Honglin Li
Multi-modal Learning with Missing Modality in Predicting Axillary Lymph Node Metastasis
Jan 03, 2024



Abstract:Multi-modal Learning has attracted widespread attention in medical image analysis. Using multi-modal data, whole slide images (WSIs) and clinical information, can improve the performance of deep learning models in the diagnosis of axillary lymph node metastasis. However, clinical information is not easy to collect in clinical practice due to privacy concerns, limited resources, lack of interoperability, etc. Although patient selection can ensure the training set to have multi-modal data for model development, missing modality of clinical information can appear during test. This normally leads to performance degradation, which limits the use of multi-modal models in the clinic. To alleviate this problem, we propose a bidirectional distillation framework consisting of a multi-modal branch and a single-modal branch. The single-modal branch acquires the complete multi-modal knowledge from the multi-modal branch, while the multi-modal learns the robust features of WSI from the single-modal. We conduct experiments on a public dataset of Lymph Node Metastasis in Early Breast Cancer to validate the method. Our approach not only achieves state-of-the-art performance with an AUC of 0.861 on the test set without missing data, but also yields an AUC of 0.842 when the rate of missing modality is 80\%. This shows the effectiveness of the approach in dealing with multi-modal data and missing modality. Such a model has the potential to improve treatment decision-making for early breast cancer patients who have axillary lymph node metastatic status.
MI-Gen: Multiple Instance Generation of Pathology Reports for Gigapixel Whole-Slide Images
Nov 27, 2023



Abstract:Whole slide images are the foundation of digital pathology for the diagnosis and treatment of carcinomas. Writing pathology reports is laborious and error-prone for inexperienced pathologists. To reduce the workload and improve clinical automation, we investigate how to generate pathology reports given whole slide images. On the data end, we curated the largest WSI-text dataset (TCGA-PathoText). In specific, we collected nearly 10000 high-quality WSI-text pairs for visual-language models by recognizing and cleaning pathology reports which narrate diagnostic slides in TCGA. On the model end, we propose the multiple instance generative model (MI-Gen) which can produce pathology reports for gigapixel WSIs. We benchmark our model on the largest subset of TCGA-PathoText. Experimental results show our model can generate pathology reports which contain multiple clinical clues. Furthermore, WSI-text prediction can be seen as an approach of visual-language pre-training, which enables our model to be transferred to downstream diagnostic tasks like carcinoma grading and phenotyping. We observe that simple semantic extraction from the pathology reports can achieve the best performance (0.838 of F1 score) on BRCA subtyping without adding extra parameters or tricky fine-tuning. Our collected dataset and related code will all be publicly available.
Long-MIL: Scaling Long Contextual Multiple Instance Learning for Histopathology Whole Slide Image Analysis
Nov 21, 2023Abstract:Histopathology image analysis is the golden standard of clinical diagnosis for Cancers. In doctors daily routine and computer-aided diagnosis, the Whole Slide Image (WSI) of histopathology tissue is used for analysis. Because of the extremely large scale of resolution, previous methods generally divide the WSI into a large number of patches, then aggregate all patches within a WSI by Multi-Instance Learning (MIL) to make the slide-level prediction when developing computer-aided diagnosis tools. However, most previous WSI-MIL models using global-attention without pairwise interaction and any positional information, or self-attention with absolute position embedding can not well handle shape varying large WSIs, e.g. testing WSIs after model deployment may be larger than training WSIs, since the model development set is always limited due to the difficulty of histopathology WSIs collection. To deal with the problem, in this paper, we propose to amend position embedding for shape varying long-contextual WSI by introducing Linear Bias into Attention, and adapt it from 1-d long sequence into 2-d long-contextual WSI which helps model extrapolate position embedding to unseen or under-fitted positions. We further utilize Flash-Attention module to tackle the computational complexity of Transformer, which also keep full self-attention performance compared to previous attention approximation work. Our method, Long-contextual MIL (Long-MIL) are evaluated on extensive experiments including 4 dataset including WSI classification and survival prediction tasks to validate the superiority on shape varying WSIs. The source code will be open-accessed soon.
Attention-Challenging Multiple Instance Learning for Whole Slide Image Classification
Nov 13, 2023Abstract:Overfitting remains a significant challenge in the application of Multiple Instance Learning (MIL) methods for Whole Slide Image (WSI) analysis. Visualizing heatmaps reveals that current MIL methods focus on a subset of predictive instances, hindering effective model generalization. To tackle this, we propose Attention-Challenging MIL (ACMIL), aimed at forcing the attention mechanism to capture more challenging predictive instances. ACMIL incorporates two techniques, Multiple Branch Attention (MBA) to capture richer predictive instances and Stochastic Top-K Instance Masking (STKIM) to suppress simple predictive instances. Evaluation on three WSI datasets outperforms state-of-the-art methods. Additionally, through heatmap visualization, UMAP visualization, and attention value statistics, this paper comprehensively illustrates ACMIL's effectiveness in overcoming the overfitting challenge. The source code is available at \url{https://github.com/dazhangyu123/ACMIL}.
Masked conditional variational autoencoders for chromosome straightening
Jun 25, 2023



Abstract:Karyotyping is of importance for detecting chromosomal aberrations in human disease. However, chromosomes easily appear curved in microscopic images, which prevents cytogeneticists from analyzing chromosome types. To address this issue, we propose a framework for chromosome straightening, which comprises a preliminary processing algorithm and a generative model called masked conditional variational autoencoders (MC-VAE). The processing method utilizes patch rearrangement to address the difficulty in erasing low degrees of curvature, providing reasonable preliminary results for the MC-VAE. The MC-VAE further straightens the results by leveraging chromosome patches conditioned on their curvatures to learn the mapping between banding patterns and conditions. During model training, we apply a masking strategy with a high masking ratio to train the MC-VAE with eliminated redundancy. This yields a non-trivial reconstruction task, allowing the model to effectively preserve chromosome banding patterns and structure details in the reconstructed results. Extensive experiments on three public datasets with two stain styles show that our framework surpasses the performance of state-of-the-art methods in retaining banding patterns and structure details. Compared to using real-world bent chromosomes, the use of high-quality straightened chromosomes generated by our proposed method can improve the performance of various deep learning models for chromosome classification by a large margin. Such a straightening approach has the potential to be combined with other karyotyping systems to assist cytogeneticists in chromosome analysis.
Semi-supervised Cell Recognition under Point Supervision
Jun 14, 2023Abstract:Cell recognition is a fundamental task in digital histopathology image analysis. Point-based cell recognition (PCR) methods normally require a vast number of annotations, which is extremely costly, time-consuming and labor-intensive. Semi-supervised learning (SSL) can provide a shortcut to make full use of cell information in gigapixel whole slide images without exhaustive labeling. However, research into semi-supervised point-based cell recognition (SSPCR) remains largely overlooked. Previous SSPCR works are all built on density map-based PCR models, which suffer from unsatisfactory accuracy, slow inference speed and high sensitivity to hyper-parameters. To address these issues, end-to-end PCR models are proposed recently. In this paper, we develop a SSPCR framework suitable for the end-to-end PCR models for the first time. Overall, we use the current models to generate pseudo labels for unlabeled images, which are in turn utilized to supervise the models training. Besides, we introduce a co-teaching strategy to overcome the confirmation bias problem that generally exists in self-training. A distribution alignment technique is also incorporated to produce high-quality, unbiased pseudo labels for unlabeled data. Experimental results on four histopathology datasets concerning different types of staining styles show the effectiveness and versatility of the proposed framework. Code is available at \textcolor{magenta}{\url{https://github.com/windygooo/SSPCR}
PathAsst: Redefining Pathology through Generative Foundation AI Assistant for Pathology
May 24, 2023



Abstract:As advances in large language models (LLMs) and multimodal techniques continue to mature, the development of general-purpose multimodal large language models (MLLMs) has surged, with significant applications in natural image interpretation. However, the field of pathology has largely remained untapped in this regard, despite the growing need for accurate, timely, and personalized diagnostics. To bridge the gap in pathology MLLMs, we present the PathAsst in this study, which is a generative foundation AI assistant to revolutionize diagnostic and predictive analytics in pathology. To develop PathAsst, we collect over 142K high-quality pathology image-text pairs from a variety of reliable sources, including PubMed, comprehensive pathology textbooks, reputable pathology websites, and private data annotated by pathologists. Leveraging the advanced capabilities of ChatGPT/GPT-4, we generate over 180K instruction-following samples. Furthermore, we devise additional instruction-following data, specifically tailored for the invocation of the pathology-specific models, allowing the PathAsst to effectively interact with these models based on the input image and user intent, consequently enhancing the model's diagnostic capabilities. Subsequently, our PathAsst is trained based on Vicuna-13B language model in coordination with the CLIP vision encoder. The results of PathAsst show the potential of harnessing the AI-powered generative foundation model to improve pathology diagnosis and treatment processes. We are committed to open-sourcing our meticulously curated dataset, as well as a comprehensive toolkit designed to aid researchers in the extensive collection and preprocessing of their own datasets. Resources can be obtained at https://github.com/superjamessyx/Generative-Foundation-AI-Assistant-for-Pathology.
Task-specific Fine-tuning via Variational Information Bottleneck for Weakly-supervised Pathology Whole Slide Image Classification
Mar 15, 2023Abstract:While Multiple Instance Learning (MIL) has shown promising results in digital Pathology Whole Slide Image (WSI) classification, such a paradigm still faces performance and generalization problems due to challenges in high computational costs on Gigapixel WSIs and limited sample size for model training. To deal with the computation problem, most MIL methods utilize a frozen pretrained model from ImageNet to obtain representations first. This process may lose essential information owing to the large domain gap and hinder the generalization of model due to the lack of image-level training-time augmentations. Though Self-supervised Learning (SSL) proposes viable representation learning schemes, the improvement of the downstream task still needs to be further explored in the conversion from the task-agnostic features of SSL to the task-specifics under the partial label supervised learning. To alleviate the dilemma of computation cost and performance, we propose an efficient WSI fine-tuning framework motivated by the Information Bottleneck theory. The theory enables the framework to find the minimal sufficient statistics of WSI, thus supporting us to fine-tune the backbone into a task-specific representation only depending on WSI-level weak labels. The WSI-MIL problem is further analyzed to theoretically deduce our fine-tuning method. Our framework is evaluated on five pathology WSI datasets on various WSI heads. The experimental results of our fine-tuned representations show significant improvements in both accuracy and generalization compared with previous works. Source code will be available at https://github.com/invoker-LL/WSI-finetuning.
Deformable Proposal-Aware P2PNet: A Universal Network for Cell Recognition under Point Supervision
Mar 05, 2023



Abstract:Point-based cell recognition, which aims to localize and classify cells present in a pathology image, is a fundamental task in digital pathology image analysis. The recently developed point-to-point network (P2PNet) has achieved unprecedented cell recognition accuracy and efficiency compared to methods that rely on intermediate density map representations. However, P2PNet could not leverage multi-scale information since it can only decode a single feature map. Moreover, the distribution of predefined point proposals, which is determined by data properties, restricts the resolution of the feature map to decode, i.e., the encoder design. To lift these limitations, we propose a variant of P2PNet named deformable proposal-aware P2PNet (DPA-P2PNet) in this study. The proposed method uses coordinates of point proposals to directly extract multi-scale region-of-interest (ROI) features for feature enhancement. Such a design also opens up possibilities to exploit dynamic distributions of proposals. We further devise a deformation module to improve the proposal quality. Extensive experiments on four datasets with various staining styles demonstrate that DPA-P2PNet outperforms the state-of-the-art methods on point-based cell recognition, which reveals the high potentiality in assisting pathologist assessments.
Benchmarking the Robustness of Deep Neural Networks to Common Corruptions in Digital Pathology
Jun 30, 2022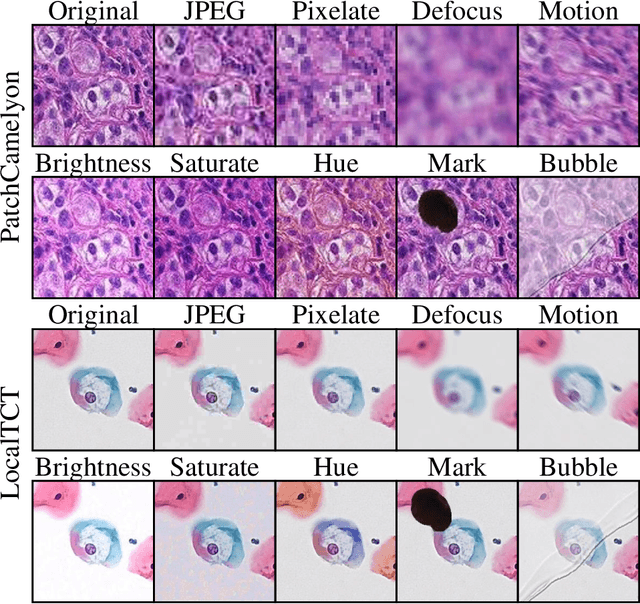
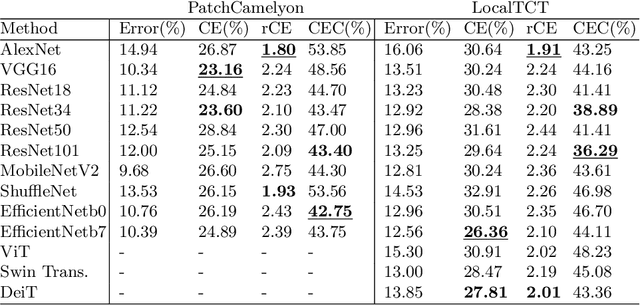
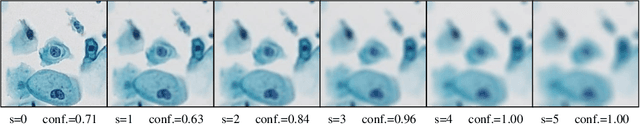
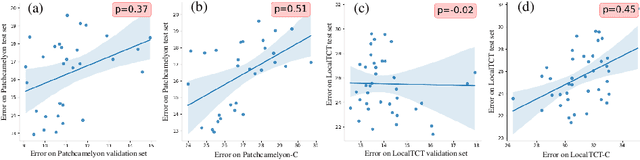
Abstract:When designing a diagnostic model for a clinical application, it is crucial to guarantee the robustness of the model with respect to a wide range of image corruptions. Herein, an easy-to-use benchmark is established to evaluate how deep neural networks perform on corrupted pathology images. Specifically, corrupted images are generated by injecting nine types of common corruptions into validation images. Besides, two classification and one ranking metrics are designed to evaluate the prediction and confidence performance under corruption. Evaluated on two resulting benchmark datasets, we find that (1) a variety of deep neural network models suffer from a significant accuracy decrease (double the error on clean images) and the unreliable confidence estimation on corrupted images; (2) A low correlation between the validation and test errors while replacing the validation set with our benchmark can increase the correlation. Our codes are available on https://github.com/superjamessyx/robustness_benchmark.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge