Michael A. Riegler
PolypConnect: Image inpainting for generating realistic gastrointestinal tract images with polyps
May 30, 2022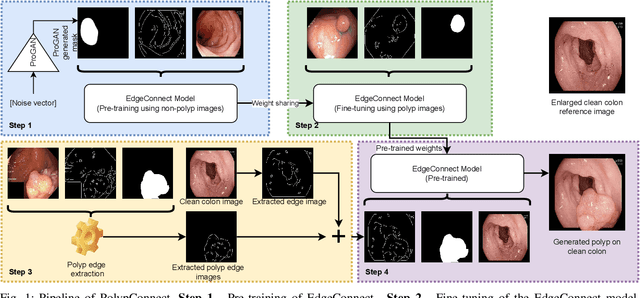
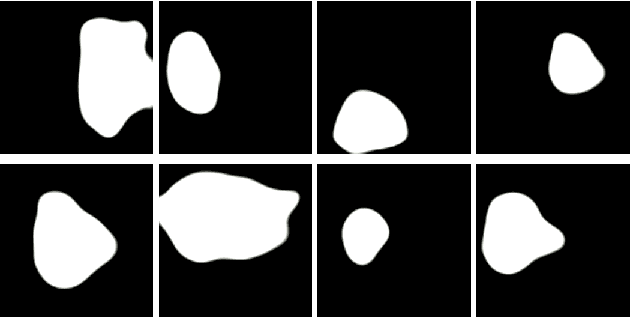
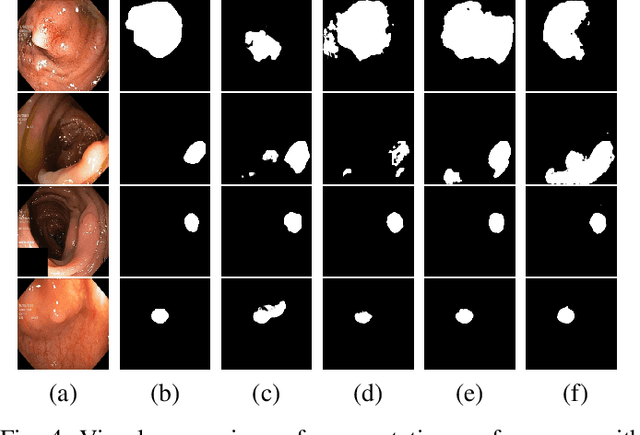



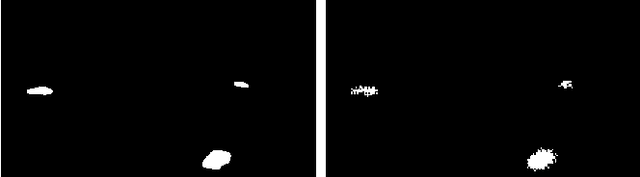
Abstract:Early identification of a polyp in the lower gastrointestinal (GI) tract can lead to prevention of life-threatening colorectal cancer. Developing computer-aided diagnosis (CAD) systems to detect polyps can improve detection accuracy and efficiency and save the time of the domain experts called endoscopists. Lack of annotated data is a common challenge when building CAD systems. Generating synthetic medical data is an active research area to overcome the problem of having relatively few true positive cases in the medical domain. To be able to efficiently train machine learning (ML) models, which are the core of CAD systems, a considerable amount of data should be used. In this respect, we propose the PolypConnect pipeline, which can convert non-polyp images into polyp images to increase the size of training datasets for training. We present the whole pipeline with quantitative and qualitative evaluations involving endoscopists. The polyp segmentation model trained using synthetic data, and real data shows a 5.1% improvement of mean intersection over union (mIOU), compared to the model trained only using real data. The codes of all the experiments are available on GitHub to reproduce the results.
Grid HTM: Hierarchical Temporal Memory for Anomaly Detection in Videos
May 30, 2022

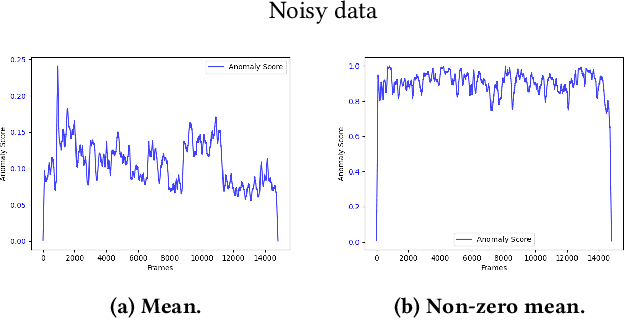
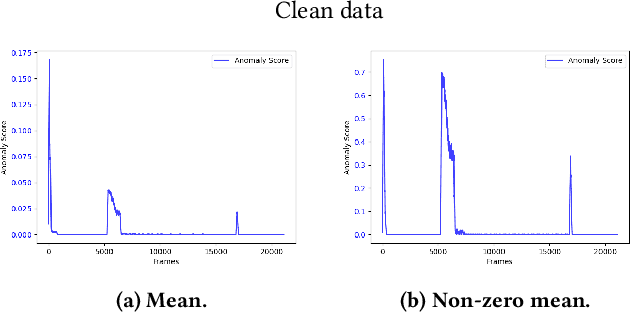
Abstract:The interest for video anomaly detection systems has gained traction for the past few years. The current approaches use deep learning to perform anomaly detection in videos, but this approach has multiple problems. For starters, deep learning in general has issues with noise, concept drift, explainability, and training data volumes. Additionally, anomaly detection in itself is a complex task and faces challenges such as unknowness, heterogeneity, and class imbalance. Anomaly detection using deep learning is therefore mainly constrained to generative models such as generative adversarial networks and autoencoders due to their unsupervised nature, but even they suffer from general deep learning issues and are hard to train properly. In this paper, we explore the capabilities of the Hierarchical Temporal Memory (HTM) algorithm to perform anomaly detection in videos, as it has favorable properties such as noise tolerance and online learning which combats concept drift. We introduce a novel version of HTM, namely, Grid HTM, which is an HTM-based architecture specifically for anomaly detection in complex videos such as surveillance footage.
Predicting tacrolimus exposure in kidney transplanted patients using machine learning
May 09, 2022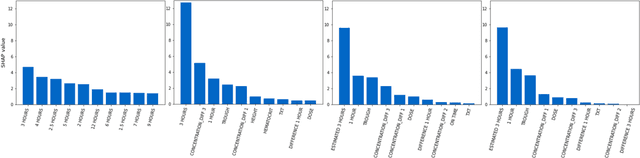
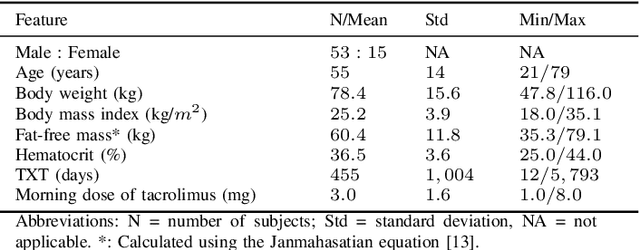
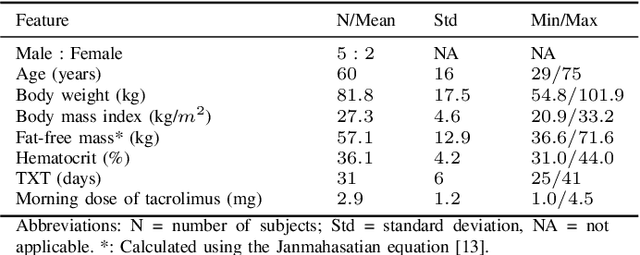
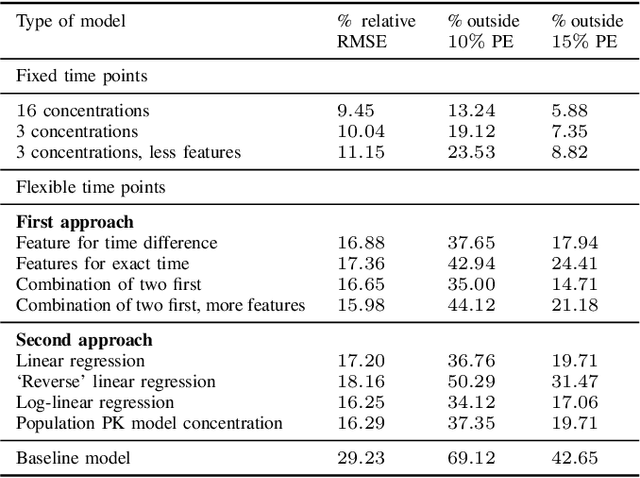
Abstract:Tacrolimus is one of the cornerstone immunosuppressive drugs in most transplantation centers worldwide following solid organ transplantation. Therapeutic drug monitoring of tacrolimus is necessary in order to avoid rejection of the transplanted organ or severe side effects. However, finding the right dose for a given patient is challenging, even for experienced clinicians. Consequently, a tool that can accurately estimate the drug exposure for individual dose adaptions would be of high clinical value. In this work, we propose a new technique using machine learning to estimate the tacrolimus exposure in kidney transplant recipients. Our models achieve predictive errors that are at the same level as an established population pharmacokinetic model, but are faster to develop and require less knowledge about the pharmacokinetic properties of the drug.
Assessing generalisability of deep learning-based polyp detection and segmentation methods through a computer vision challenge
Feb 24, 2022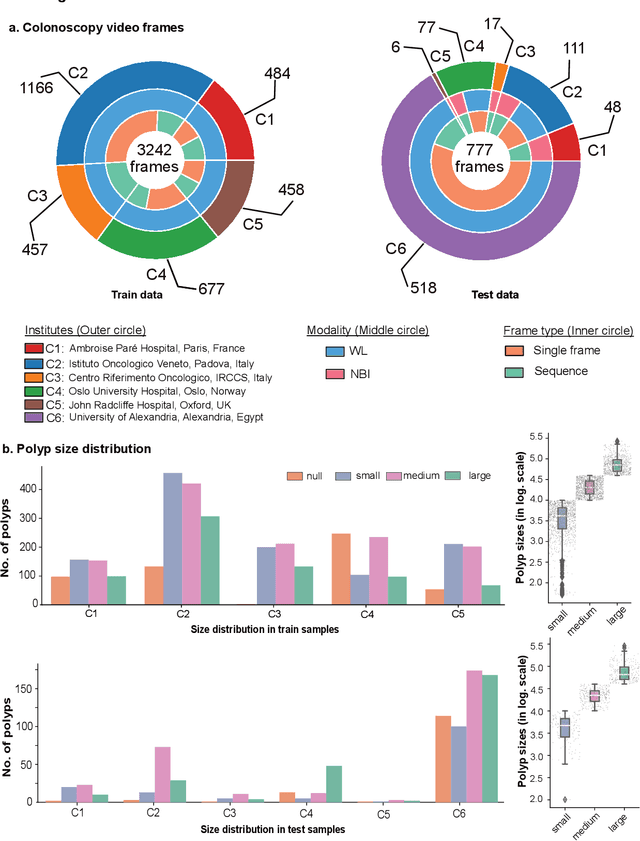
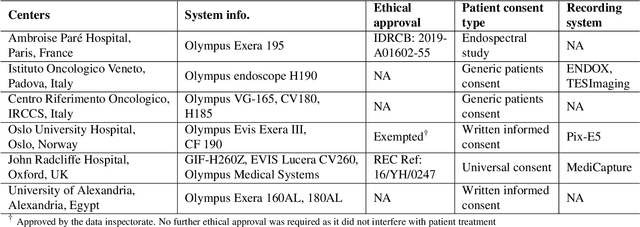
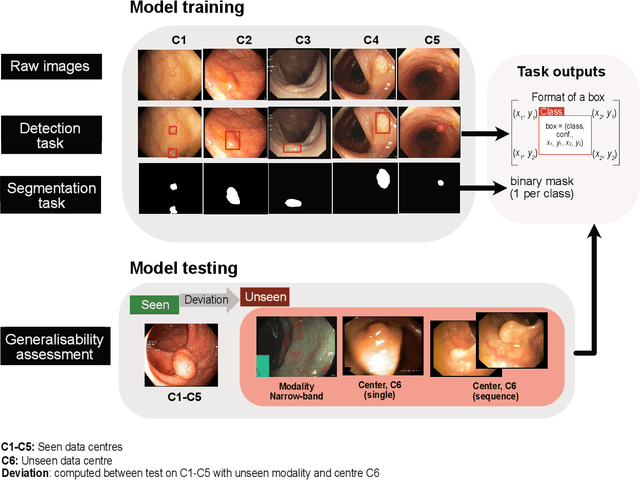
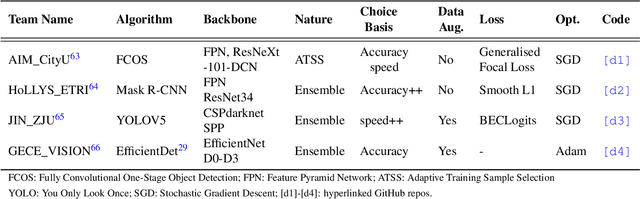
Abstract:Polyps are well-known cancer precursors identified by colonoscopy. However, variability in their size, location, and surface largely affect identification, localisation, and characterisation. Moreover, colonoscopic surveillance and removal of polyps (referred to as polypectomy ) are highly operator-dependent procedures. There exist a high missed detection rate and incomplete removal of colonic polyps due to their variable nature, the difficulties to delineate the abnormality, the high recurrence rates, and the anatomical topography of the colon. There have been several developments in realising automated methods for both detection and segmentation of these polyps using machine learning. However, the major drawback in most of these methods is their ability to generalise to out-of-sample unseen datasets that come from different centres, modalities and acquisition systems. To test this hypothesis rigorously we curated a multi-centre and multi-population dataset acquired from multiple colonoscopy systems and challenged teams comprising machine learning experts to develop robust automated detection and segmentation methods as part of our crowd-sourcing Endoscopic computer vision challenge (EndoCV) 2021. In this paper, we analyse the detection results of the four top (among seven) teams and the segmentation results of the five top teams (among 16). Our analyses demonstrate that the top-ranking teams concentrated on accuracy (i.e., accuracy > 80% on overall Dice score on different validation sets) over real-time performance required for clinical applicability. We further dissect the methods and provide an experiment-based hypothesis that reveals the need for improved generalisability to tackle diversity present in multi-centre datasets.
Visual Sentiment Analysis: A Natural DisasterUse-case Task at MediaEval 2021
Nov 22, 2021Abstract:The Visual Sentiment Analysis task is being offered for the first time at MediaEval. The main purpose of the task is to predict the emotional response to images of natural disasters shared on social media. Disaster-related images are generally complex and often evoke an emotional response, making them an ideal use case of visual sentiment analysis. We believe being able to perform meaningful analysis of natural disaster-related data could be of great societal importance, and a joint effort in this regard can open several interesting directions for future research. The task is composed of three sub-tasks, each aiming to explore a different aspect of the challenge. In this paper, we provide a detailed overview of the task, the general motivation of the task, and an overview of the dataset and the metrics to be used for the evaluation of the proposed solutions.
PAANet: Progressive Alternating Attention for Automatic Medical Image Segmentation
Nov 20, 2021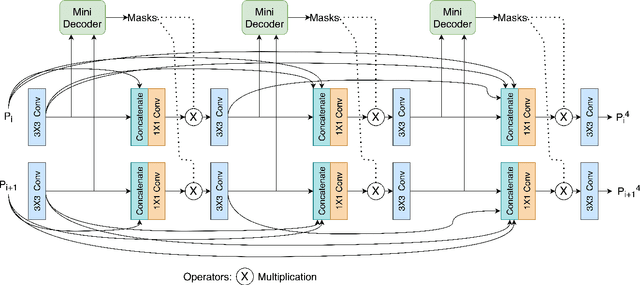
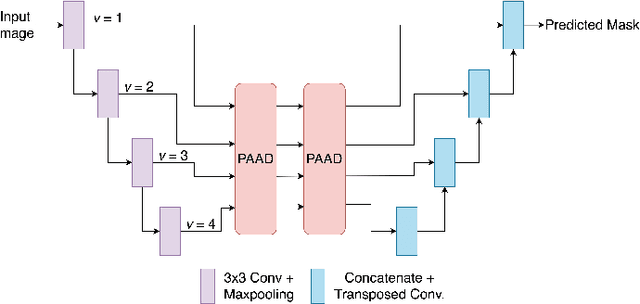
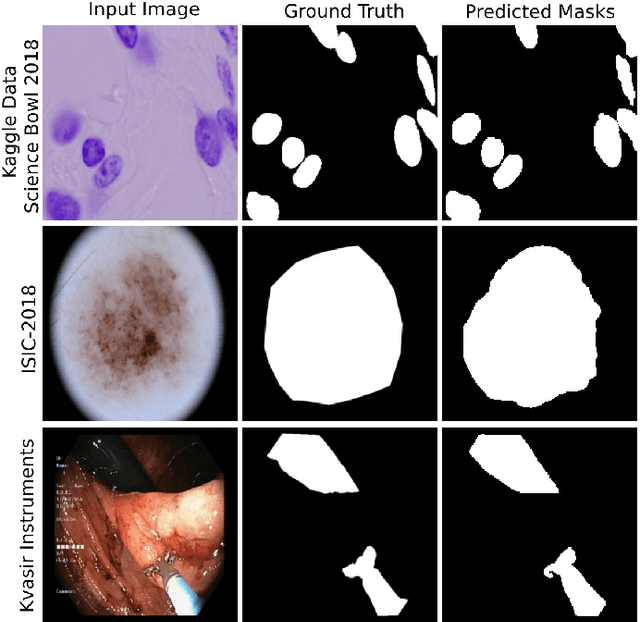
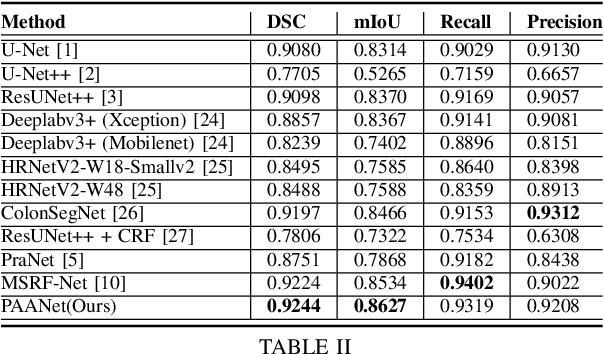
Abstract:Medical image segmentation can provide detailed information for clinical analysis which can be useful for scenarios where the detailed location of a finding is important. Knowing the location of disease can play a vital role in treatment and decision-making. Convolutional neural network (CNN) based encoder-decoder techniques have advanced the performance of automated medical image segmentation systems. Several such CNN-based methodologies utilize techniques such as spatial- and channel-wise attention to enhance performance. Another technique that has drawn attention in recent years is residual dense blocks (RDBs). The successive convolutional layers in densely connected blocks are capable of extracting diverse features with varied receptive fields and thus, enhancing performance. However, consecutive stacked convolutional operators may not necessarily generate features that facilitate the identification of the target structures. In this paper, we propose a progressive alternating attention network (PAANet). We develop progressive alternating attention dense (PAAD) blocks, which construct a guiding attention map (GAM) after every convolutional layer in the dense blocks using features from all scales. The GAM allows the following layers in the dense blocks to focus on the spatial locations relevant to the target region. Every alternate PAAD block inverts the GAM to generate a reverse attention map which guides ensuing layers to extract boundary and edge-related information, refining the segmentation process. Our experiments on three different biomedical image segmentation datasets exhibit that our PAANet achieves favourable performance when compared to other state-of-the-art methods.
2020 CATARACTS Semantic Segmentation Challenge
Oct 21, 2021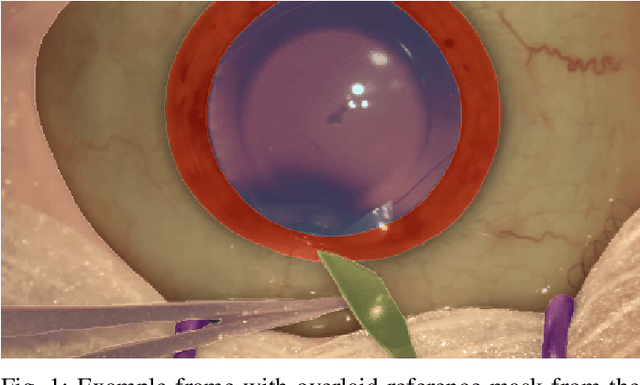
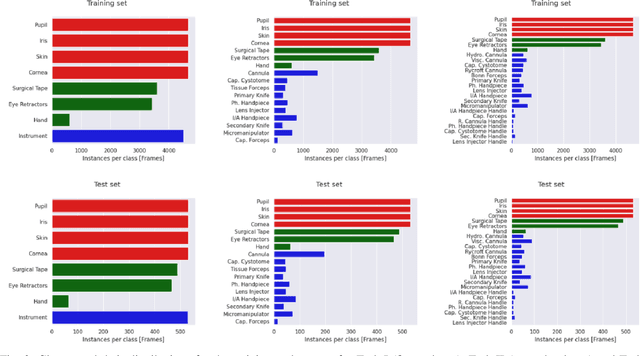
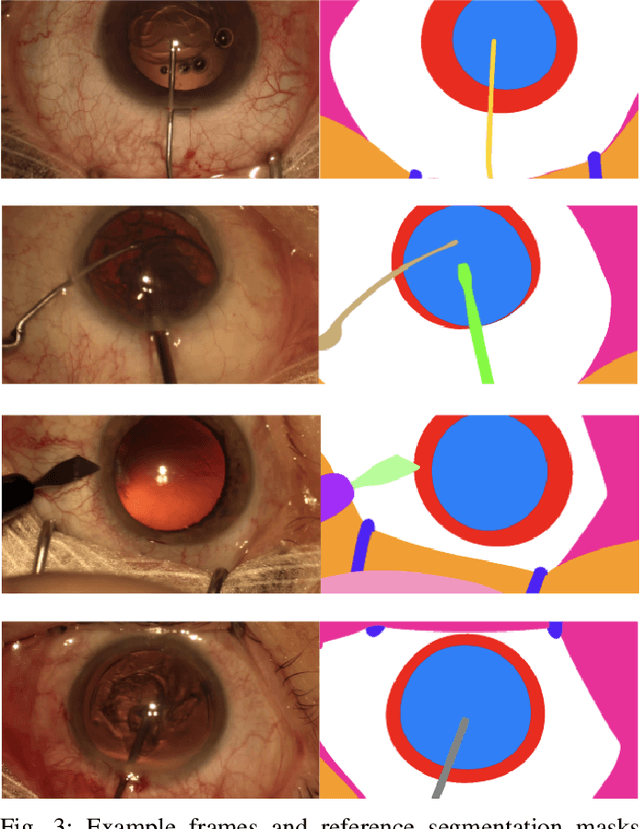
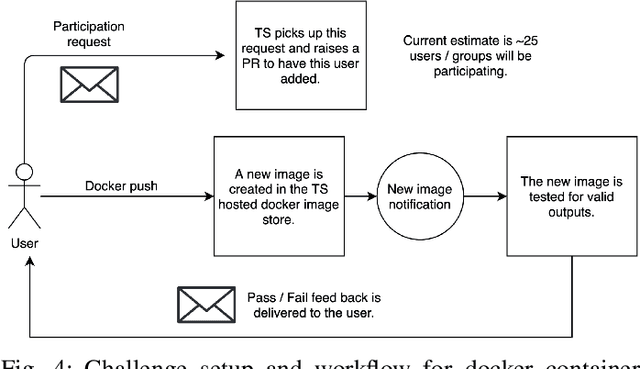
Abstract:Surgical scene segmentation is essential for anatomy and instrument localization which can be further used to assess tissue-instrument interactions during a surgical procedure. In 2017, the Challenge on Automatic Tool Annotation for cataRACT Surgery (CATARACTS) released 50 cataract surgery videos accompanied by instrument usage annotations. These annotations included frame-level instrument presence information. In 2020, we released pixel-wise semantic annotations for anatomy and instruments for 4670 images sampled from 25 videos of the CATARACTS training set. The 2020 CATARACTS Semantic Segmentation Challenge, which was a sub-challenge of the 2020 MICCAI Endoscopic Vision (EndoVis) Challenge, presented three sub-tasks to assess participating solutions on anatomical structure and instrument segmentation. Their performance was assessed on a hidden test set of 531 images from 10 videos of the CATARACTS test set.
Artificial Intelligence in Dry Eye Disease
Sep 02, 2021Abstract:Dry eye disease (DED) has a prevalence of between 5 and 50\%, depending on the diagnostic criteria used and population under study. However, it remains one of the most underdiagnosed and undertreated conditions in ophthalmology. Many tests used in the diagnosis of DED rely on an experienced observer for image interpretation, which may be considered subjective and result in variation in diagnosis. Since artificial intelligence (AI) systems are capable of advanced problem solving, use of such techniques could lead to more objective diagnosis. Although the term `AI' is commonly used, recent success in its applications to medicine is mainly due to advancements in the sub-field of machine learning, which has been used to automatically classify images and predict medical outcomes. Powerful machine learning techniques have been harnessed to understand nuances in patient data and medical images, aiming for consistent diagnosis and stratification of disease severity. This is the first literature review on the use of AI in DED. We provide a brief introduction to AI, report its current use in DED research and its potential for application in the clinic. Our review found that AI has been employed in a wide range of DED clinical tests and research applications, primarily for interpretation of interferometry, slit-lamp and meibography images. While initial results are promising, much work is still needed on model development, clinical testing and standardisation.
Exploring Deep Learning Methods for Real-Time Surgical Instrument Segmentation in Laparoscopy
Aug 03, 2021
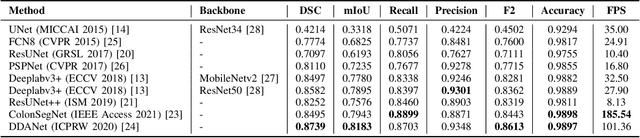
Abstract:Minimally invasive surgery is a surgical intervention used to examine the organs inside the abdomen and has been widely used due to its effectiveness over open surgery. Due to the hardware improvements such as high definition cameras, this procedure has significantly improved and new software methods have demonstrated potential for computer-assisted procedures. However, there exists challenges and requirements to improve detection and tracking of the position of the instruments during these surgical procedures. To this end, we evaluate and compare some popular deep learning methods that can be explored for the automated segmentation of surgical instruments in laparoscopy, an important step towards tool tracking. Our experimental results exhibit that the Dual decoder attention network (DDANet) produces a superior result compared to other recent deep learning methods. DDANet yields a Dice coefficient of 0.8739 and mean intersection-over-union of 0.8183 for the Robust Medical Instrument Segmentation (ROBUST-MIS) Challenge 2019 dataset, at a real-time speed of 101.36 frames-per-second that is critical for such procedures.
A Comprehensive Study on Colorectal Polyp Segmentation with ResUNet++, Conditional Random Field and Test-Time Augmentation
Jul 26, 2021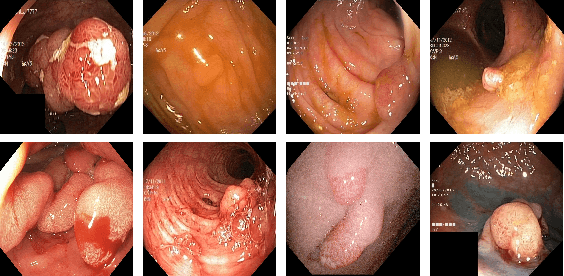
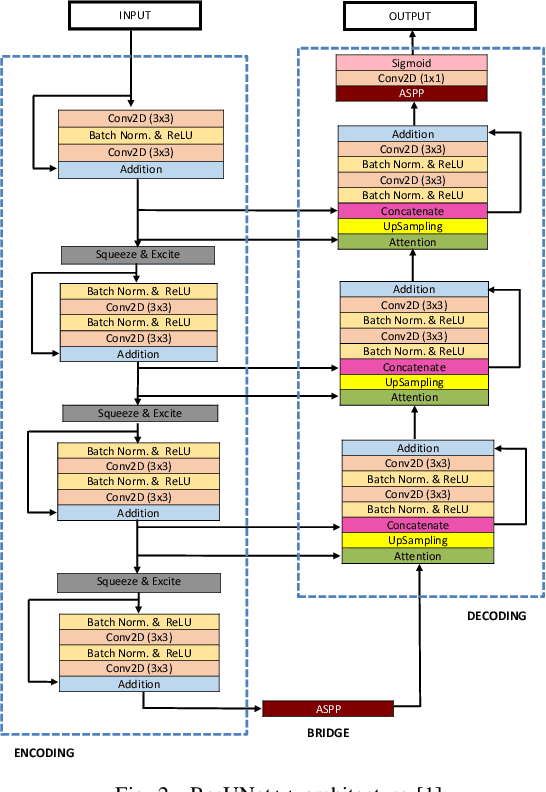
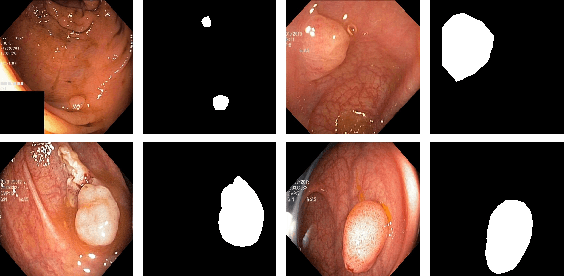
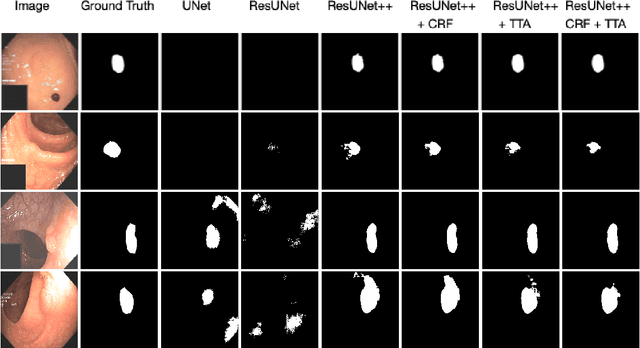
Abstract:Colonoscopy is considered the gold standard for detection of colorectal cancer and its precursors. Existing examination methods are, however, hampered by high overall miss-rate, and many abnormalities are left undetected. Computer-Aided Diagnosis systems based on advanced machine learning algorithms are touted as a game-changer that can identify regions in the colon overlooked by the physicians during endoscopic examinations, and help detect and characterize lesions. In previous work, we have proposed the ResUNet++ architecture and demonstrated that it produces more efficient results compared with its counterparts U-Net and ResUNet. In this paper, we demonstrate that further improvements to the overall prediction performance of the ResUNet++ architecture can be achieved by using conditional random field and test-time augmentation. We have performed extensive evaluations and validated the improvements using six publicly available datasets: Kvasir-SEG, CVC-ClinicDB, CVC-ColonDB, ETIS-Larib Polyp DB, ASU-Mayo Clinic Colonoscopy Video Database, and CVC-VideoClinicDB. Moreover, we compare our proposed architecture and resulting model with other State-of-the-art methods. To explore the generalization capability of ResUNet++ on different publicly available polyp datasets, so that it could be used in a real-world setting, we performed an extensive cross-dataset evaluation. The experimental results show that applying CRF and TTA improves the performance on various polyp segmentation datasets both on the same dataset and cross-dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge