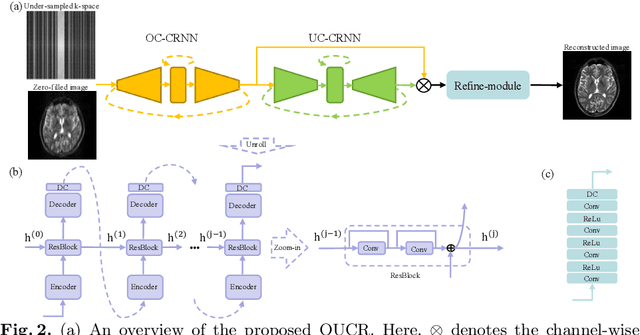
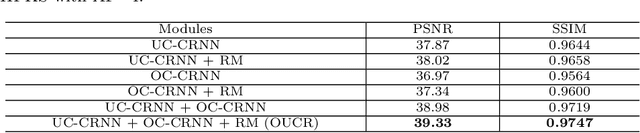
Abstract:Reconstructing magnetic resonance (MR) images from undersampled data is a challenging problem due to various artifacts introduced by the under-sampling operation. Recent deep learning-based methods for MR image reconstruction usually leverage a generic auto-encoder architecture which captures low-level features at the initial layers and high-level features at the deeper layers. Such networks focus much on global features which may not be optimal to reconstruct the fully-sampled image. In this paper, we propose an Over-and-Under Complete Convolutional Recurrent Neural Network (OUCR), which consists of an overcomplete and an undercomplete Convolutional Recurrent Neural Network(CRNN). The overcomplete branch gives special attention in learning local structures by restraining the receptive field of the network. Combining it with the undercomplete branch leads to a network which focuses more on low-level features without losing out on the global structures. Extensive experiments on two datasets demonstrate that the proposed method achieves significant improvements over the compressed sensing and popular deep learning-based methods with less number of trainable parameters.
Multi-institutional Collaborations for Improving Deep Learning-based Magnetic Resonance Image Reconstruction Using Federated Learning
Mar 10, 2021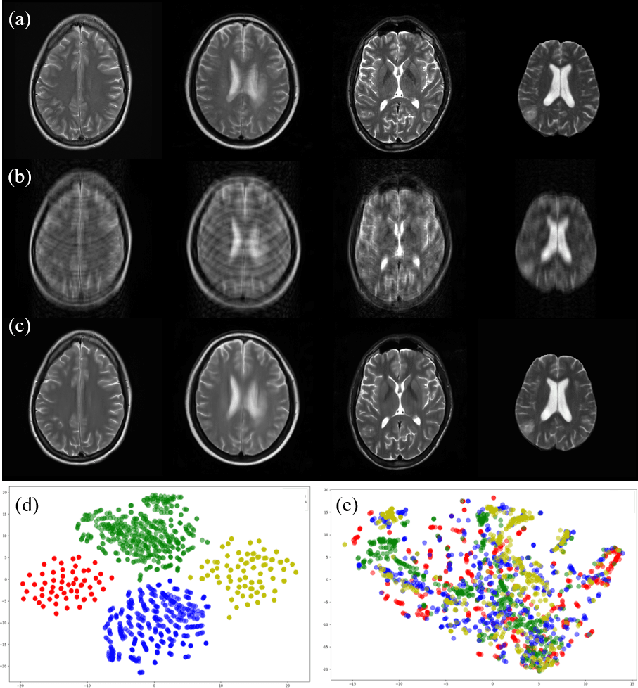
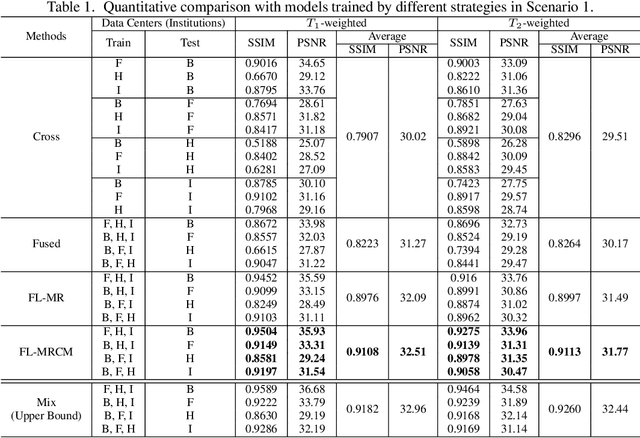
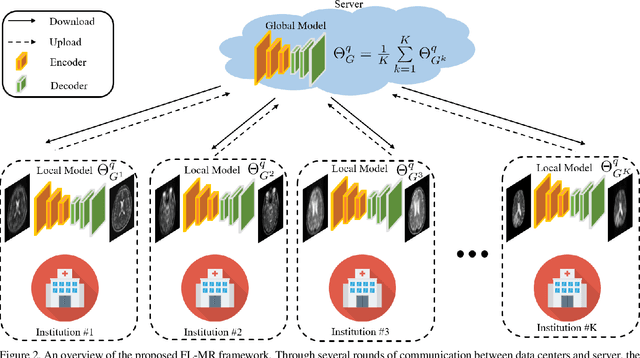
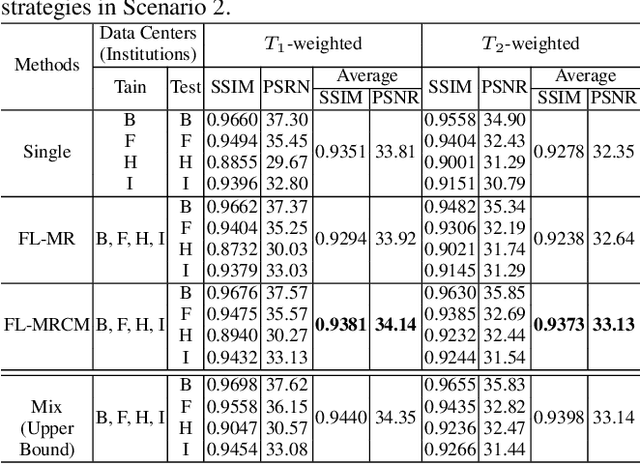
Abstract:Fast and accurate reconstruction of magnetic resonance (MR) images from under-sampled data is important in many clinical applications. In recent years, deep learning-based methods have been shown to produce superior performance on MR image reconstruction. However, these methods require large amounts of data which is difficult to collect and share due to the high cost of acquisition and medical data privacy regulations. In order to overcome this challenge, we propose a federated learning (FL) based solution in which we take advantage of the MR data available at different institutions while preserving patients' privacy. However, the generalizability of models trained with the FL setting can still be suboptimal due to domain shift, which results from the data collected at multiple institutions with different sensors, disease types, and acquisition protocols, etc. With the motivation of circumventing this challenge, we propose a cross-site modeling for MR image reconstruction in which the learned intermediate latent features among different source sites are aligned with the distribution of the latent features at the target site. Extensive experiments are conducted to provide various insights about FL for MR image reconstruction. Experimental results demonstrate that the proposed framework is a promising direction to utilize multi-institutional data without compromising patients' privacy for achieving improved MR image reconstruction. Our code will be available at https://github.com/guopengf/FLMRCM.
Cuid: A new study of perceived image quality and its subjective assessment
Sep 28, 2020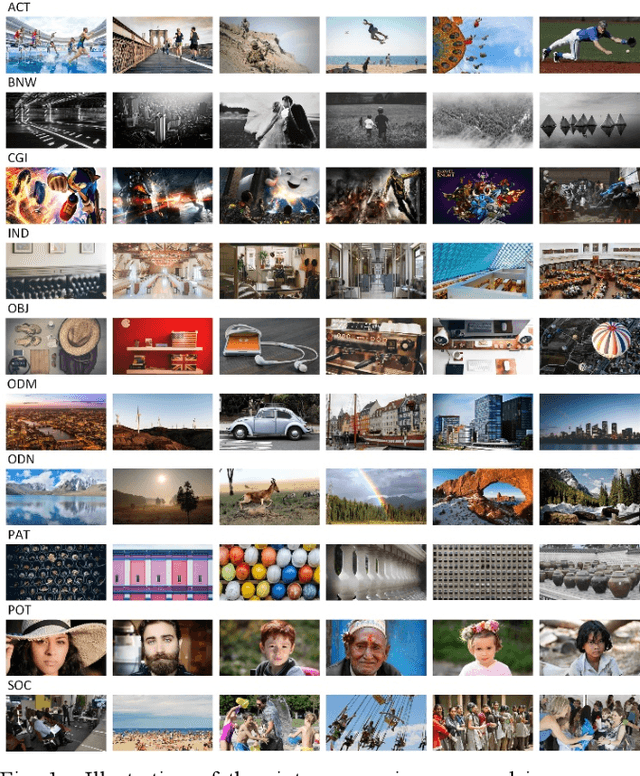
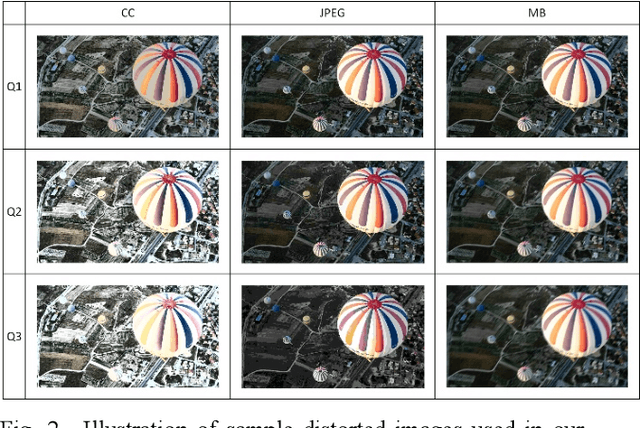
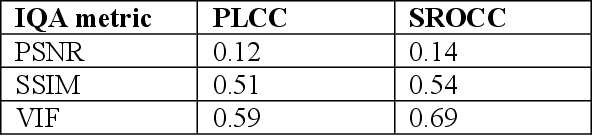


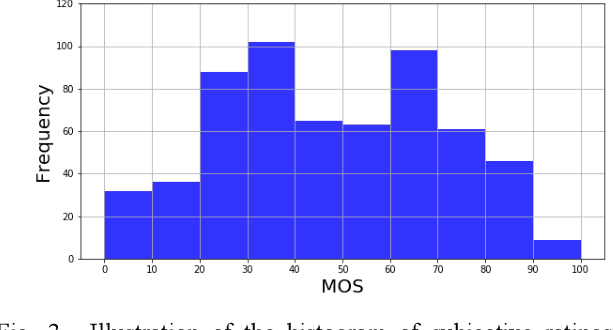
Abstract:Research on image quality assessment (IQA) remains limited mainly due to our incomplete knowledge about human visual perception. Existing IQA algorithms have been designed or trained with insufficient subjective data with a small degree of stimulus variability. This has led to challenges for those algorithms to handle complexity and diversity of real-world digital content. Perceptual evidence from human subjects serves as a grounding for the development of advanced IQA algorithms. It is thus critical to acquire reliable subjective data with controlled perception experiments that faithfully reflect human behavioural responses to distortions in visual signals. In this paper, we present a new study of image quality perception where subjective ratings were collected in a controlled lab environment. We investigate how quality perception is affected by a combination of different categories of images and different types and levels of distortions. The database will be made publicly available to facilitate calibration and validation of IQA algorithms.
Confidence-guided Lesion Mask-based Simultaneous Synthesis of Anatomic and Molecular MR Images in Patients with Post-treatment Malignant Gliomas
Aug 06, 2020Abstract:Data-driven automatic approaches have demonstrated their great potential in resolving various clinical diagnostic dilemmas in neuro-oncology, especially with the help of standard anatomic and advanced molecular MR images. However, data quantity and quality remain a key determinant of, and a significant limit on, the potential of such applications. In our previous work, we explored synthesis of anatomic and molecular MR image network (SAMR) in patients with post-treatment malignant glioms. Now, we extend it and propose Confidence Guided SAMR (CG-SAMR) that synthesizes data from lesion information to multi-modal anatomic sequences, including T1-weighted (T1w), gadolinium enhanced T1w (Gd-T1w), T2-weighted (T2w), and fluid-attenuated inversion recovery (FLAIR), and the molecular amide proton transfer-weighted (APTw) sequence. We introduce a module which guides the synthesis based on confidence measure about the intermediate results. Furthermore, we extend the proposed architecture for unsupervised synthesis so that unpaired data can be used for training the network. Extensive experiments on real clinical data demonstrate that the proposed model can perform better than the state-of-theart synthesis methods.
Multi-person 3D Pose Estimation in Crowded Scenes Based on Multi-View Geometry
Jul 21, 2020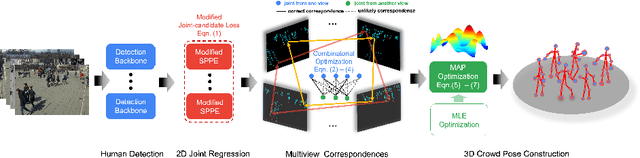
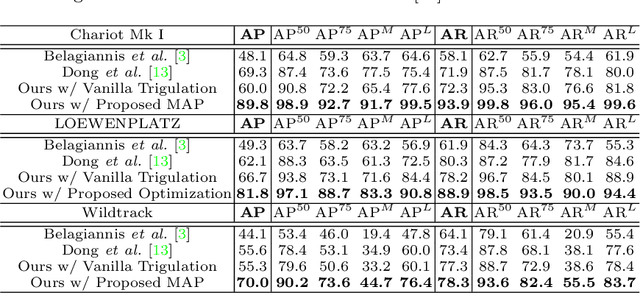
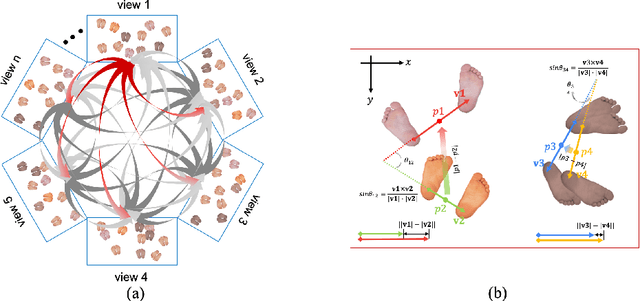
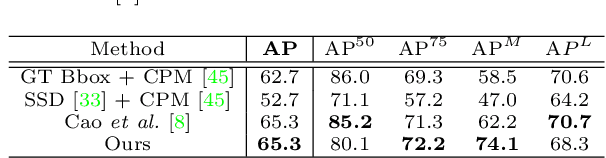
Abstract:Epipolar constraints are at the core of feature matching and depth estimation in current multi-person multi-camera 3D human pose estimation methods. Despite the satisfactory performance of this formulation in sparser crowd scenes, its effectiveness is frequently challenged under denser crowd circumstances mainly due to two sources of ambiguity. The first is the mismatch of human joints resulting from the simple cues provided by the Euclidean distances between joints and epipolar lines. The second is the lack of robustness from the naive formulation of the problem as a least squares minimization. In this paper, we depart from the multi-person 3D pose estimation formulation, and instead reformulate it as crowd pose estimation. Our method consists of two key components: a graph model for fast cross-view matching, and a maximum a posteriori (MAP) estimator for the reconstruction of the 3D human poses. We demonstrate the effectiveness and superiority of our proposed method on four benchmark datasets.
Lesion Mask-based Simultaneous Synthesis of Anatomic and MolecularMR Images using a GAN
Jul 05, 2020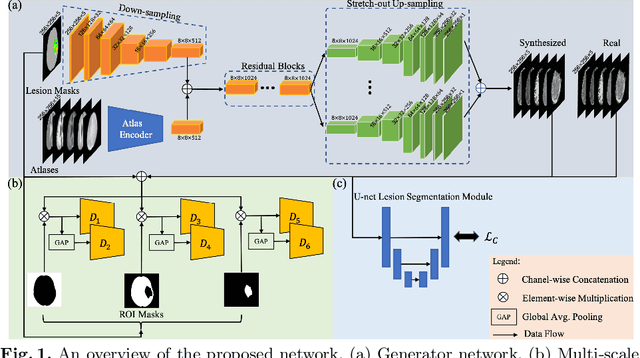

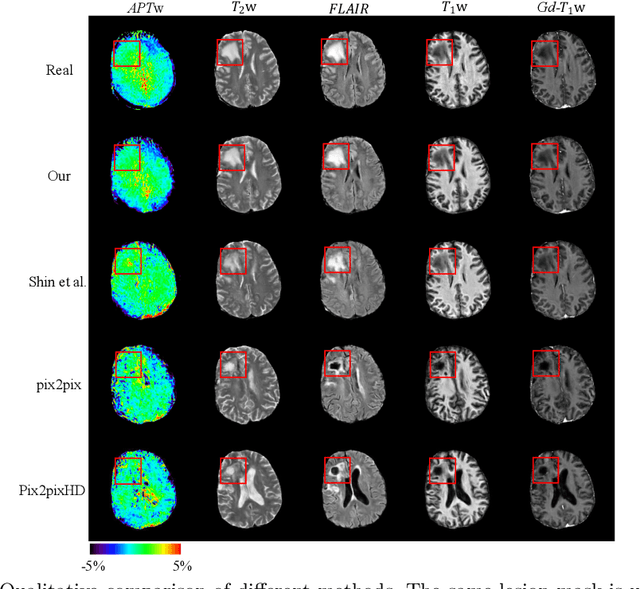
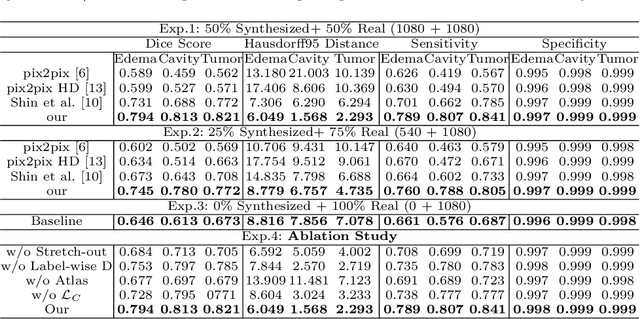
Abstract:Data-driven automatic approaches have demonstrated their great potential in resolving various clinical diagnostic dilemmas for patients with malignant gliomas in neuro-oncology with the help of conventional and advanced molecular MR images. However, the lack of sufficient annotated MRI data has vastly impeded the development of such automatic methods. Conventional data augmentation approaches, including flipping, scaling, rotation, and distortion are not capable of generating data with diverse image content. In this paper, we propose a generative adversarial network (GAN), which can simultaneously synthesize data from arbitrary manipulated lesion information on multiple anatomic and molecular MRI sequences, including T1-weighted (T1w), gadolinium enhanced T1w (Gd-T1w), T2-weighted (T2w), fluid-attenuated inversion recovery (FLAIR), and amide proton transfer-weighted (APTw). The proposed framework consists of a stretch-out up-sampling module, a brain atlas encoder, a segmentation consistency module, and multi-scale labelwise discriminators. Extensive experiments on real clinical data demonstrate that the proposed model can perform significantly better than the state-of-the-art synthesis methods.
Customized OCT images compression scheme with deep neural network
Aug 27, 2019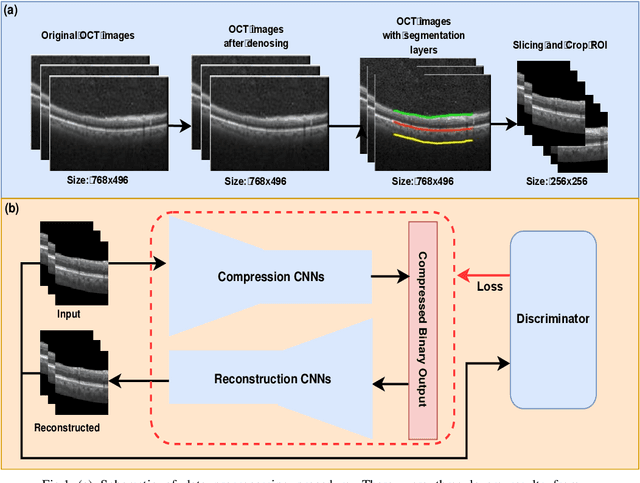
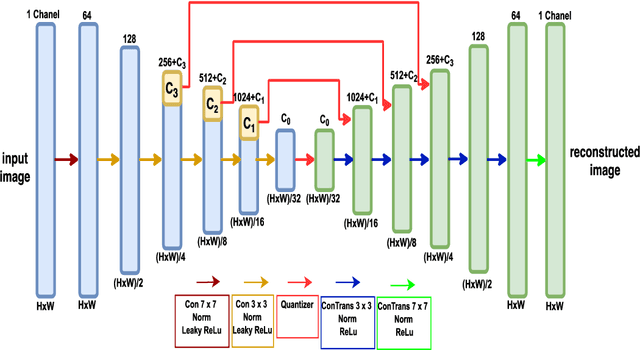
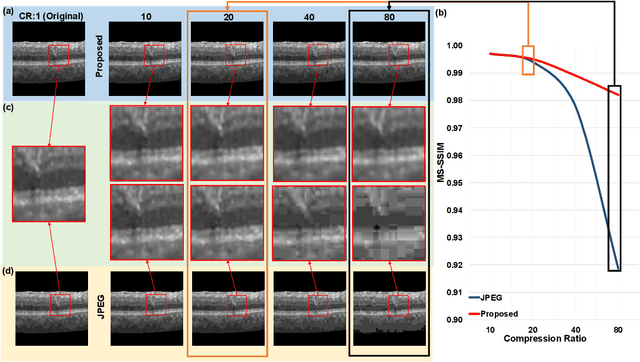
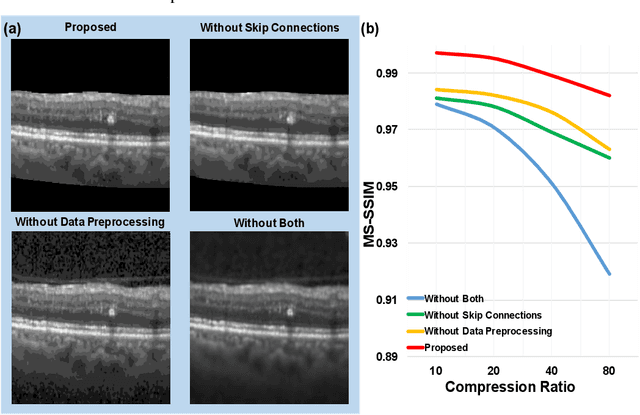
Abstract:We customize an end-to-end image compression framework for retina OCT images based on deep convolutional neural networks (CNNs). The customized compression scheme consists of three parts: data Preprocessing, compression CNNs, and reconstruction CNNs. Data preprocessing module reduces the speckle noise of the OCT images and the segments out the region of interest. We added customized skip connections between the compression CNNs and the reconstruction CNNs to reserve the detail information and trained the two nets together with the semantic segmented image patches from data preprocessing module. To train the two networks sensitive to both low frequency information and high frequency information, we adopted an objective function with two parts: A PatchGAN discriminator to judge the high frequency information and a differentiable MS-SSIM penalty to evaluate the low frequency information. The proposed framework was trained and evaluated on a publicly available OCT dataset. The evaluation showed above 99% similarity in terms of multi-scale structural similarity (MS-SSIM) when the compression ratio is as high as 40. Furthermore, the reconstructed images of compression ratio 80 from the proposed framework even have better quality than that of compression ratio 20 from JPEG by visual comparison. The testing result outperforms JPEG in term of both of MS-SSIM and visualization, which is more obvious as the increase of compression ratio. Our preliminary result indicates the huge potential of deep neural networks on customized medical image compression.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge