Haocheng Tang
Learning Structure, Energy, and Dynamics: A Survey of Artificial Intelligence for Protein Dynamics
Apr 28, 2026Abstract:Protein dynamics underlie many biological functions, yet remain difficult to characterize due to the high computational cost of molecular dynamics simulations and the scarcity of dynamic structural data. This survey reviews recent advances in artificial intelligence for protein dynamics from three perspectives: learning from structural ensembles and trajectories, learning from physical energy signals, and learning to accelerate molecular simulations. We summarize representative methods for conformation ensemble generation, trajectory generation, Boltzmann generators, physics-aware adaptation, machine learning potentials, coarse-grained modeling, and collective variable discovery. We further discuss available datasets and key open challenges, such as scalability, thermodynamic consistency, kinetic fidelity, and integration with experimental constraints.
Fine-tuning DeepSeek-OCR-2 for Molecular Structure Recognition
Apr 03, 2026Abstract:Optical Chemical Structure Recognition (OCSR) is critical for converting 2D molecular diagrams from printed literature into machine-readable formats. While Vision-Language Models have shown promise in end-to-end OCR tasks, their direct application to OCSR remains challenging, and direct full-parameter supervised fine-tuning often fails. In this work, we adapt DeepSeek-OCR-2 for molecular optical recognition by formulating the task as image-conditioned SMILES generation. To overcome training instabilities, we propose a two-stage progressive supervised fine-tuning strategy: starting with parameter-efficient LoRA and transitioning to selective full-parameter fine-tuning with split learning rates. We train our model on a large-scale corpus combining synthetic renderings from PubChem and realistic patent images from USPTO-MOL to improve coverage and robustness. Our fine-tuned model, MolSeek-OCR, demonstrates competitive capabilities, achieving exact matching accuracies comparable to the best-performing image-to-sequence model. However, it remains inferior to state-of-the-art image-to-graph modelS. Furthermore, we explore reinforcement-style post-training and data-curation-based refinement, finding that they fail to improve the strict sequence-level fidelity required for exact SMILES matching.
Parallax to Align Them All: An OmniParallax Attention Mechanism for Distributed Multi-View Image Compression
Mar 04, 2026Abstract:Multi-view image compression (MIC) aims to achieve high compression efficiency by exploiting inter-image correlations, playing a crucial role in 3D applications. As a subfield of MIC, distributed multi-view image compression (DMIC) offers performance comparable to MIC while eliminating the need for inter-view information at the encoder side. However, existing methods in DMIC typically treat all images equally, overlooking the varying degrees of correlation between different views during decoding, which leads to suboptimal coding performance. To address this limitation, we propose a novel $\textbf{OmniParallax Attention Mechanism}$ (OPAM), which is a general mechanism for explicitly modeling correlations and aligned features between arbitrary pairs of information sources. Building upon OPAM, we propose a Parallax Multi Information Fusion Module (PMIFM) to adaptively integrate information from different sources. PMIFM is incorporated into both the joint decoder and the entropy model to construct our end-to-end DMIC framework, $\textbf{ParaHydra}$. Extensive experiments demonstrate that $\textbf{ParaHydra}$ is $\textbf{the first DMIC method}$ to significantly surpass state-of-the-art MIC codecs, while maintaining low computational overhead. Performance gains become more pronounced as the number of input views increases. Compared with LDMIC, $\textbf{ParaHydra}$ achieves bitrate savings of $\textbf{19.72%}$ on WildTrack(3) and up to $\textbf{24.18%}$ on WildTrack(6), while significantly improving coding efficiency (as much as $\textbf{65}\times$ in decoding and $\textbf{34}\times$ in encoding).
Auxiliary Discrminator Sequence Generative Adversarial Networks (ADSeqGAN) for Few Sample Molecule Generation
Feb 23, 2025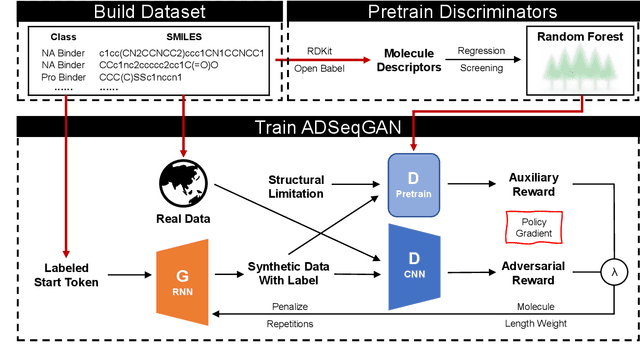



Abstract:In this work, we introduce Auxiliary Discriminator Sequence Generative Adversarial Networks (ADSeqGAN), a novel approach for molecular generation in small-sample datasets. Traditional generative models often struggle with limited training data, particularly in drug discovery, where molecular datasets for specific therapeutic targets, such as nucleic acids binders and central nervous system (CNS) drugs, are scarce. ADSeqGAN addresses this challenge by integrating an auxiliary random forest classifier as an additional discriminator into the GAN framework, significantly improves molecular generation quality and class specificity. Our method incorporates pretrained generator and Wasserstein distance to enhance training stability and diversity. We evaluate ADSeqGAN on a dataset comprising nucleic acid-targeting and protein-targeting small molecules, demonstrating its superior ability to generate nucleic acid binders compared to baseline models such as SeqGAN, ORGAN, and MolGPT. Through an oversampling strategy, ADSeqGAN also significantly improves CNS drug generation, achieving a higher yield than traditional de novo models. Critical assessments, including docking simulations and molecular property analysis, confirm that ADSeqGAN-generated molecules exhibit strong binding affinities, enhanced chemical diversity, and improved synthetic feasibility. Overall, ADSeqGAN presents a novel framework for generative molecular design in data-scarce scenarios, offering potential applications in computational drug discovery. We have demonstrated the successful applications of ADSeqGAN in generating synthetic nucleic acid-targeting and CNS drugs in this work.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge