Indrani Bhattacharya
ProsDectNet: Bridging the Gap in Prostate Cancer Detection via Transrectal B-mode Ultrasound Imaging
Dec 08, 2023



Abstract:Interpreting traditional B-mode ultrasound images can be challenging due to image artifacts (e.g., shadowing, speckle), leading to low sensitivity and limited diagnostic accuracy. While Magnetic Resonance Imaging (MRI) has been proposed as a solution, it is expensive and not widely available. Furthermore, most biopsies are guided by Transrectal Ultrasound (TRUS) alone and can miss up to 52% cancers, highlighting the need for improved targeting. To address this issue, we propose ProsDectNet, a multi-task deep learning approach that localizes prostate cancer on B-mode ultrasound. Our model is pre-trained using radiologist-labeled data and fine-tuned using biopsy-confirmed labels. ProsDectNet includes a lesion detection and patch classification head, with uncertainty minimization using entropy to improve model performance and reduce false positive predictions. We trained and validated ProsDectNet using a cohort of 289 patients who underwent MRI-TRUS fusion targeted biopsy. We then tested our approach on a group of 41 patients and found that ProsDectNet outperformed the average expert clinician in detecting prostate cancer on B-mode ultrasound images, achieving a patient-level ROC-AUC of 82%, a sensitivity of 74%, and a specificity of 67%. Our results demonstrate that ProsDectNet has the potential to be used as a computer-aided diagnosis system to improve targeted biopsy and treatment planning.
BLOOM: A 176B-Parameter Open-Access Multilingual Language Model
Nov 09, 2022Abstract:Large language models (LLMs) have been shown to be able to perform new tasks based on a few demonstrations or natural language instructions. While these capabilities have led to widespread adoption, most LLMs are developed by resource-rich organizations and are frequently kept from the public. As a step towards democratizing this powerful technology, we present BLOOM, a 176B-parameter open-access language model designed and built thanks to a collaboration of hundreds of researchers. BLOOM is a decoder-only Transformer language model that was trained on the ROOTS corpus, a dataset comprising hundreds of sources in 46 natural and 13 programming languages (59 in total). We find that BLOOM achieves competitive performance on a wide variety of benchmarks, with stronger results after undergoing multitask prompted finetuning. To facilitate future research and applications using LLMs, we publicly release our models and code under the Responsible AI License.
Correlated Feature Aggregation by Region Helps Distinguish Aggressive from Indolent Clear Cell Renal Cell Carcinoma Subtypes on CT
Sep 29, 2022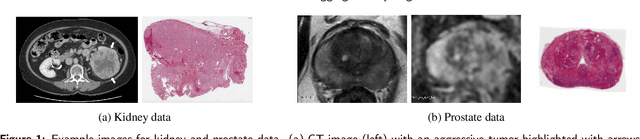
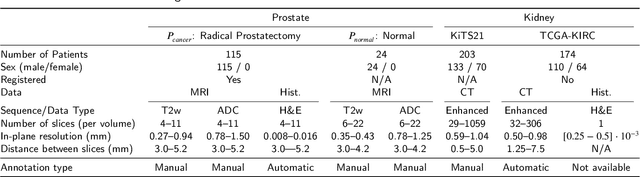
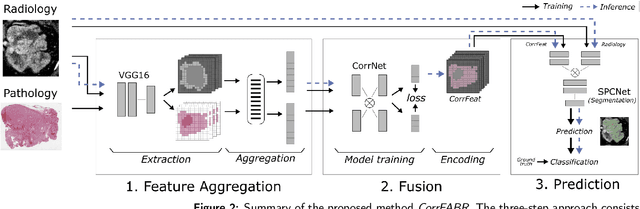

Abstract:Renal cell carcinoma (RCC) is a common cancer that varies in clinical behavior. Indolent RCC is often low-grade without necrosis and can be monitored without treatment. Aggressive RCC is often high-grade and can cause metastasis and death if not promptly detected and treated. While most kidney cancers are detected on CT scans, grading is based on histology from invasive biopsy or surgery. Determining aggressiveness on CT images is clinically important as it facilitates risk stratification and treatment planning. This study aims to use machine learning methods to identify radiology features that correlate with features on pathology to facilitate assessment of cancer aggressiveness on CT images instead of histology. This paper presents a novel automated method, Correlated Feature Aggregation By Region (CorrFABR), for classifying aggressiveness of clear cell RCC by leveraging correlations between radiology and corresponding unaligned pathology images. CorrFABR consists of three main steps: (1) Feature Aggregation where region-level features are extracted from radiology and pathology images, (2) Fusion where radiology features correlated with pathology features are learned on a region level, and (3) Prediction where the learned correlated features are used to distinguish aggressive from indolent clear cell RCC using CT alone as input. Thus, during training, CorrFABR learns from both radiology and pathology images, but during inference, CorrFABR will distinguish aggressive from indolent clear cell RCC using CT alone, in the absence of pathology images. CorrFABR improved classification performance over radiology features alone, with an increase in binary classification F1-score from 0.68 (0.04) to 0.73 (0.03). This demonstrates the potential of incorporating pathology disease characteristics for improved classification of aggressiveness of clear cell RCC on CT images.
Domain Generalization for Prostate Segmentation in Transrectal Ultrasound Images: A Multi-center Study
Sep 05, 2022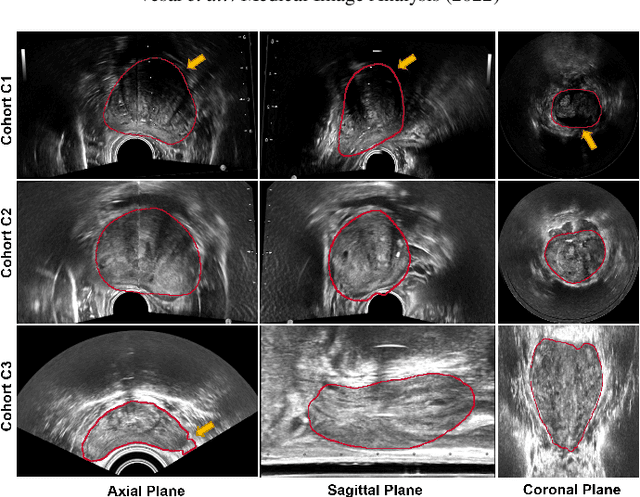
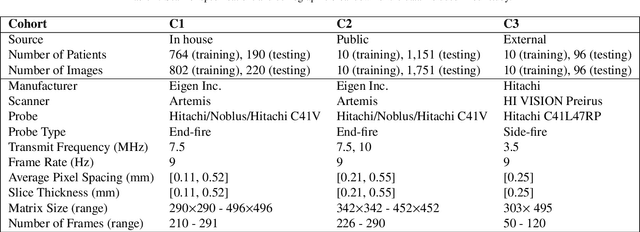
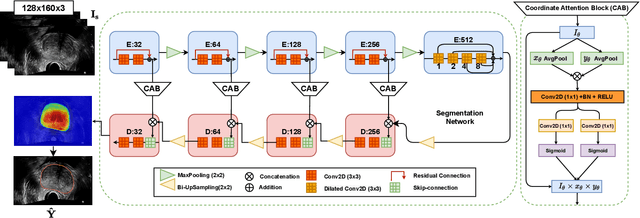
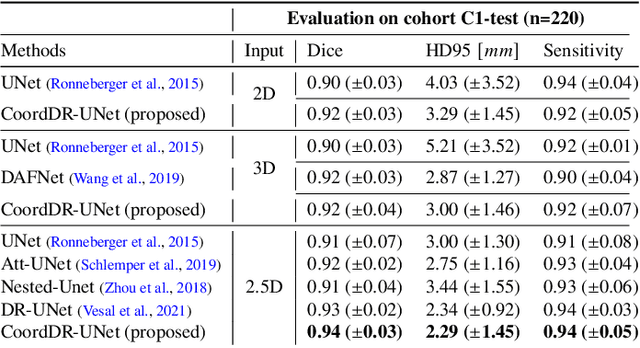
Abstract:Prostate biopsy and image-guided treatment procedures are often performed under the guidance of ultrasound fused with magnetic resonance images (MRI). Accurate image fusion relies on accurate segmentation of the prostate on ultrasound images. Yet, the reduced signal-to-noise ratio and artifacts (e.g., speckle and shadowing) in ultrasound images limit the performance of automated prostate segmentation techniques and generalizing these methods to new image domains is inherently difficult. In this study, we address these challenges by introducing a novel 2.5D deep neural network for prostate segmentation on ultrasound images. Our approach addresses the limitations of transfer learning and finetuning methods (i.e., drop in performance on the original training data when the model weights are updated) by combining a supervised domain adaptation technique and a knowledge distillation loss. The knowledge distillation loss allows the preservation of previously learned knowledge and reduces the performance drop after model finetuning on new datasets. Furthermore, our approach relies on an attention module that considers model feature positioning information to improve the segmentation accuracy. We trained our model on 764 subjects from one institution and finetuned our model using only ten subjects from subsequent institutions. We analyzed the performance of our method on three large datasets encompassing 2067 subjects from three different institutions. Our method achieved an average Dice Similarity Coefficient (Dice) of $94.0\pm0.03$ and Hausdorff Distance (HD95) of 2.28 $mm$ in an independent set of subjects from the first institution. Moreover, our model generalized well in the studies from the other two institutions (Dice: $91.0\pm0.03$; HD95: 3.7$mm$ and Dice: $82.0\pm0.03$; HD95: 7.1 $mm$).
Bridging the gap between prostate radiology and pathology through machine learning
Dec 03, 2021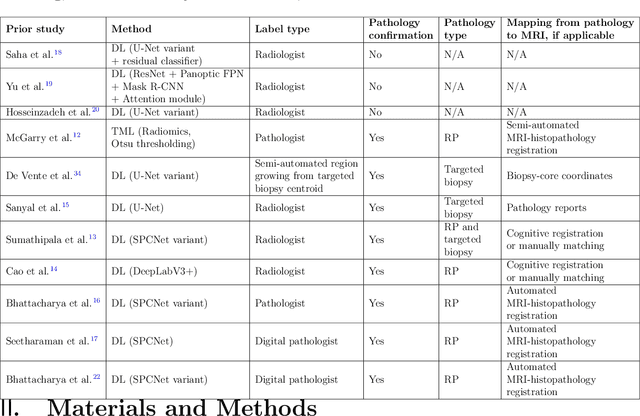
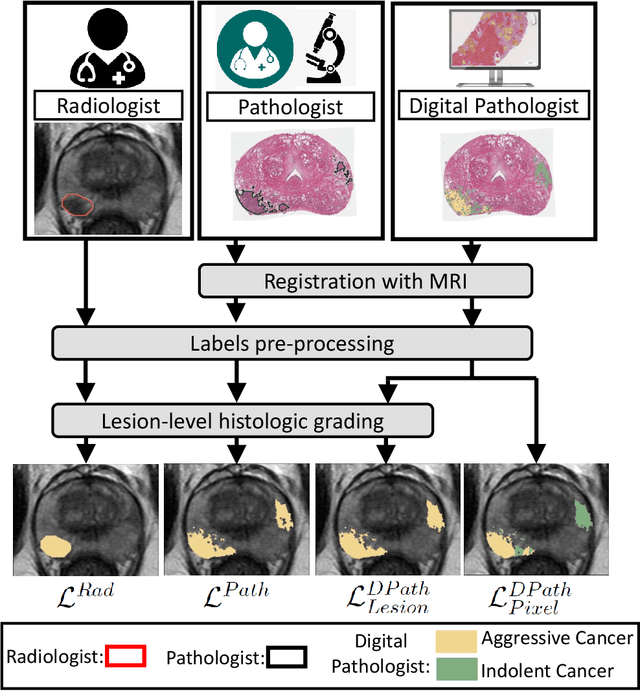
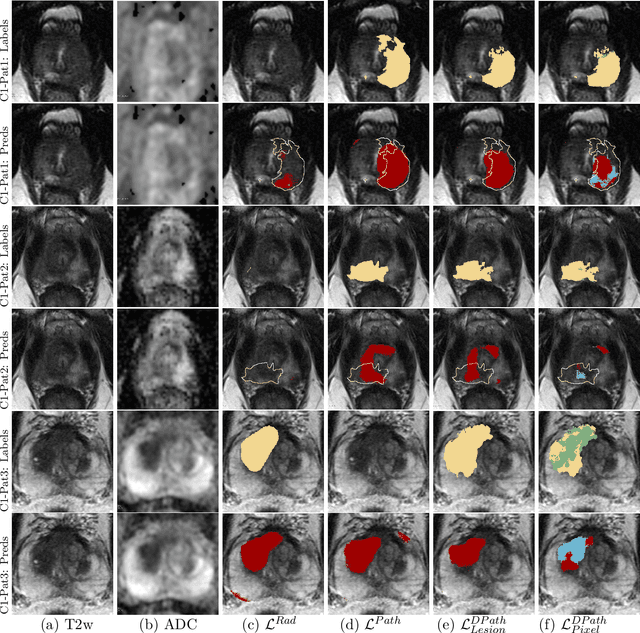
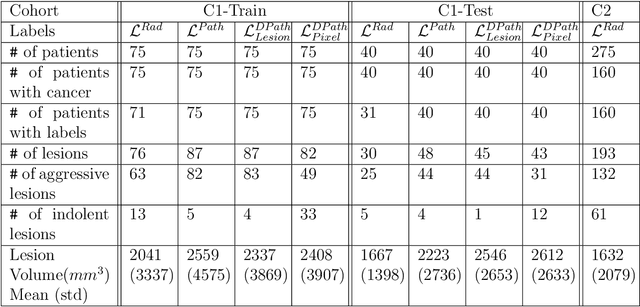
Abstract:Prostate cancer is the second deadliest cancer for American men. While Magnetic Resonance Imaging (MRI) is increasingly used to guide targeted biopsies for prostate cancer diagnosis, its utility remains limited due to high rates of false positives and false negatives as well as low inter-reader agreements. Machine learning methods to detect and localize cancer on prostate MRI can help standardize radiologist interpretations. However, existing machine learning methods vary not only in model architecture, but also in the ground truth labeling strategies used for model training. In this study, we compare different labeling strategies, namely, pathology-confirmed radiologist labels, pathologist labels on whole-mount histopathology images, and lesion-level and pixel-level digital pathologist labels (previously validated deep learning algorithm on histopathology images to predict pixel-level Gleason patterns) on whole-mount histopathology images. We analyse the effects these labels have on the performance of the trained machine learning models. Our experiments show that (1) radiologist labels and models trained with them can miss cancers, or underestimate cancer extent, (2) digital pathologist labels and models trained with them have high concordance with pathologist labels, and (3) models trained with digital pathologist labels achieve the best performance in prostate cancer detection in two different cohorts with different disease distributions, irrespective of the model architecture used. Digital pathologist labels can reduce challenges associated with human annotations, including labor, time, inter- and intra-reader variability, and can help bridge the gap between prostate radiology and pathology by enabling the training of reliable machine learning models to detect and localize prostate cancer on MRI.
Weakly Supervised Registration of Prostate MRI and Histopathology Images
Jun 23, 2021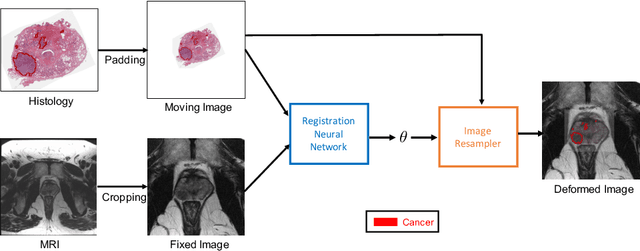
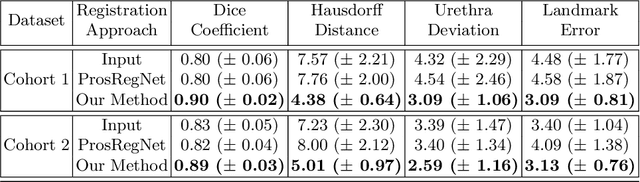
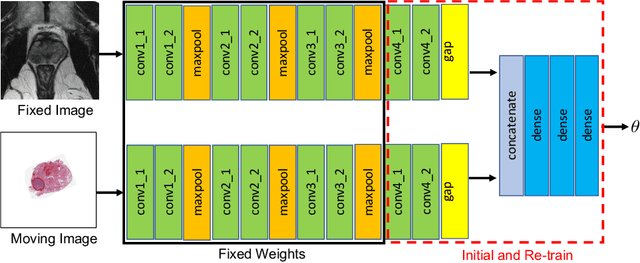
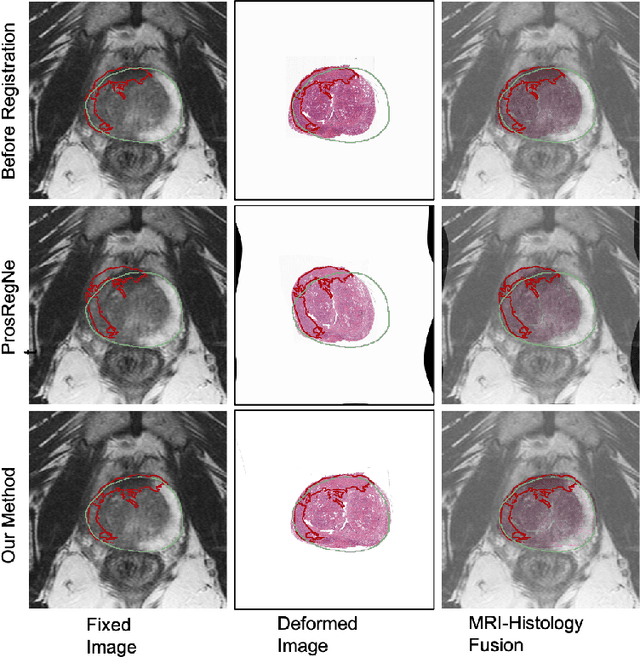
Abstract:The interpretation of prostate MRI suffers from low agreement across radiologists due to the subtle differences between cancer and normal tissue. Image registration addresses this issue by accurately mapping the ground-truth cancer labels from surgical histopathology images onto MRI. Cancer labels achieved by image registration can be used to improve radiologists' interpretation of MRI by training deep learning models for early detection of prostate cancer. A major limitation of current automated registration approaches is that they require manual prostate segmentations, which is a time-consuming task, prone to errors. This paper presents a weakly supervised approach for affine and deformable registration of MRI and histopathology images without requiring prostate segmentations. We used manual prostate segmentations and mono-modal synthetic image pairs to train our registration networks to align prostate boundaries and local prostate features. Although prostate segmentations were used during the training of the network, such segmentations were not needed when registering unseen images at inference time. We trained and validated our registration network with 135 and 10 patients from an internal cohort, respectively. We tested the performance of our method using 16 patients from the internal cohort and 22 patients from an external cohort. The results show that our weakly supervised method has achieved significantly higher registration accuracy than a state-of-the-art method run without prostate segmentations. Our deep learning framework will ease the registration of MRI and histopathology images by obviating the need for prostate segmentations.
CorrSigNet: Learning CORRelated Prostate Cancer SIGnatures from Radiology and Pathology Images for Improved Computer Aided Diagnosis
Jul 31, 2020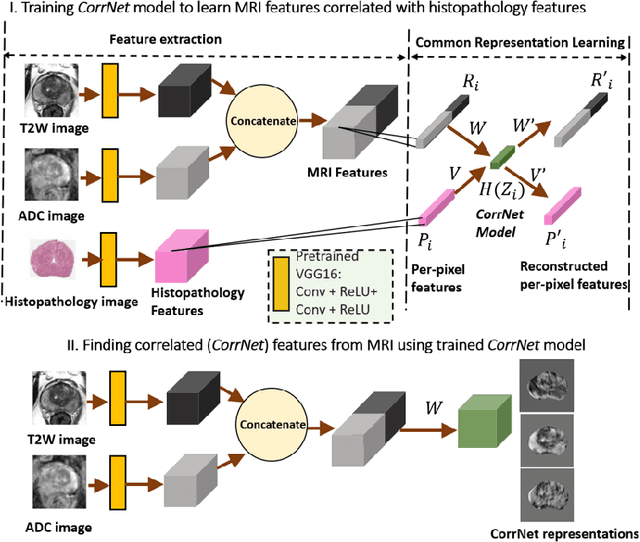
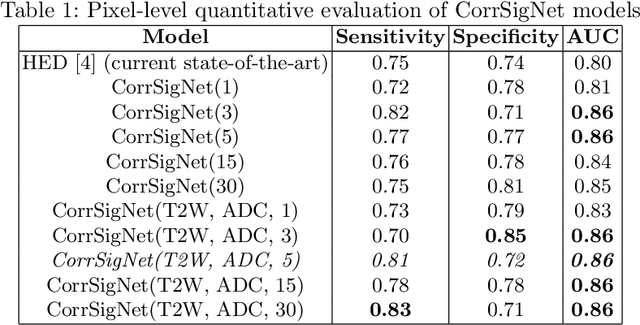
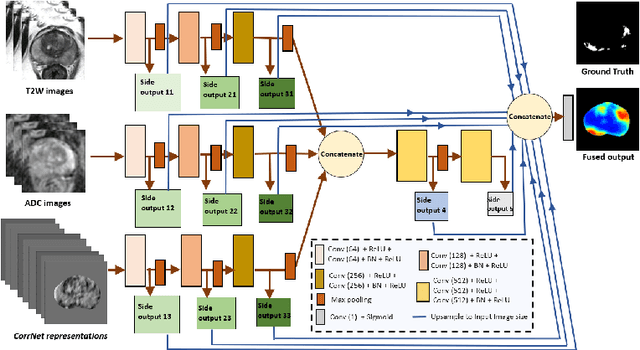
Abstract:Magnetic Resonance Imaging (MRI) is widely used for screening and staging prostate cancer. However, many prostate cancers have subtle features which are not easily identifiable on MRI, resulting in missed diagnoses and alarming variability in radiologist interpretation. Machine learning models have been developed in an effort to improve cancer identification, but current models localize cancer using MRI-derived features, while failing to consider the disease pathology characteristics observed on resected tissue. In this paper, we propose CorrSigNet, an automated two-step model that localizes prostate cancer on MRI by capturing the pathology features of cancer. First, the model learns MRI signatures of cancer that are correlated with corresponding histopathology features using Common Representation Learning. Second, the model uses the learned correlated MRI features to train a Convolutional Neural Network to localize prostate cancer. The histopathology images are used only in the first step to learn the correlated features. Once learned, these correlated features can be extracted from MRI of new patients (without histopathology or surgery) to localize cancer. We trained and validated our framework on a unique dataset of 75 patients with 806 slices who underwent MRI followed by prostatectomy surgery. We tested our method on an independent test set of 20 prostatectomy patients (139 slices, 24 cancerous lesions, 1.12M pixels) and achieved a per-pixel sensitivity of 0.81, specificity of 0.71, AUC of 0.86 and a per-lesion AUC of $0.96 \pm 0.07$, outperforming the current state-of-the-art accuracy in predicting prostate cancer using MRI.
Multi-modal dialog for browsing large visual catalogs using exploration-exploitation paradigm in a joint embedding space
Jan 29, 2019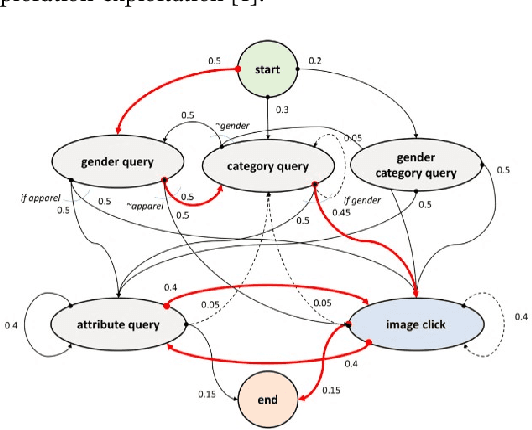
Abstract:We present a multi-modal dialog system to assist online shoppers in visually browsing through large catalogs. Visual browsing is different from visual search in that it allows the user to explore the wide range of products in a catalog, beyond the exact search matches. We focus on a slightly asymmetric version of the complete multi-modal dialog where the system can understand both text and image queries but responds only in images. We formulate our problem of "showing $k$ best images to a user" based on the dialog context so far, as sampling from a Gaussian Mixture Model in a high dimensional joint multi-modal embedding space, that embed both the text and the image queries. Our system remembers the context of the dialog and uses an exploration-exploitation paradigm to assist in visual browsing. We train and evaluate the system on a multi-modal dialog dataset that we generate from large catalog data. Our experiments are promising and show that the agent is capable of learning and can display relevant results with an average cosine similarity of 0.85 to the ground truth. Our preliminary human evaluation also corroborates the fact that such a multi-modal dialog system for visual browsing is well-received and is capable of engaging human users.
Designing Intelligent Automation based Solutions for Complex Social Problems
Jun 16, 2016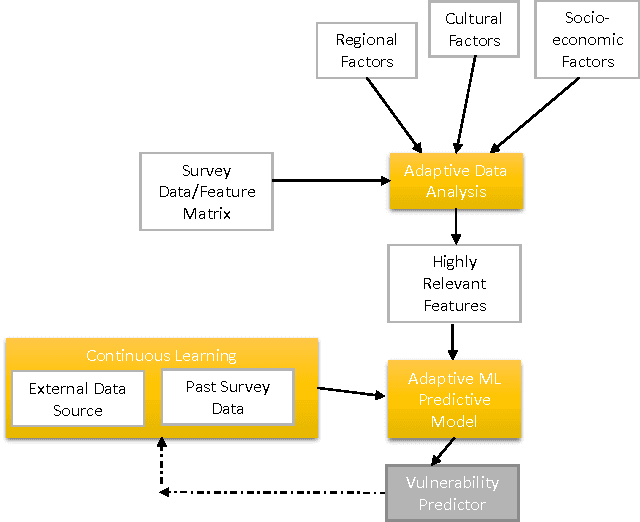
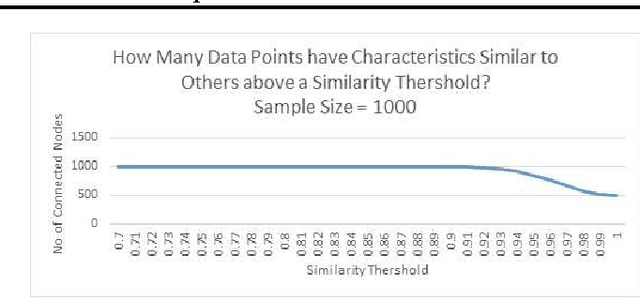
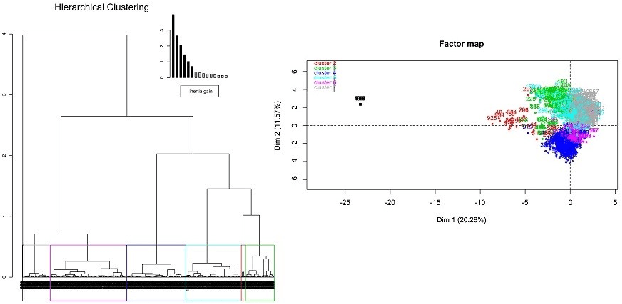

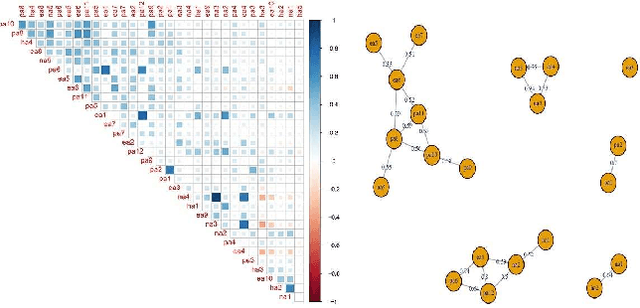
Abstract:Deciding effective and timely preventive measures against complex social problems affecting relatively low income geographies is a difficult challenge. There is a strong need to adopt intelligent automation based solutions with low cost imprints to tackle these problems at larger scales. Starting with the hypothesis that analytical modelling and analysis of social phenomena with high accuracy is in general inherently hard, in this paper we propose design framework to enable data-driven machine learning based adaptive solution approach towards enabling more effective preventive measures. We use survey data collected from a socio-economically backward region of India about adolescent girls to illustrate the design approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge