Jens Petersen
Transformer Utilization in Medical Image Segmentation Networks
Apr 09, 2023Abstract:Owing to success in the data-rich domain of natural images, Transformers have recently become popular in medical image segmentation. However, the pairing of Transformers with convolutional blocks in varying architectural permutations leaves their relative effectiveness to open interpretation. We introduce Transformer Ablations that replace the Transformer blocks with plain linear operators to quantify this effectiveness. With experiments on 8 models on 2 medical image segmentation tasks, we explore -- 1) the replaceable nature of Transformer-learnt representations, 2) Transformer capacity alone cannot prevent representational replaceability and works in tandem with effective design, 3) The mere existence of explicit feature hierarchies in transformer blocks is more beneficial than accompanying self-attention modules, 4) Major spatial downsampling before Transformer modules should be used with caution.
MedNeXt: Transformer-driven Scaling of ConvNets for Medical Image Segmentation
Mar 22, 2023



Abstract:There has been exploding interest in embracing Transformer-based architectures for medical image segmentation. However, the lack of large-scale annotated medical datasets make achieving performances equivalent to those in natural images challenging. Convolutional networks, in contrast, have higher inductive biases and consequently, are easily trainable to high performance. Recently, the ConvNeXt architecture attempted to modernize the standard ConvNet by mirroring Transformer blocks. In this work, we improve upon this to design a modernized and scalable convolutional architecture customized to challenges of data-scarce medical settings. We introduce MedNeXt, a Transformer-inspired large kernel segmentation network which introduces - 1) A fully ConvNeXt 3D Encoder-Decoder Network for medical image segmentation, 2) Residual ConvNeXt up and downsampling blocks to preserve semantic richness across scales, 3) A novel technique to iteratively increase kernel sizes by upsampling small kernel networks, to prevent performance saturation on limited medical data, 4) Compound scaling at multiple levels (depth, width, kernel size) of MedNeXt. This leads to state-of-the-art performance on 4 tasks on CT and MRI modalities and varying dataset sizes, representing a modernized deep architecture for medical image segmentation.
Understanding metric-related pitfalls in image analysis validation
Feb 09, 2023Abstract:Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice. However, increasing evidence shows that particularly in image analysis, metrics are often chosen inadequately in relation to the underlying research problem. This could be attributed to a lack of accessibility of metric-related knowledge: While taking into account the individual strengths, weaknesses, and limitations of validation metrics is a critical prerequisite to making educated choices, the relevant knowledge is currently scattered and poorly accessible to individual researchers. Based on a multi-stage Delphi process conducted by a multidisciplinary expert consortium as well as extensive community feedback, the present work provides the first reliable and comprehensive common point of access to information on pitfalls related to validation metrics in image analysis. Focusing on biomedical image analysis but with the potential of transfer to other fields, the addressed pitfalls generalize across application domains and are categorized according to a newly created, domain-agnostic taxonomy. To facilitate comprehension, illustrations and specific examples accompany each pitfall. As a structured body of information accessible to researchers of all levels of expertise, this work enhances global comprehension of a key topic in image analysis validation.
Neural Image Compression with a Diffusion-Based Decoder
Jan 23, 2023



Abstract:Diffusion probabilistic models have recently achieved remarkable success in generating high quality image and video data. In this work, we build on this class of generative models and introduce a method for lossy compression of high resolution images. The resulting codec, which we call DIffuson-based Residual Augmentation Codec (DIRAC),is the first neural codec to allow smooth traversal of the rate-distortion-perception tradeoff at test time, while obtaining competitive performance with GAN-based methods in perceptual quality. Furthermore, while sampling from diffusion probabilistic models is notoriously expensive, we show that in the compression setting the number of steps can be drastically reduced.
Explicit Temporal Embedding in Deep Generative Latent Models for Longitudinal Medical Image Synthesis
Jan 13, 2023



Abstract:Medical imaging plays a vital role in modern diagnostics and treatment. The temporal nature of disease or treatment progression often results in longitudinal data. Due to the cost and potential harm, acquiring large medical datasets necessary for deep learning can be difficult. Medical image synthesis could help mitigate this problem. However, until now, the availability of GANs capable of synthesizing longitudinal volumetric data has been limited. To address this, we use the recent advances in latent space-based image editing to propose a novel joint learning scheme to explicitly embed temporal dependencies in the latent space of GANs. This, in contrast to previous methods, allows us to synthesize continuous, smooth, and high-quality longitudinal volumetric data with limited supervision. We show the effectiveness of our approach on three datasets containing different longitudinal dependencies. Namely, modeling a simple image transformation, breathing motion, and tumor regression, all while showing minimal disentanglement. The implementation is made available online at https://github.com/julschoen/Temp-GAN.
CRADL: Contrastive Representations for Unsupervised Anomaly Detection and Localization
Jan 05, 2023



Abstract:Unsupervised anomaly detection in medical imaging aims to detect and localize arbitrary anomalies without requiring annotated anomalous data during training. Often, this is achieved by learning a data distribution of normal samples and detecting anomalies as regions in the image which deviate from this distribution. Most current state-of-the-art methods use latent variable generative models operating directly on the images. However, generative models have been shown to mostly capture low-level features, s.a. pixel-intensities, instead of rich semantic features, which also applies to their representations. We circumvent this problem by proposing CRADL whose core idea is to model the distribution of normal samples directly in the low-dimensional representation space of an encoder trained with a contrastive pretext-task. By utilizing the representations of contrastive learning, we aim to fix the over-fixation on low-level features and learn more semantic-rich representations. Our experiments on anomaly detection and localization tasks using three distinct evaluation datasets show that 1) contrastive representations are superior to representations of generative latent variable models and 2) the CRADL framework shows competitive or superior performance to state-of-the-art.
Interpreting Latent Spaces of Generative Models for Medical Images using Unsupervised Methods
Jul 20, 2022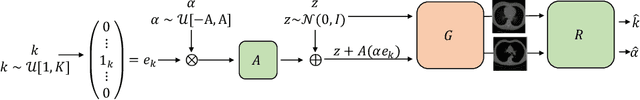
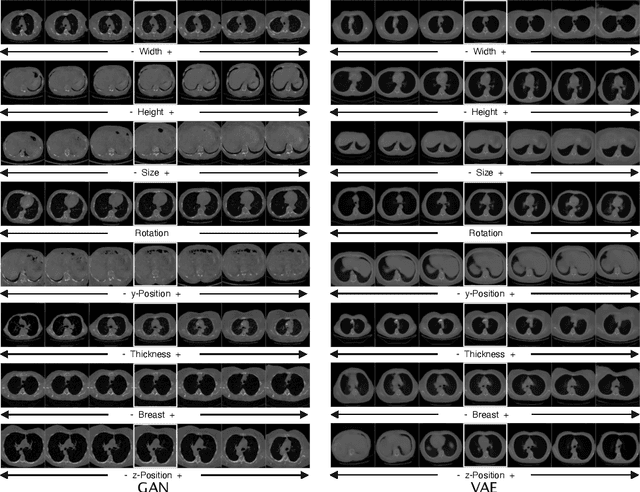

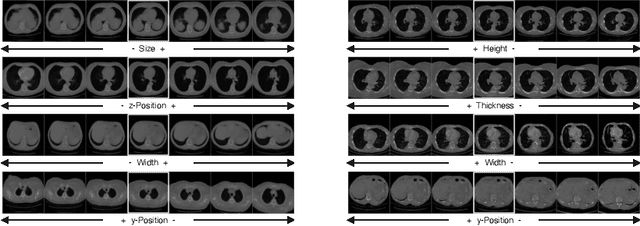
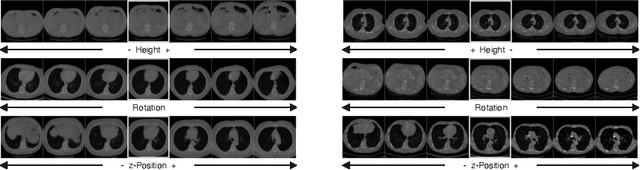
Abstract:Generative models such as Generative Adversarial Networks (GANs) and Variational Autoencoders (VAEs) play an increasingly important role in medical image analysis. The latent spaces of these models often show semantically meaningful directions corresponding to human-interpretable image transformations. However, until now, their exploration for medical images has been limited due to the requirement of supervised data. Several methods for unsupervised discovery of interpretable directions in GAN latent spaces have shown interesting results on natural images. This work explores the potential of applying these techniques on medical images by training a GAN and a VAE on thoracic CT scans and using an unsupervised method to discover interpretable directions in the resulting latent space. We find several directions corresponding to non-trivial image transformations, such as rotation or breast size. Furthermore, the directions show that the generative models capture 3D structure despite being presented only with 2D data. The results show that unsupervised methods to discover interpretable directions in GANs generalize to VAEs and can be applied to medical images. This opens a wide array of future work using these methods in medical image analysis.
Metrics reloaded: Pitfalls and recommendations for image analysis validation
Jun 03, 2022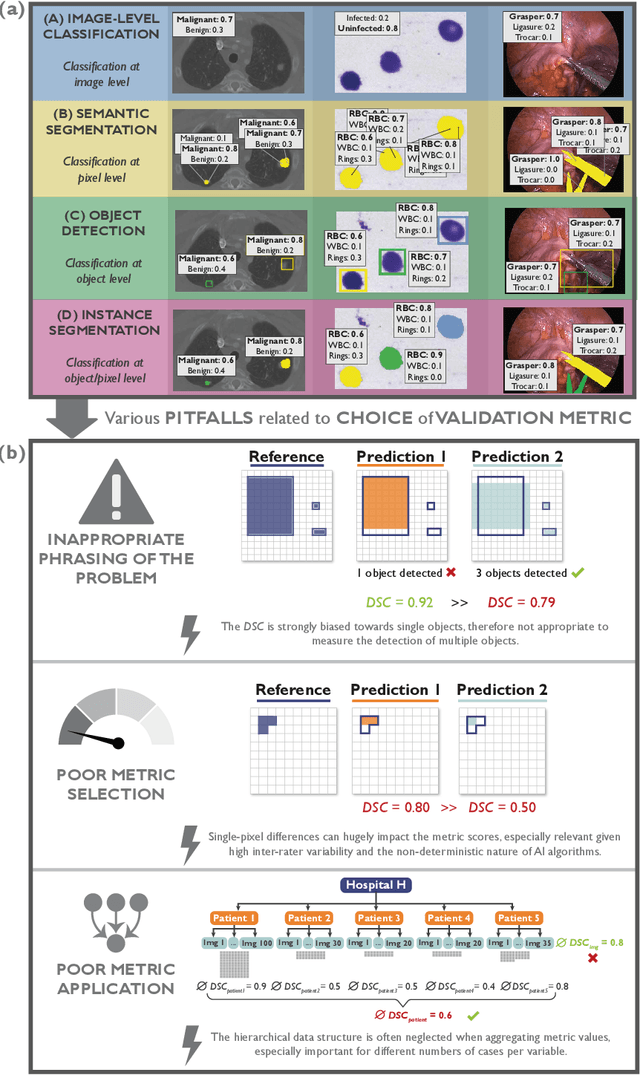
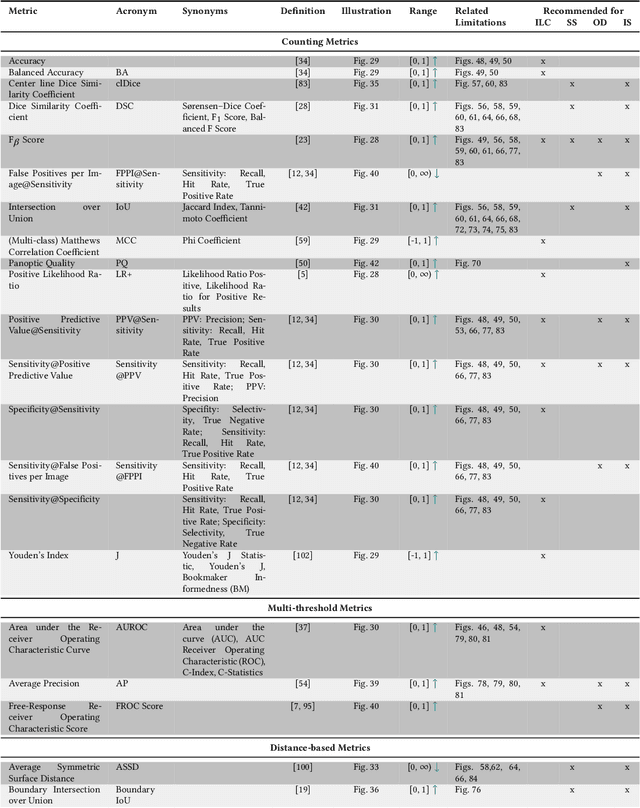
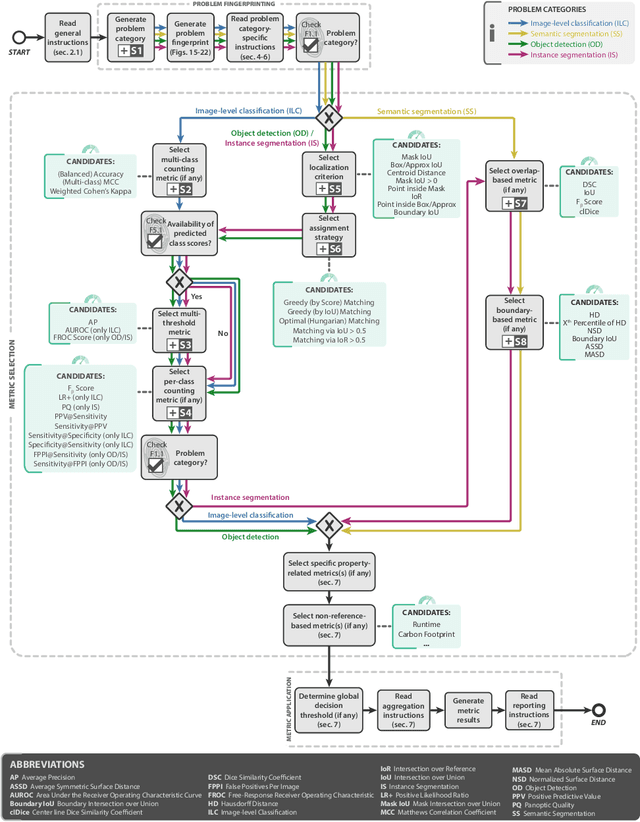
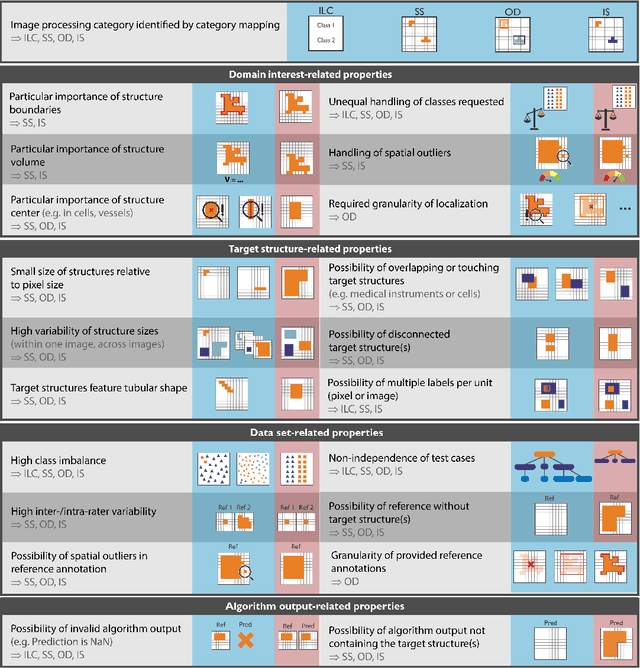
Abstract:The field of automatic biomedical image analysis crucially depends on robust and meaningful performance metrics for algorithm validation. Current metric usage, however, is often ill-informed and does not reflect the underlying domain interest. Here, we present a comprehensive framework that guides researchers towards choosing performance metrics in a problem-aware manner. Specifically, we focus on biomedical image analysis problems that can be interpreted as a classification task at image, object or pixel level. The framework first compiles domain interest-, target structure-, data set- and algorithm output-related properties of a given problem into a problem fingerprint, while also mapping it to the appropriate problem category, namely image-level classification, semantic segmentation, instance segmentation, or object detection. It then guides users through the process of selecting and applying a set of appropriate validation metrics while making them aware of potential pitfalls related to individual choices. In this paper, we describe the current status of the Metrics Reloaded recommendation framework, with the goal of obtaining constructive feedback from the image analysis community. The current version has been developed within an international consortium of more than 60 image analysis experts and will be made openly available as a user-friendly toolkit after community-driven optimization.
Patch-based medical image segmentation using Quantum Tensor Networks
Sep 15, 2021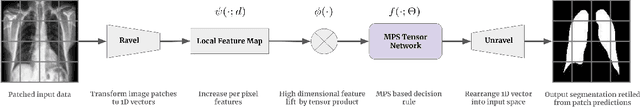
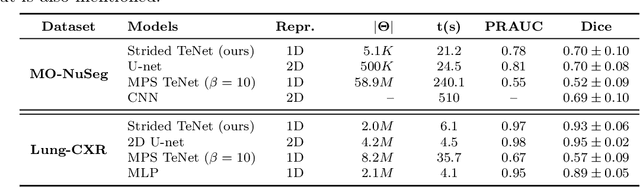
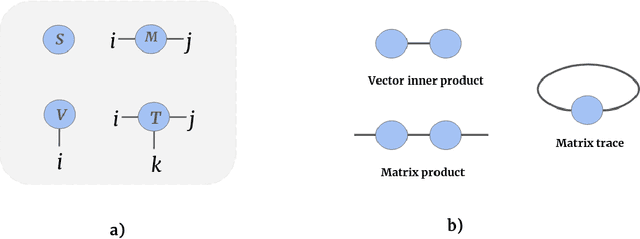
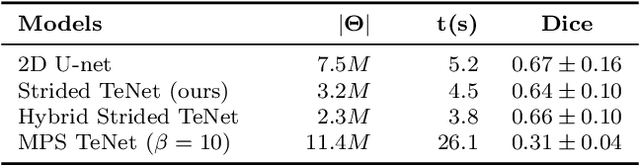
Abstract:Tensor networks are efficient factorisations of high dimensional tensors into a network of lower order tensors. They have been most commonly used to model entanglement in quantum many-body systems and more recently are witnessing increased applications in supervised machine learning. In this work, we formulate image segmentation in a supervised setting with tensor networks. The key idea is to first lift the pixels in image patches to exponentially high dimensional feature spaces and using a linear decision hyper-plane to classify the input pixels into foreground and background classes. The high dimensional linear model itself is approximated using the matrix product state (MPS) tensor network. The MPS is weight-shared between the non-overlapping image patches resulting in our strided tensor network model. The performance of the proposed model is evaluated on three 2D- and one 3D- biomedical imaging datasets. The performance of the proposed tensor network segmentation model is compared with relevant baseline methods. In the 2D experiments, the tensor network model yeilds competitive performance compared to the baseline methods while being more resource efficient.
Continuous-Time Deep Glioma Growth Models
Jul 02, 2021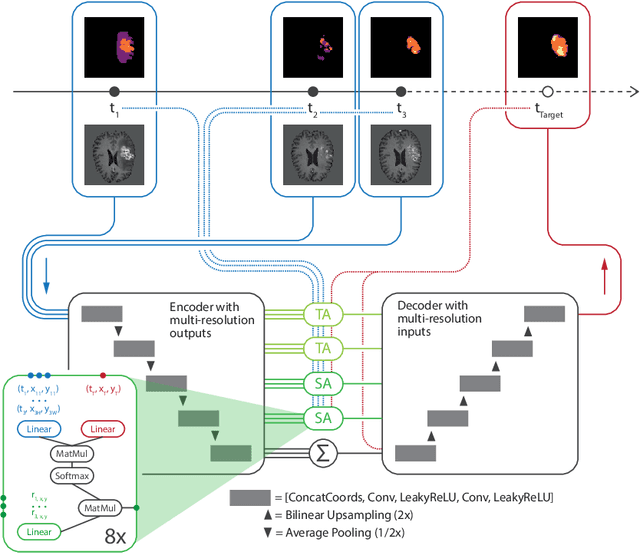
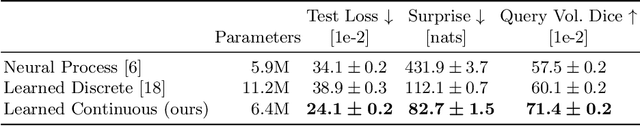


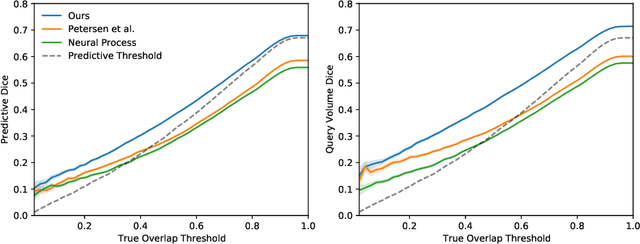
Abstract:The ability to estimate how a tumor might evolve in the future could have tremendous clinical benefits, from improved treatment decisions to better dose distribution in radiation therapy. Recent work has approached the glioma growth modeling problem via deep learning and variational inference, thus learning growth dynamics entirely from a real patient data distribution. So far, this approach was constrained to predefined image acquisition intervals and sequences of fixed length, which limits its applicability in more realistic scenarios. We overcome these limitations by extending Neural Processes, a class of conditional generative models for stochastic time series, with a hierarchical multi-scale representation encoding including a spatio-temporal attention mechanism. The result is a learned growth model that can be conditioned on an arbitrary number of observations, and that can produce a distribution of temporally consistent growth trajectories on a continuous time axis. On a dataset of 379 patients, the approach successfully captures both global and finer-grained variations in the images, exhibiting superior performance compared to other learned growth models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge