Dean C Barratt
Trackerless freehand ultrasound with sequence modelling and auxiliary transformation over past and future frames
Nov 09, 2022



Abstract:Three-dimensional (3D) freehand ultrasound (US) reconstruction without a tracker can be advantageous over its two-dimensional or tracked counterparts in many clinical applications. In this paper, we propose to estimate 3D spatial transformation between US frames from both past and future 2D images, using feed-forward and recurrent neural networks (RNNs). With the temporally available frames, a further multi-task learning algorithm is proposed to utilise a large number of auxiliary transformation-predicting tasks between them. Using more than 40,000 US frames acquired from 228 scans on 38 forearms of 19 volunteers in a volunteer study, the hold-out test performance is quantified by frame prediction accuracy, volume reconstruction overlap, accumulated tracking error and final drift, based on ground-truth from an optical tracker. The results show the importance of modelling the temporal-spatially correlated input frames as well as output transformations, with further improvement owing to additional past and/or future frames. The best performing model was associated with predicting transformation between moderately-spaced frames, with an interval of less than ten frames at 20 frames per second (fps). Little benefit was observed by adding frames more than one second away from the predicted transformation, with or without LSTM-based RNNs. Interestingly, with the proposed approach, explicit within-sequence loss that encourages consistency in composing transformations or minimises accumulated error may no longer be required. The implementation code and volunteer data will be made publicly available ensuring reproducibility and further research.
Domain Generalization for Prostate Segmentation in Transrectal Ultrasound Images: A Multi-center Study
Sep 05, 2022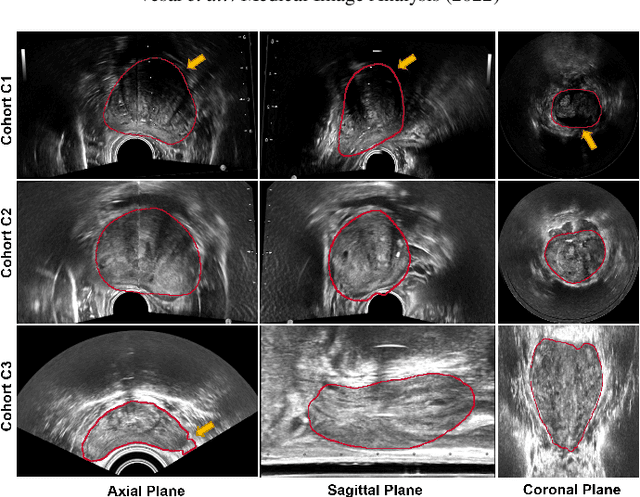
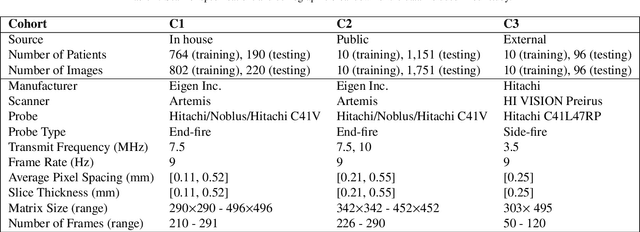
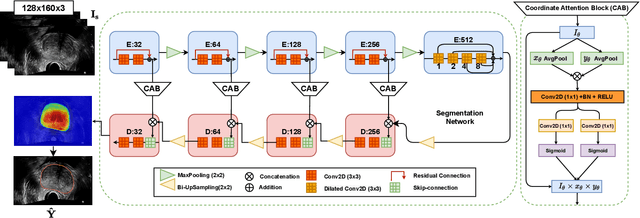
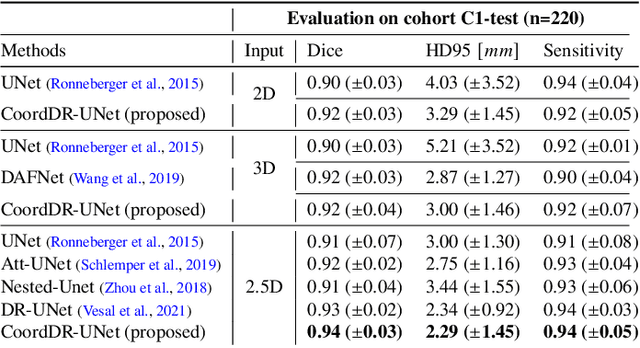
Abstract:Prostate biopsy and image-guided treatment procedures are often performed under the guidance of ultrasound fused with magnetic resonance images (MRI). Accurate image fusion relies on accurate segmentation of the prostate on ultrasound images. Yet, the reduced signal-to-noise ratio and artifacts (e.g., speckle and shadowing) in ultrasound images limit the performance of automated prostate segmentation techniques and generalizing these methods to new image domains is inherently difficult. In this study, we address these challenges by introducing a novel 2.5D deep neural network for prostate segmentation on ultrasound images. Our approach addresses the limitations of transfer learning and finetuning methods (i.e., drop in performance on the original training data when the model weights are updated) by combining a supervised domain adaptation technique and a knowledge distillation loss. The knowledge distillation loss allows the preservation of previously learned knowledge and reduces the performance drop after model finetuning on new datasets. Furthermore, our approach relies on an attention module that considers model feature positioning information to improve the segmentation accuracy. We trained our model on 764 subjects from one institution and finetuned our model using only ten subjects from subsequent institutions. We analyzed the performance of our method on three large datasets encompassing 2067 subjects from three different institutions. Our method achieved an average Dice Similarity Coefficient (Dice) of $94.0\pm0.03$ and Hausdorff Distance (HD95) of 2.28 $mm$ in an independent set of subjects from the first institution. Moreover, our model generalized well in the studies from the other two institutions (Dice: $91.0\pm0.03$; HD95: 3.7$mm$ and Dice: $82.0\pm0.03$; HD95: 7.1 $mm$).
Meta-Registration: Learning Test-Time Optimization for Single-Pair Image Registration
Jul 22, 2022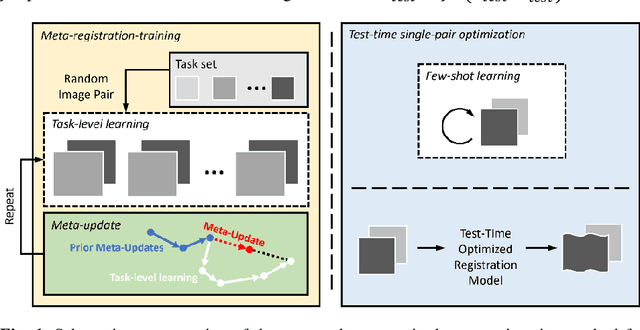
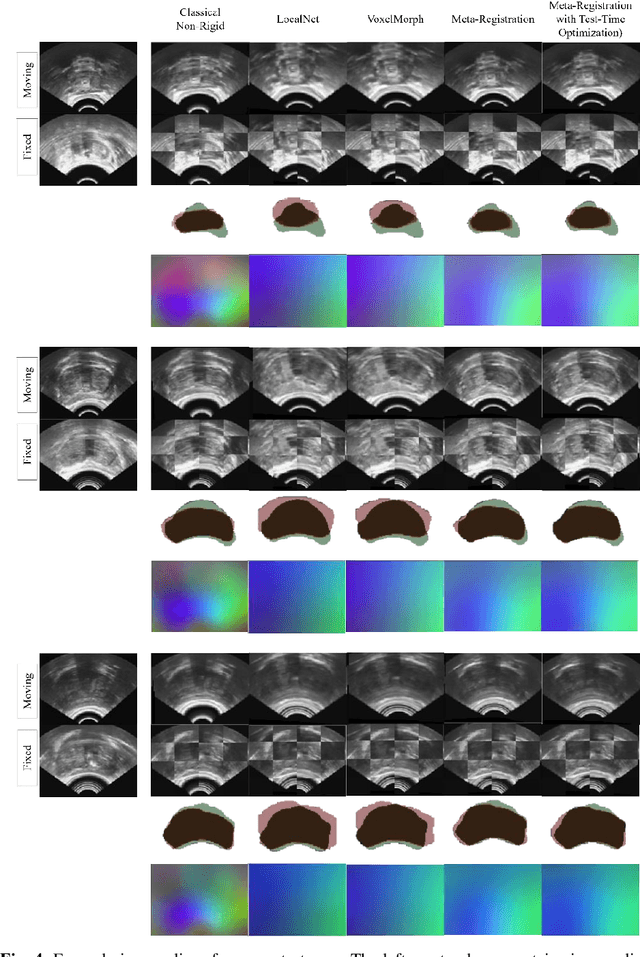
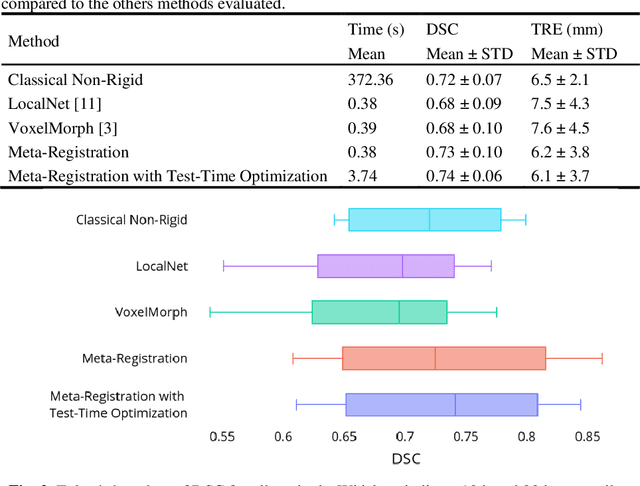
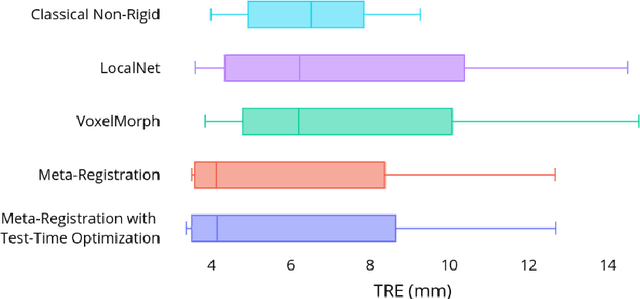

Abstract:Neural networks have been proposed for medical image registration by learning, with a substantial amount of training data, the optimal transformations between image pairs. These trained networks can further be optimized on a single pair of test images - known as test-time optimization. This work formulates image registration as a meta-learning algorithm. Such networks can be trained by aligning the training image pairs while simultaneously improving test-time optimization efficacy; tasks which were previously considered two independent training and optimization processes. The proposed meta-registration is hypothesized to maximize the efficiency and effectiveness of the test-time optimization in the "outer" meta-optimization of the networks. For image guidance applications that often are time-critical yet limited in training data, the potentially gained speed and accuracy are compared with classical registration algorithms, registration networks without meta-learning, and single-pair optimization without test-time optimization data. Experiments are presented in this paper using clinical transrectal ultrasound image data from 108 prostate cancer patients. These experiments demonstrate the effectiveness of a meta-registration protocol, which yields significantly improved performance relative to existing learning-based methods. Furthermore, the meta-registration achieves comparable results to classical iterative methods in a fraction of the time, owing to its rapid test-time optimization process.
Learning Generalized Non-Rigid Multimodal Biomedical Image Registration from Generic Point Set Data
Jul 22, 2022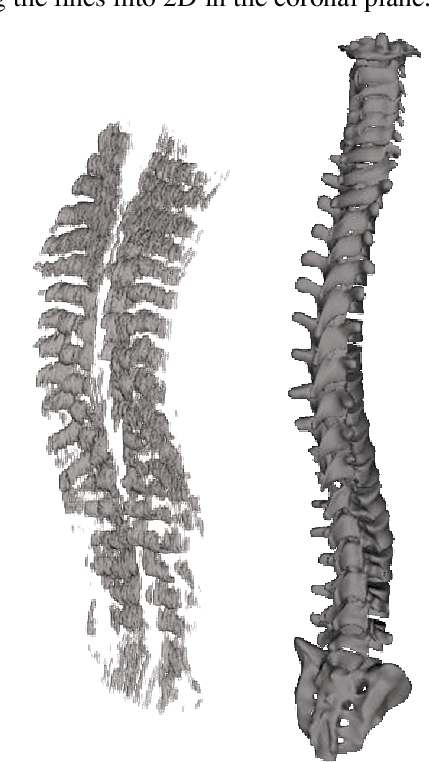
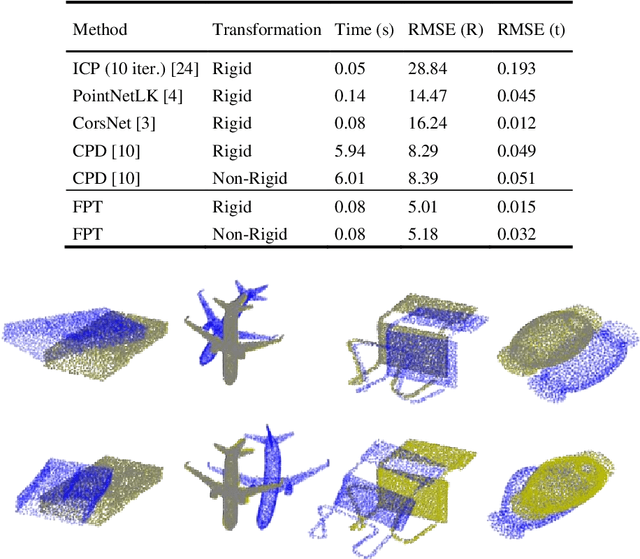
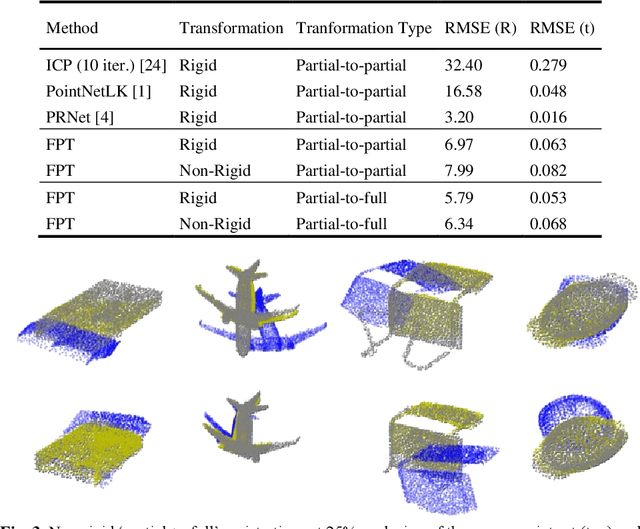
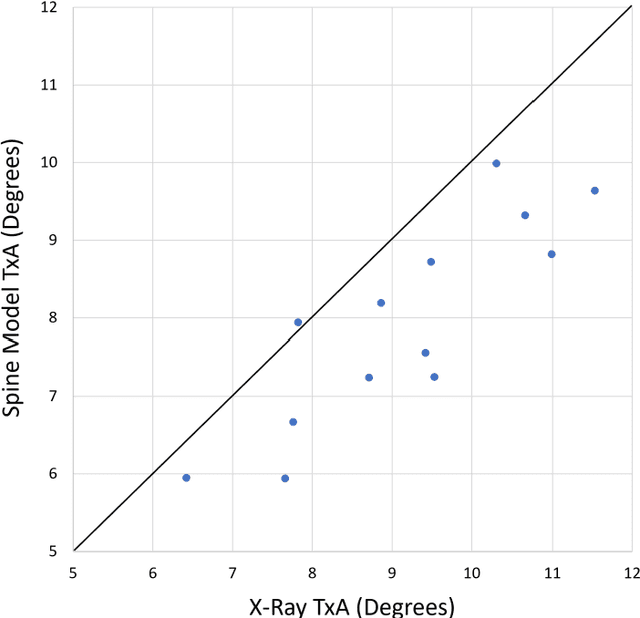
Abstract:Free Point Transformer (FPT) has been proposed as a data-driven, non-rigid point set registration approach using deep neural networks. As FPT does not assume constraints based on point vicinity or correspondence, it may be trained simply and in a flexible manner by minimizing an unsupervised loss based on the Chamfer Distance. This makes FPT amenable to real-world medical imaging applications where ground-truth deformations may be infeasible to obtain, or in scenarios where only a varying degree of completeness in the point sets to be aligned is available. To test the limit of the correspondence finding ability of FPT and its dependency on training data sets, this work explores the generalizability of the FPT from well-curated non-medical data sets to medical imaging data sets. First, we train FPT on the ModelNet40 dataset to demonstrate its effectiveness and the superior registration performance of FPT over iterative and learning-based point set registration methods. Second, we demonstrate superior performance in rigid and non-rigid registration and robustness to missing data. Last, we highlight the interesting generalizability of the ModelNet-trained FPT by registering reconstructed freehand ultrasound scans of the spine and generic spine models without additional training, whereby the average difference to the ground truth curvatures is 1.3 degrees, across 13 patients.
Real-time multimodal image registration with partial intraoperative point-set data
Sep 20, 2021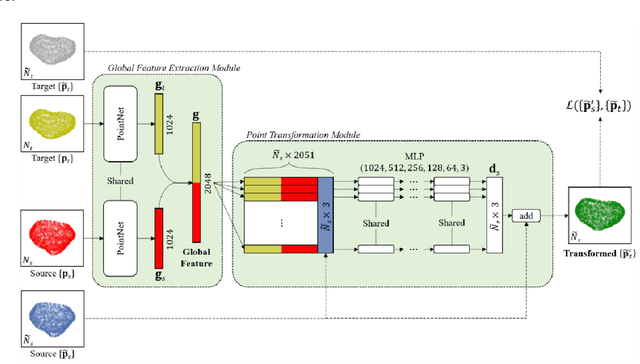
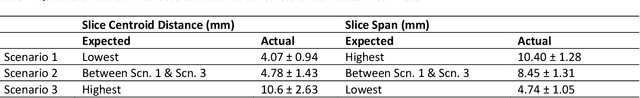

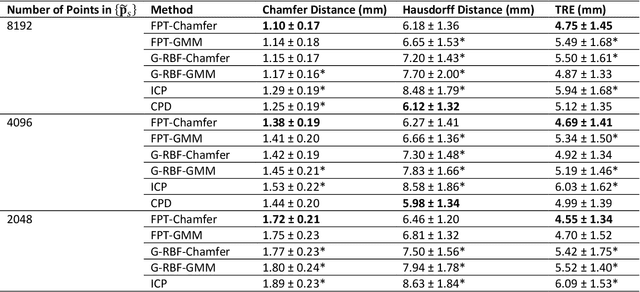
Abstract:We present Free Point Transformer (FPT) - a deep neural network architecture for non-rigid point-set registration. Consisting of two modules, a global feature extraction module and a point transformation module, FPT does not assume explicit constraints based on point vicinity, thereby overcoming a common requirement of previous learning-based point-set registration methods. FPT is designed to accept unordered and unstructured point-sets with a variable number of points and uses a "model-free" approach without heuristic constraints. Training FPT is flexible and involves minimizing an intuitive unsupervised loss function, but supervised, semi-supervised, and partially- or weakly-supervised training are also supported. This flexibility makes FPT amenable to multimodal image registration problems where the ground-truth deformations are difficult or impossible to measure. In this paper, we demonstrate the application of FPT to non-rigid registration of prostate magnetic resonance (MR) imaging and sparsely-sampled transrectal ultrasound (TRUS) images. The registration errors were 4.71 mm and 4.81 mm for complete TRUS imaging and sparsely-sampled TRUS imaging, respectively. The results indicate superior accuracy to the alternative rigid and non-rigid registration algorithms tested and substantially lower computation time. The rapid inference possible with FPT makes it particularly suitable for applications where real-time registration is beneficial.
Image quality assessment for closed-loop computer-assisted lung ultrasound
Aug 20, 2020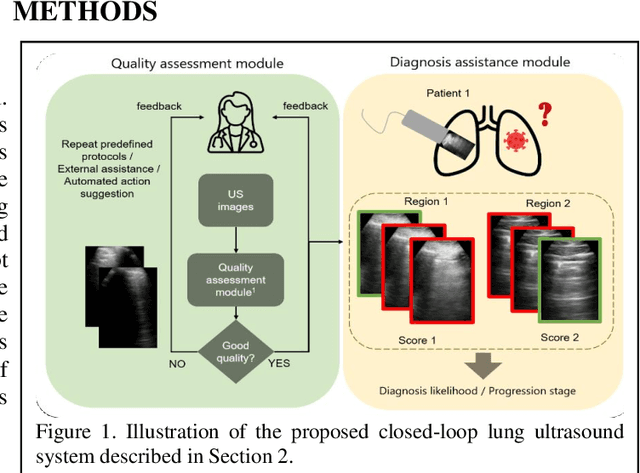
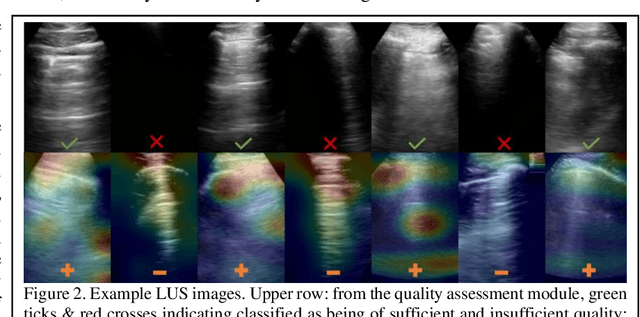
Abstract:We describe a novel, two-stage computer assistance system for lung anomaly detection using ultrasound imaging in the intensive care setting to improve operator performance and patient stratification during coronavirus pandemics. The proposed system consists of two deep-learning-based models. A quality assessment module automates predictions of image quality, and a diagnosis assistance module determines the likelihood-of-anomaly in ultrasound images of sufficient quality. Our two-stage strategy uses a novelty detection algorithm to address the lack of control cases available for training a quality assessment classifier. The diagnosis assistance module can then be trained with data that are deemed of sufficient quality, guaranteed by the closed-loop feedback mechanism from the quality assessment module. Integrating the two modules yields accurate, fast, and practical acquisition guidance and diagnostic assistance for patients with suspected respiratory conditions at the point-of-care. Using more than 25,000 ultrasound images from 37 COVID-19-positive patients scanned at two hospitals, plus 12 control cases, this study demonstrates the feasibility of using the proposed machine learning approach. We report an accuracy of 86% when classifying between sufficient and insufficient quality images by the quality assessment module. For data of sufficient quality, the mean classification accuracy in detecting COVID-19-positive cases was 95% on five holdout test data sets, unseen during the training of any networks within the proposed system.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge