Oula Puonti
Danish Research Centre for Magnetic Resonance, Copenhagen University Hospital Hvidovre, Denmark
Conditional diffusion models for guided anomaly detection in brain images using fluid-driven anomaly randomization
Jun 11, 2025Abstract:Supervised machine learning has enabled accurate pathology detection in brain MRI, but requires training data from diseased subjects that may not be readily available in some scenarios, for example, in the case of rare diseases. Reconstruction-based unsupervised anomaly detection, in particular using diffusion models, has gained popularity in the medical field as it allows for training on healthy images alone, eliminating the need for large disease-specific cohorts. These methods assume that a model trained on normal data cannot accurately represent or reconstruct anomalies. However, this assumption often fails with models failing to reconstruct healthy tissue or accurately reconstruct abnormal regions i.e., failing to remove anomalies. In this work, we introduce a novel conditional diffusion model framework for anomaly detection and healthy image reconstruction in brain MRI. Our weakly supervised approach integrates synthetically generated pseudo-pathology images into the modeling process to better guide the reconstruction of healthy images. To generate these pseudo-pathologies, we apply fluid-driven anomaly randomization to augment real pathology segmentation maps from an auxiliary dataset, ensuring that the synthetic anomalies are both realistic and anatomically coherent. We evaluate our model's ability to detect pathology, using both synthetic anomaly datasets and real pathology from the ATLAS dataset. In our extensive experiments, our model: (i) consistently outperforms variational autoencoders, and conditional and unconditional latent diffusion; and (ii) surpasses on most datasets, the performance of supervised inpainting methods with access to paired diseased/healthy images.
Scalable Segmentation for Ultra-High-Resolution Brain MR Images
May 27, 2025Abstract:Although deep learning has shown great success in 3D brain MRI segmentation, achieving accurate and efficient segmentation of ultra-high-resolution brain images remains challenging due to the lack of labeled training data for fine-scale anatomical structures and high computational demands. In this work, we propose a novel framework that leverages easily accessible, low-resolution coarse labels as spatial references and guidance, without incurring additional annotation cost. Instead of directly predicting discrete segmentation maps, our approach regresses per-class signed distance transform maps, enabling smooth, boundary-aware supervision. Furthermore, to enhance scalability, generalizability, and efficiency, we introduce a scalable class-conditional segmentation strategy, where the model learns to segment one class at a time conditioned on a class-specific input. This novel design not only reduces memory consumption during both training and testing, but also allows the model to generalize to unseen anatomical classes. We validate our method through comprehensive experiments on both synthetic and real-world datasets, demonstrating its superior performance and scalability compared to conventional segmentation approaches.
End-to-end Cortical Surface Reconstruction from Clinical Magnetic Resonance Images
May 20, 2025Abstract:Surface-based cortical analysis is valuable for a variety of neuroimaging tasks, such as spatial normalization, parcellation, and gray matter (GM) thickness estimation. However, most tools for estimating cortical surfaces work exclusively on scans with at least 1 mm isotropic resolution and are tuned to a specific magnetic resonance (MR) contrast, often T1-weighted (T1w). This precludes application using most clinical MR scans, which are very heterogeneous in terms of contrast and resolution. Here, we use synthetic domain-randomized data to train the first neural network for explicit estimation of cortical surfaces from scans of any contrast and resolution, without retraining. Our method deforms a template mesh to the white matter (WM) surface, which guarantees topological correctness. This mesh is further deformed to estimate the GM surface. We compare our method to recon-all-clinical (RAC), an implicit surface reconstruction method which is currently the only other tool capable of processing heterogeneous clinical MR scans, on ADNI and a large clinical dataset (n=1,332). We show a approximately 50 % reduction in cortical thickness error (from 0.50 to 0.24 mm) with respect to RAC and better recovery of the aging-related cortical thinning patterns detected by FreeSurfer on high-resolution T1w scans. Our method enables fast and accurate surface reconstruction of clinical scans, allowing studies (1) with sample sizes far beyond what is feasible in a research setting, and (2) of clinical populations that are difficult to enroll in research studies. The code is publicly available at https://github.com/simnibs/brainnet.
From Low Field to High Value: Robust Cortical Mapping from Low-Field MRI
May 18, 2025Abstract:Three-dimensional reconstruction of cortical surfaces from MRI for morphometric analysis is fundamental for understanding brain structure. While high-field MRI (HF-MRI) is standard in research and clinical settings, its limited availability hinders widespread use. Low-field MRI (LF-MRI), particularly portable systems, offers a cost-effective and accessible alternative. However, existing cortical surface analysis tools are optimized for high-resolution HF-MRI and struggle with the lower signal-to-noise ratio and resolution of LF-MRI. In this work, we present a machine learning method for 3D reconstruction and analysis of portable LF-MRI across a range of contrasts and resolutions. Our method works "out of the box" without retraining. It uses a 3D U-Net trained on synthetic LF-MRI to predict signed distance functions of cortical surfaces, followed by geometric processing to ensure topological accuracy. We evaluate our method using paired HF/LF-MRI scans of the same subjects, showing that LF-MRI surface reconstruction accuracy depends on acquisition parameters, including contrast type (T1 vs T2), orientation (axial vs isotropic), and resolution. A 3mm isotropic T2-weighted scan acquired in under 4 minutes, yields strong agreement with HF-derived surfaces: surface area correlates at r=0.96, cortical parcellations reach Dice=0.98, and gray matter volume achieves r=0.93. Cortical thickness remains more challenging with correlations up to r=0.70, reflecting the difficulty of sub-mm precision with 3mm voxels. We further validate our method on challenging postmortem LF-MRI, demonstrating its robustness. Our method represents a step toward enabling cortical surface analysis on portable LF-MRI. Code is available at https://surfer.nmr.mgh.harvard.edu/fswiki/ReconAny
Reference-Free 3D Reconstruction of Brain Dissection Photographs with Machine Learning
Mar 13, 2025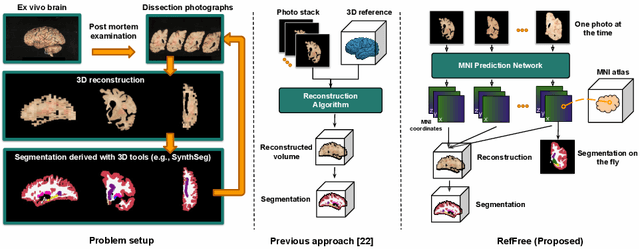
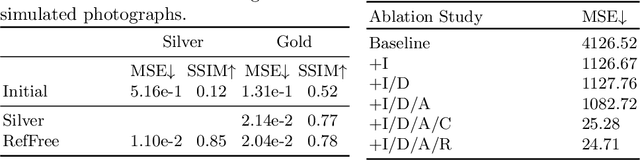
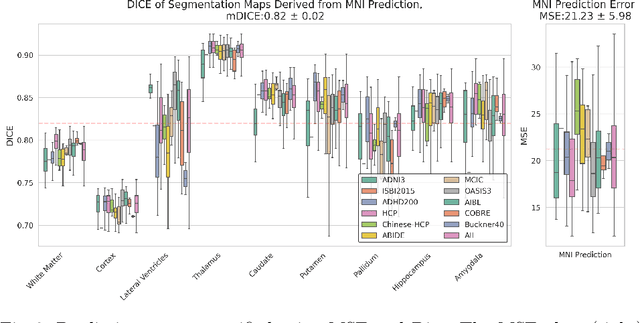
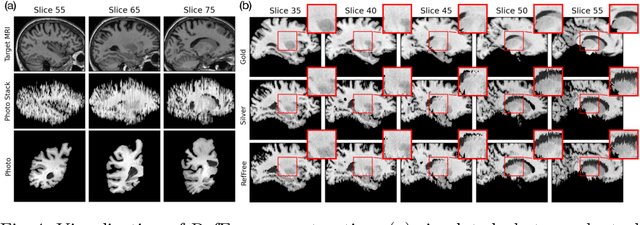
Abstract:Correlation of neuropathology with MRI has the potential to transfer microscopic signatures of pathology to invivo scans. Recently, a classical registration method has been proposed, to build these correlations from 3D reconstructed stacks of dissection photographs, which are routinely taken at brain banks. These photographs bypass the need for exvivo MRI, which is not widely accessible. However, this method requires a full stack of brain slabs and a reference mask (e.g., acquired with a surface scanner), which severely limits the applicability of the technique. Here we propose RefFree, a dissection photograph reconstruction method without external reference. RefFree is a learning approach that estimates the 3D coordinates in the atlas space for every pixel in every photograph; simple least-squares fitting can then be used to compute the 3D reconstruction. As a by-product, RefFree also produces an atlas-based segmentation of the reconstructed stack. RefFree is trained on synthetic photographs generated from digitally sliced 3D MRI data, with randomized appearance for enhanced generalization ability. Experiments on simulated and real data show that RefFree achieves performance comparable to the baseline method without an explicit reference while also enabling reconstruction of partial stacks. Our code is available at https://github.com/lintian-a/reffree.
Learn2Synth: Learning Optimal Data Synthesis Using Hypergradients
Nov 23, 2024



Abstract:Domain randomization through synthesis is a powerful strategy to train networks that are unbiased with respect to the domain of the input images. Randomization allows networks to see a virtually infinite range of intensities and artifacts during training, thereby minimizing overfitting to appearance and maximizing generalization to unseen data. While powerful, this approach relies on the accurate tuning of a large set of hyper-parameters governing the probabilistic distribution of the synthesized images. Instead of manually tuning these parameters, we introduce Learn2Synth, a novel procedure in which synthesis parameters are learned using a small set of real labeled data. Unlike methods that impose constraints to align synthetic data with real data (e.g., contrastive or adversarial techniques), which risk misaligning the image and its label map, we tune an augmentation engine such that a segmentation network trained on synthetic data has optimal accuracy when applied to real data. This approach allows the training procedure to benefit from real labeled examples, without ever using these real examples to train the segmentation network, which avoids biasing the network towards the properties of the training set. Specifically, we develop both parametric and nonparametric strategies to augment the synthetic images, enhancing the segmentation network's performance. Experimental results on both synthetic and real-world datasets demonstrate the effectiveness of this learning strategy. Code is available at: https://github.com/HuXiaoling/Learn2Synth.
Recon-all-clinical: Cortical surface reconstruction and analysis of heterogeneous clinical brain MRI
Sep 05, 2024



Abstract:Surface-based analysis of the cerebral cortex is ubiquitous in human neuroimaging with MRI. It is crucial for cortical registration, parcellation, and thickness estimation. Traditionally, these analyses require high-resolution, isotropic scans with good gray-white matter contrast, typically a 1mm T1-weighted scan. This excludes most clinical MRI scans, which are often anisotropic and lack the necessary T1 contrast. To enable large-scale neuroimaging studies using vast clinical data, we introduce recon-all-clinical, a novel method for cortical reconstruction, registration, parcellation, and thickness estimation in brain MRI scans of any resolution and contrast. Our approach employs a hybrid analysis method that combines a convolutional neural network (CNN) trained with domain randomization to predict signed distance functions (SDFs) and classical geometry processing for accurate surface placement while maintaining topological and geometric constraints. The method does not require retraining for different acquisitions, thus simplifying the analysis of heterogeneous clinical datasets. We tested recon-all-clinical on multiple datasets, including over 19,000 clinical scans. The method consistently produced precise cortical reconstructions and high parcellation accuracy across varied MRI contrasts and resolutions. Cortical thickness estimates are precise enough to capture aging effects independently of MRI contrast, although accuracy varies with slice thickness. Our method is publicly available at https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all-clinical, enabling researchers to perform detailed cortical analysis on the huge amounts of already existing clinical MRI scans. This advancement may be particularly valuable for studying rare diseases and underrepresented populations where research-grade MRI data is scarce.
High-resolution segmentations of the hypothalamus and its subregions for training of segmentation models
Jun 27, 2024Abstract:Segmentation of brain structures on magnetic resonance imaging (MRI) is a highly relevant neuroimaging topic, as it is a prerequisite for different analyses such as volumetry or shape analysis. Automated segmentation facilitates the study of brain structures in larger cohorts when compared with manual segmentation, which is time-consuming. However, the development of most automated methods relies on large and manually annotated datasets, which limits the generalizability of these methods. Recently, new techniques using synthetic images have emerged, reducing the need for manual annotation. Here we provide HELM, Hypothalamic ex vivo Label Maps, a dataset composed of label maps built from publicly available ultra-high resolution ex vivo MRI from 10 whole hemispheres, which can be used to develop segmentation methods using synthetic data. The label maps are obtained with a combination of manual labels for the hypothalamic regions and automated segmentations for the rest of the brain, and mirrored to simulate entire brains. We also provide the pre-processed ex vivo scans, as this dataset can support future projects to include other structures after these are manually segmented.
Registration by Regression (RbR): a framework for interpretable and flexible atlas registration
Apr 25, 2024


Abstract:In human neuroimaging studies, atlas registration enables mapping MRI scans to a common coordinate frame, which is necessary to aggregate data from multiple subjects. Machine learning registration methods have achieved excellent speed and accuracy but lack interpretability. More recently, keypoint-based methods have been proposed to tackle this issue, but their accuracy is still subpar, particularly when fitting nonlinear transforms. Here we propose Registration by Regression (RbR), a novel atlas registration framework that is highly robust and flexible, conceptually simple, and can be trained with cheaply obtained data. RbR predicts the (x,y,z) atlas coordinates for every voxel of the input scan (i.e., every voxel is a keypoint), and then uses closed-form expressions to quickly fit transforms using a wide array of possible deformation models, including affine and nonlinear (e.g., Bspline, Demons, invertible diffeomorphic models, etc.). Robustness is provided by the large number of voxels informing the registration and can be further increased by robust estimators like RANSAC. Experiments on independent public datasets show that RbR yields more accurate registration than competing keypoint approaches, while providing full control of the deformation model.
P-Count: Persistence-based Counting of White Matter Hyperintensities in Brain MRI
Mar 20, 2024



Abstract:White matter hyperintensities (WMH) are a hallmark of cerebrovascular disease and multiple sclerosis. Automated WMH segmentation methods enable quantitative analysis via estimation of total lesion load, spatial distribution of lesions, and number of lesions (i.e., number of connected components after thresholding), all of which are correlated with patient outcomes. While the two former measures can generally be estimated robustly, the number of lesions is highly sensitive to noise and segmentation mistakes -- even when small connected components are eroded or disregarded. In this article, we present P-Count, an algebraic WMH counting tool based on persistent homology that accounts for the topological features of WM lesions in a robust manner. Using computational geometry, P-Count takes the persistence of connected components into consideration, effectively filtering out the noisy WMH positives, resulting in a more accurate count of true lesions. We validated P-Count on the ISBI2015 longitudinal lesion segmentation dataset, where it produces significantly more accurate results than direct thresholding.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge