Ni Ou
Native-Domain Cross-Attention for Camera-LiDAR Extrinsic Calibration Under Large Initial Perturbations
Mar 31, 2026Abstract:Accurate camera-LiDAR fusion relies on precise extrinsic calibration, which fundamentally depends on establishing reliable cross-modal correspondences under potentially large misalignments. Existing learning-based methods typically project LiDAR points into depth maps for feature fusion, which distorts 3D geometry and degrades performance when the extrinsic initialization is far from the ground truth. To address this issue, we propose an extrinsic-aware cross-attention framework that directly aligns image patches and LiDAR point groups in their native domains. The proposed attention mechanism explicitly injects extrinsic parameter hypotheses into the correspondence modeling process, enabling geometry-consistent cross-modal interaction without relying on projected 2D depth maps. Extensive experiments on the KITTI and nuScenes benchmarks demonstrate that our method consistently outperforms state-of-the-art approaches in both accuracy and robustness. Under large extrinsic perturbations, our approach achieves accurate calibration in 88% of KITTI cases and 99% of nuScenes cases, substantially surpassing the second-best baseline. We have open sourced our code on https://github.com/gitouni/ProjFusion to benefit the community.
General Force Sensation for Tactile Robot
Mar 02, 2025



Abstract:Robotic tactile sensors, including vision-based and taxel-based sensors, enable agile manipulation and safe human-robot interaction through force sensation. However, variations in structural configurations, measured signals, and material properties create domain gaps that limit the transferability of learned force sensation across different tactile sensors. Here, we introduce GenForce, a general framework for achieving transferable force sensation across both homogeneous and heterogeneous tactile sensors in robotic systems. By unifying tactile signals into marker-based binary tactile images, GenForce enables the transfer of existing force labels to arbitrary target sensors using a marker-to-marker translation technique with a few paired data. This process equips uncalibrated tactile sensors with force prediction capabilities through spatiotemporal force prediction models trained on the transferred data. Extensive experimental results validate GenForce's generalizability, accuracy, and robustness across sensors with diverse marker patterns, structural designs, material properties, and sensing principles. The framework significantly reduces the need for costly and labor-intensive labeled data collection, enabling the rapid deployment of multiple tactile sensors on robotic hands requiring force sensing capabilities.
Iterative Camera-LiDAR Extrinsic Optimization via Surrogate Diffusion
Nov 17, 2024



Abstract:Cameras and LiDAR are essential sensors for autonomous vehicles. Camera-LiDAR data fusion compensate for deficiencies of stand-alone sensors but relies on precise extrinsic calibration. Many learning-based calibration methods predict extrinsic parameters in a single step. Driven by the growing demand for higher accuracy, a few approaches utilize multi-range models or integrate multiple methods to improve extrinsic parameter predictions, but these strategies incur extended training times and require additional storage for separate models. To address these issues, we propose a single-model iterative approach based on surrogate diffusion to significantly enhance the capacity of individual calibration methods. By applying a buffering technique proposed by us, the inference time of our surrogate diffusion is 43.7% less than that of multi-range models. Additionally, we create a calibration network as our denoiser, featuring both projection-first and encoding-first branches for effective point feature extraction. Extensive experiments demonstrate that our diffusion model outperforms other single-model iterative methods and delivers competitive results compared to multi-range models. Our denoiser exceeds state-of-the-art calibration methods, reducing the rotation error by 24.5% compared to the second-best method. Furthermore, with the proposed diffusion applied, it achieves 20.4% less rotation error and 9.6% less translation error.
TransForce: Transferable Force Prediction for Vision-based Tactile Sensors with Sequential Image Translation
Sep 15, 2024



Abstract:Vision-based tactile sensors (VBTSs) provide high-resolution tactile images crucial for robot in-hand manipulation. However, force sensing in VBTSs is underutilized due to the costly and time-intensive process of acquiring paired tactile images and force labels. In this study, we introduce a transferable force prediction model, TransForce, designed to leverage collected image-force paired data for new sensors under varying illumination colors and marker patterns while improving the accuracy of predicted forces, especially in the shear direction. Our model effectively achieves translation of tactile images from the source domain to the target domain, ensuring that the generated tactile images reflect the illumination colors and marker patterns of the new sensors while accurately aligning the elastomer deformation observed in existing sensors, which is beneficial to force prediction of new sensors. As such, a recurrent force prediction model trained with generated sequential tactile images and existing force labels is employed to estimate higher-accuracy forces for new sensors with lowest average errors of 0.69N (5.8\% in full work range) in $x$-axis, 0.70N (5.8\%) in $y$-axis, and 1.11N (6.9\%) in $z$-axis compared with models trained with single images. The experimental results also reveal that pure marker modality is more helpful than the RGB modality in improving the accuracy of force in the shear direction, while the RGB modality show better performance in the normal direction.
Marker or Markerless? Mode-Switchable Optical Tactile Sensing for Diverse Robot Tasks
Aug 15, 2024Abstract:Optical tactile sensors play a pivotal role in robot perception and manipulation tasks. The membrane of these sensors can be painted with markers or remain markerless, enabling them to function in either marker or markerless mode. However, this uni-modal selection means the sensor is only suitable for either manipulation or perception tasks. While markers are vital for manipulation, they can also obstruct the camera, thereby impeding perception. The dilemma of selecting between marker and markerless modes presents a significant obstacle. To address this issue, we propose a novel mode-switchable optical tactile sensing approach that facilitates transitions between the two modes. The marker-to-markerless transition is achieved through a generative model, whereas its inverse transition is realized using a sparsely supervised regressive model. Our approach allows a single-mode optical sensor to operate effectively in both marker and markerless modes without the need for additional hardware, making it well-suited for both perception and manipulation tasks. Extensive experiments validate the effectiveness of our method. For perception tasks, our approach decreases the number of categories that include misclassified samples by 2 and improves contact area segmentation IoU by 3.53%. For manipulation tasks, our method attains a high success rate of 92.59% in slip detection. Code, dataset and demo videos are available at the project website: https://gitouni.github.io/Marker-Markerless-Transition/
Deep Domain Adaptation Regression for Force Calibration of Optical Tactile Sensors
Jul 19, 2024Abstract:Optical tactile sensors provide robots with rich force information for robot grasping in unstructured environments. The fast and accurate calibration of three-dimensional contact forces holds significance for new sensors and existing tactile sensors which may have incurred damage or aging. However, the conventional neural-network-based force calibration method necessitates a large volume of force-labeled tactile images to minimize force prediction errors, with the need for accurate Force/Torque measurement tools as well as a time-consuming data collection process. To address this challenge, we propose a novel deep domain-adaptation force calibration method, designed to transfer the force prediction ability from a calibrated optical tactile sensor to uncalibrated ones with various combinations of domain gaps, including marker presence, illumination condition, and elastomer modulus. Experimental results show the effectiveness of the proposed unsupervised force calibration method, with lowest force prediction errors of 0.102N (3.4\% in full force range) for normal force, and 0.095N (6.3\%) and 0.062N (4.1\%) for shear forces along the x-axis and y-axis, respectively. This study presents a promising, general force calibration methodology for optical tactile sensors.
A Foundation Model for Brain Lesion Segmentation with Mixture of Modality Experts
May 16, 2024



Abstract:Brain lesion segmentation plays an essential role in neurological research and diagnosis. As brain lesions can be caused by various pathological alterations, different types of brain lesions tend to manifest with different characteristics on different imaging modalities. Due to this complexity, brain lesion segmentation methods are often developed in a task-specific manner. A specific segmentation model is developed for a particular lesion type and imaging modality. However, the use of task-specific models requires predetermination of the lesion type and imaging modality, which complicates their deployment in real-world scenarios. In this work, we propose a universal foundation model for 3D brain lesion segmentation, which can automatically segment different types of brain lesions for input data of various imaging modalities. We formulate a novel Mixture of Modality Experts (MoME) framework with multiple expert networks attending to different imaging modalities. A hierarchical gating network combines the expert predictions and fosters expertise collaboration. Furthermore, we introduce a curriculum learning strategy during training to avoid the degeneration of each expert network and preserve their specialization. We evaluated the proposed method on nine brain lesion datasets, encompassing five imaging modalities and eight lesion types. The results show that our model outperforms state-of-the-art universal models and provides promising generalization to unseen datasets.
Visual-LiDAR Odometry and Mapping with Monocular Scale Correction and Motion Compensation
Apr 18, 2023



Abstract:This paper presents a novel visual-LiDAR odometry and mapping method with low-drift characteristics. The proposed method is based on two popular approaches, ORB-SLAM and A-LOAM, with monocular scale correction and visual-assisted LiDAR motion compensation modifications. The scale corrector calculates the proportion between the depth of image keypoints recovered by triangulation and that provided by LiDAR, using an outlier rejection process for accuracy improvement. Concerning LiDAR motion compensation, the visual odometry approach gives the initial guesses of LiDAR motions for better performance. This methodology is not only applicable to high-resolution LiDAR but can also adapt to low-resolution LiDAR. To evaluate the proposed SLAM system's robustness and accuracy, we conducted experiments on the KITTI Odometry and S3E datasets. Experimental results illustrate that our method significantly outperforms standalone ORB-SLAM2 and A-LOAM. Furthermore, regarding the accuracy of visual odometry with scale correction, our method performs similarly to the stereo-mode ORB-SLAM2.
Unsupervised Brain Tumor Segmentation with Image-based Prompts
Apr 04, 2023



Abstract:Automated brain tumor segmentation based on deep learning (DL) has achieved promising performance. However, it generally relies on annotated images for model training, which is not always feasible in clinical settings. Therefore, the development of unsupervised DL-based brain tumor segmentation approaches without expert annotations is desired. Motivated by the success of prompt learning (PL) in natural language processing, we propose an approach to unsupervised brain tumor segmentation by designing image-based prompts that allow indication of brain tumors, and this approach is dubbed as PL-based Brain Tumor Segmentation (PL-BTS). Specifically, instead of directly training a model for brain tumor segmentation with a large amount of annotated data, we seek to train a model that can answer the question: is a voxel in the input image associated with tumor-like hyper-/hypo-intensity? Such a model can be trained by artificially generating tumor-like hyper-/hypo-intensity on images without tumors with hand-crafted designs. Since the hand-crafted designs may be too simplistic to represent all kinds of real tumors, the trained model may overfit the simplistic hand-crafted task rather than actually answer the question of abnormality. To address this problem, we propose the use of a validation task, where we generate a different hand-crafted task to monitor overfitting. In addition, we propose PL-BTS+ that further improves PL-BTS by exploiting unannotated images with brain tumors. Compared with competing unsupervised methods, the proposed method has achieved marked improvements on both public and in-house datasets, and we have also demonstrated its possible extension to other brain lesion segmentation tasks.
CarveMix: A Simple Data Augmentation Method for Brain Lesion Segmentation
Aug 17, 2021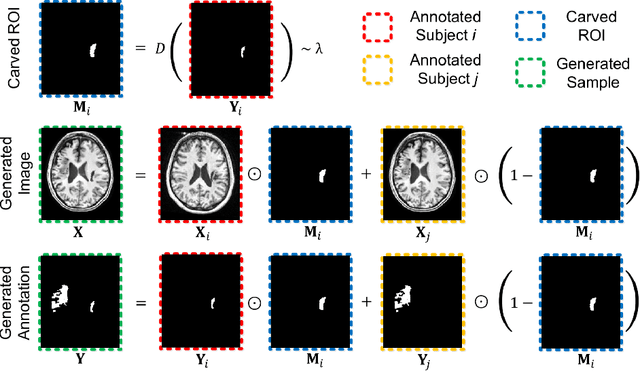
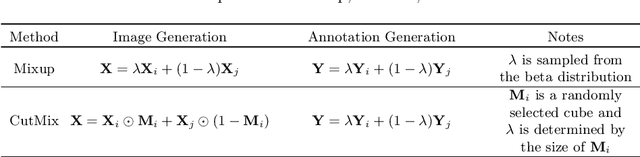

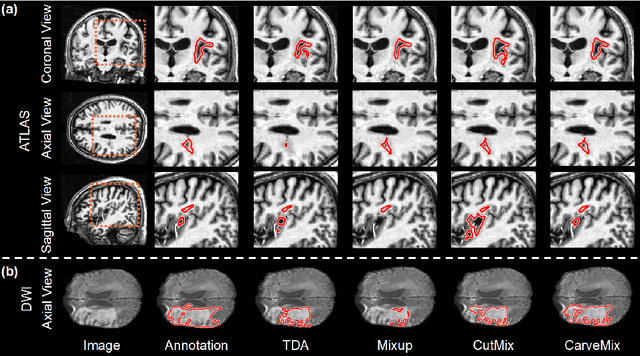
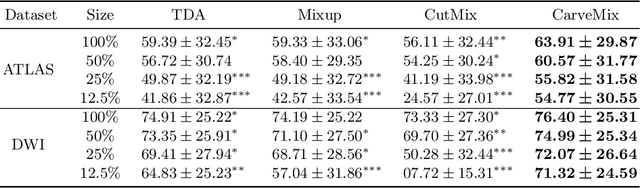
Abstract:Brain lesion segmentation provides a valuable tool for clinical diagnosis, and convolutional neural networks (CNNs) have achieved unprecedented success in the task. Data augmentation is a widely used strategy that improves the training of CNNs, and the design of the augmentation method for brain lesion segmentation is still an open problem. In this work, we propose a simple data augmentation approach, dubbed as CarveMix, for CNN-based brain lesion segmentation. Like other "mix"-based methods, such as Mixup and CutMix, CarveMix stochastically combines two existing labeled images to generate new labeled samples. Yet, unlike these augmentation strategies based on image combination, CarveMix is lesion-aware, where the combination is performed with an attention on the lesions and a proper annotation is created for the generated image. Specifically, from one labeled image we carve a region of interest (ROI) according to the lesion location and geometry, and the size of the ROI is sampled from a probability distribution. The carved ROI then replaces the corresponding voxels in a second labeled image, and the annotation of the second image is replaced accordingly as well. In this way, we generate new labeled images for network training and the lesion information is preserved. To evaluate the proposed method, experiments were performed on two brain lesion datasets. The results show that our method improves the segmentation accuracy compared with other simple data augmentation approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge