Hua Li
DongYuan: An LLM-Based Framework for Integrative Chinese and Western Medicine Spleen-Stomach Disorders Diagnosis
Mar 30, 2026Abstract:The clinical burden of spleen-stomach disorders is substantial. While large language models (LLMs) offer new potential for medical applications, they face three major challenges in the context of integrative Chinese and Western medicine (ICWM): a lack of high-quality data, the absence of models capable of effectively integrating the reasoning logic of traditional Chinese medicine (TCM) syndrome differentiation with that of Western medical (WM) disease diagnosis, and the shortage of a standardized evaluation benchmark. To address these interrelated challenges, we propose DongYuan, an ICWM spleen-stomach diagnostic framework. Specifically, three ICWM datasets (SSDF-Syndrome, SSDF-Dialogue, and SSDF-PD) were curated to fill the gap in high-quality data for spleen-stomach disorders. We then developed SSDF-Core, a core diagnostic LLM that acquires robust ICWM reasoning capabilities through a two-stage training regimen of supervised fine-tuning. tuning (SFT) and direct preference optimization (DPO), and complemented it with SSDF-Navigator, a pluggable consultation navigation model designed to optimize clinical inquiry strategies. Additionally, we established SSDF-Bench, a comprehensive evaluation benchmark focused on ICWM diagnosis of spleen-stomach disorders. Experimental results demonstrate that SSDF-Core significantly outperforms 12 mainstream baselines on SSDF-Bench. DongYuan lays a solid methodological foundation and provides practical technical references for the future development of intelligent ICWM diagnostic systems.
PerformRecast: Expression and Head Pose Disentanglement for Portrait Video Editing
Mar 20, 2026Abstract:This paper primarily investigates the task of expression-only portrait video performance editing based on a driving video, which plays a crucial role in animation and film industries. Most existing research mainly focuses on portrait animation, which aims to animate a static portrait image according to the facial motion from the driving video. As a consequence, it remains challenging for them to disentangle the facial expression from head pose rotation and thus lack the ability to edit facial expression independently. In this paper, we propose PerformRecast, a versatile expression-only video editing method which is dedicated to recast the performance in existing film and animation. The key insight of our method comes from the characteristics of 3D Morphable Face Model (3DMM), which models the face identity, facial expression and head pose of 3D face mesh with separate parameters. Therefore, we improve the keypoints transformation formula in previous methods to make it more consistent with 3DMM model, which achieves a better disentanglement and provides users with much more fine-grained control. Furthermore, to avoid the misalignment around the boundary of face in generated results, we decouple the facial and non-facial regions of input portrait images and pre-train a teacher model to provide separate supervision for them. Extensive experiments show that our method produces high-quality results which are more faithful to the driving video, outperforming existing methods in both controllability and efficiency. Our code, data and trained models are available at https://youku-aigc.github.io/PerformRecast.
Fine-Tuning Diffusion-Based Recommender Systems via Reinforcement Learning with Reward Function Optimization
Nov 10, 2025Abstract:Diffusion models recently emerged as a powerful paradigm for recommender systems, offering state-of-the-art performance by modeling the generative process of user-item interactions. However, training such models from scratch is both computationally expensive and yields diminishing returns once convergence is reached. To remedy these challenges, we propose ReFiT, a new framework that integrates Reinforcement learning (RL)-based Fine-Tuning into diffusion-based recommender systems. In contrast to prior RL approaches for diffusion models depending on external reward models, ReFiT adopts a task-aligned design: it formulates the denoising trajectory as a Markov decision process (MDP) and incorporates a collaborative signal-aware reward function that directly reflects recommendation quality. By tightly coupling the MDP structure with this reward signal, ReFiT empowers the RL agent to exploit high-order connectivity for fine-grained optimization, while avoiding the noisy or uninformative feedback common in naive reward designs. Leveraging policy gradient optimization, ReFiT maximizes exact log-likelihood of observed interactions, thereby enabling effective post hoc fine-tuning of diffusion recommenders. Comprehensive experiments on wide-ranging real-world datasets demonstrate that the proposed ReFiT framework (a) exhibits substantial performance gains over strong competitors (up to 36.3% on sequential recommendation), (b) demonstrates strong efficiency with linear complexity in the number of users or items, and (c) generalizes well across multiple diffusion-based recommendation scenarios. The source code and datasets are publicly available at https://anonymous.4open.science/r/ReFiT-4C60.
WaterFlow: Explicit Physics-Prior Rectified Flow for Underwater Saliency Mask Generation
Oct 14, 2025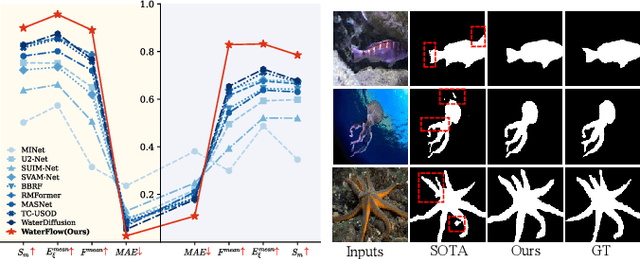
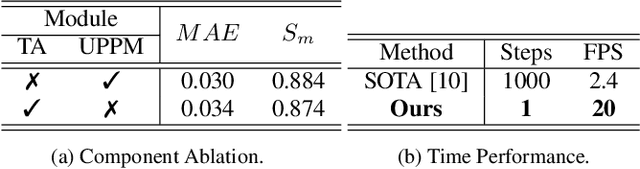
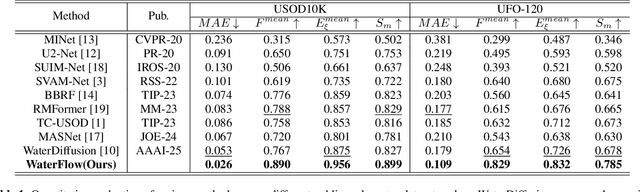
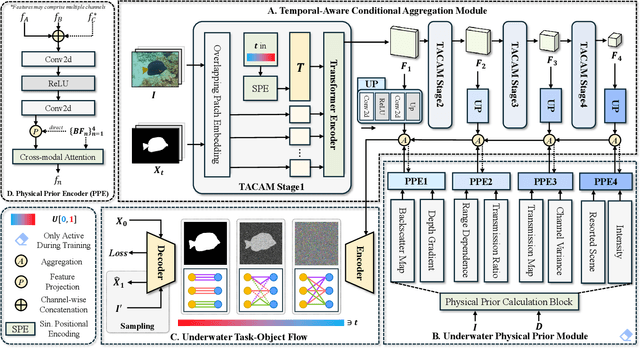

Abstract:Underwater Salient Object Detection (USOD) faces significant challenges, including underwater image quality degradation and domain gaps. Existing methods tend to ignore the physical principles of underwater imaging or simply treat degradation phenomena in underwater images as interference factors that must be eliminated, failing to fully exploit the valuable information they contain. We propose WaterFlow, a rectified flow-based framework for underwater salient object detection that innovatively incorporates underwater physical imaging information as explicit priors directly into the network training process and introduces temporal dimension modeling, significantly enhancing the model's capability for salient object identification. On the USOD10K dataset, WaterFlow achieves a 0.072 gain in S_m, demonstrating the effectiveness and superiority of our method. The code will be published after the acceptance.
UWSAM: Segment Anything Model Guided Underwater Instance Segmentation and A Large-scale Benchmark Dataset
May 21, 2025Abstract:With recent breakthroughs in large-scale modeling, the Segment Anything Model (SAM) has demonstrated significant potential in a variety of visual applications. However, due to the lack of underwater domain expertise, SAM and its variants face performance limitations in end-to-end underwater instance segmentation tasks, while their higher computational requirements further hinder their application in underwater scenarios. To address this challenge, we propose a large-scale underwater instance segmentation dataset, UIIS10K, which includes 10,048 images with pixel-level annotations for 10 categories. Then, we introduce UWSAM, an efficient model designed for automatic and accurate segmentation of underwater instances. UWSAM efficiently distills knowledge from the SAM ViT-Huge image encoder into the smaller ViT-Small image encoder via the Mask GAT-based Underwater Knowledge Distillation (MG-UKD) method for effective visual representation learning. Furthermore, we design an End-to-end Underwater Prompt Generator (EUPG) for UWSAM, which automatically generates underwater prompts instead of explicitly providing foreground points or boxes as prompts, thus enabling the network to locate underwater instances accurately for efficient segmentation. Comprehensive experimental results show that our model is effective, achieving significant performance improvements over state-of-the-art methods on multiple underwater instance datasets. Datasets and codes are available at https://github.com/LiamLian0727/UIIS10K.
TUGS: Physics-based Compact Representation of Underwater Scenes by Tensorized Gaussian
May 12, 2025Abstract:Underwater 3D scene reconstruction is crucial for undewater robotic perception and navigation. However, the task is significantly challenged by the complex interplay between light propagation, water medium, and object surfaces, with existing methods unable to model their interactions accurately. Additionally, expensive training and rendering costs limit their practical application in underwater robotic systems. Therefore, we propose Tensorized Underwater Gaussian Splatting (TUGS), which can effectively solve the modeling challenges of the complex interactions between object geometries and water media while achieving significant parameter reduction. TUGS employs lightweight tensorized higher-order Gaussians with a physics-based underwater Adaptive Medium Estimation (AME) module, enabling accurate simulation of both light attenuation and backscatter effects in underwater environments. Compared to other NeRF-based and GS-based methods designed for underwater, TUGS is able to render high-quality underwater images with faster rendering speeds and less memory usage. Extensive experiments on real-world underwater datasets have demonstrated that TUGS can efficiently achieve superior reconstruction quality using a limited number of parameters, making it particularly suitable for memory-constrained underwater UAV applications
Mamba Based Feature Extraction And Adaptive Multilevel Feature Fusion For 3D Tumor Segmentation From Multi-modal Medical Image
Apr 30, 2025Abstract:Multi-modal 3D medical image segmentation aims to accurately identify tumor regions across different modalities, facing challenges from variations in image intensity and tumor morphology. Traditional convolutional neural network (CNN)-based methods struggle with capturing global features, while Transformers-based methods, despite effectively capturing global context, encounter high computational costs in 3D medical image segmentation. The Mamba model combines linear scalability with long-distance modeling, making it a promising approach for visual representation learning. However, Mamba-based 3D multi-modal segmentation still struggles to leverage modality-specific features and fuse complementary information effectively. In this paper, we propose a Mamba based feature extraction and adaptive multilevel feature fusion for 3D tumor segmentation using multi-modal medical image. We first develop the specific modality Mamba encoder to efficiently extract long-range relevant features that represent anatomical and pathological structures present in each modality. Moreover, we design an bi-level synergistic integration block that dynamically merges multi-modal and multi-level complementary features by the modality attention and channel attention learning. Lastly, the decoder combines deep semantic information with fine-grained details to generate the tumor segmentation map. Experimental results on medical image datasets (PET/CT and MRI multi-sequence) show that our approach achieve competitive performance compared to the state-of-the-art CNN, Transformer, and Mamba-based approaches.
Spectrally-Corrected and Regularized QDA Classifier for Spiked Covariance Model
Mar 17, 2025Abstract:Quadratic discriminant analysis (QDA) is a widely used method for classification problems, particularly preferable over Linear Discriminant Analysis (LDA) for heterogeneous data. However, QDA loses its effectiveness in high-dimensional settings, where the data dimension and sample size tend to infinity. To address this issue, we propose a novel QDA method utilizing spectral correction and regularization techniques, termed SR-QDA. The regularization parameters in our method are selected by maximizing the Fisher-discriminant ratio. We compare SR-QDA with QDA, regularized quadratic discriminant analysis (R-QDA), and several other competitors. The results indicate that SR-QDA performs exceptionally well, especially in moderate and high-dimensional situations. Empirical experiments across diverse datasets further support this conclusion.
Estimating Task-based Performance Bounds for Accelerated MRI Image Reconstruction Methods by Use of Learned-Ideal Observers
Jan 16, 2025



Abstract:Medical imaging systems are commonly assessed and optimized by the use of objective measures of image quality (IQ). The performance of the ideal observer (IO) acting on imaging measurements has long been advocated as a figure-of-merit to guide the optimization of imaging systems. For computed imaging systems, the performance of the IO acting on imaging measurements also sets an upper bound on task-performance that no image reconstruction method can transcend. As such, estimation of IO performance can provide valuable guidance when designing under-sampled data-acquisition techniques by enabling the identification of designs that will not permit the reconstruction of diagnostically inappropriate images for a specified task - no matter how advanced the reconstruction method is or how plausible the reconstructed images appear. The need for such analysis is urgent because of the substantial increase of medical device submissions on deep learning-based image reconstruction methods and the fact that they may produce clean images disguising the potential loss of diagnostic information when data is aggressively under-sampled. Recently, convolutional neural network (CNN) approximated IOs (CNN-IOs) was investigated for estimating the performance of data space IOs to establish task-based performance bounds for image reconstruction, under an X-ray computed tomographic (CT) context. In this work, the application of such data space CNN-IO analysis to multi-coil magnetic resonance imaging (MRI) systems has been explored. This study utilized stylized multi-coil sensitivity encoding (SENSE) MRI systems and deep-generated stochastic brain models to demonstrate the approach. Signal-known-statistically and background-known-statistically (SKS/BKS) binary signal detection tasks were selected to study the impact of different acceleration factors on the data space IO performance.
HFH-Font: Few-shot Chinese Font Synthesis with Higher Quality, Faster Speed, and Higher Resolution
Oct 09, 2024Abstract:The challenge of automatically synthesizing high-quality vector fonts, particularly for writing systems (e.g., Chinese) consisting of huge amounts of complex glyphs, remains unsolved. Existing font synthesis techniques fall into two categories: 1) methods that directly generate vector glyphs, and 2) methods that initially synthesize glyph images and then vectorize them. However, the first category often fails to construct complete and correct shapes for complex glyphs, while the latter struggles to efficiently synthesize high-resolution (i.e., 1024 $\times$ 1024 or higher) glyph images while preserving local details. In this paper, we introduce HFH-Font, a few-shot font synthesis method capable of efficiently generating high-resolution glyph images that can be converted into high-quality vector glyphs. More specifically, our method employs a diffusion model-based generative framework with component-aware conditioning to learn different levels of style information adaptable to varying input reference sizes. We also design a distillation module based on Score Distillation Sampling for 1-step fast inference, and a style-guided super-resolution module to refine and upscale low-resolution synthesis results. Extensive experiments, including a user study with professional font designers, have been conducted to demonstrate that our method significantly outperforms existing font synthesis approaches. Experimental results show that our method produces high-fidelity, high-resolution raster images which can be vectorized into high-quality vector fonts. Using our method, for the first time, large-scale Chinese vector fonts of a quality comparable to those manually created by professional font designers can be automatically generated.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge