Junmei Wang
Fine-tuning DeepSeek-OCR-2 for Molecular Structure Recognition
Apr 03, 2026Abstract:Optical Chemical Structure Recognition (OCSR) is critical for converting 2D molecular diagrams from printed literature into machine-readable formats. While Vision-Language Models have shown promise in end-to-end OCR tasks, their direct application to OCSR remains challenging, and direct full-parameter supervised fine-tuning often fails. In this work, we adapt DeepSeek-OCR-2 for molecular optical recognition by formulating the task as image-conditioned SMILES generation. To overcome training instabilities, we propose a two-stage progressive supervised fine-tuning strategy: starting with parameter-efficient LoRA and transitioning to selective full-parameter fine-tuning with split learning rates. We train our model on a large-scale corpus combining synthetic renderings from PubChem and realistic patent images from USPTO-MOL to improve coverage and robustness. Our fine-tuned model, MolSeek-OCR, demonstrates competitive capabilities, achieving exact matching accuracies comparable to the best-performing image-to-sequence model. However, it remains inferior to state-of-the-art image-to-graph modelS. Furthermore, we explore reinforcement-style post-training and data-curation-based refinement, finding that they fail to improve the strict sequence-level fidelity required for exact SMILES matching.
Ultrasound-CLIP: Semantic-Aware Contrastive Pre-training for Ultrasound Image-Text Understanding
Apr 02, 2026Abstract:Ultrasound imaging is widely used in clinical diagnostics due to its real-time capability and radiation-free nature. However, existing vision-language pre-training models, such as CLIP, are primarily designed for other modalities, and are difficult to directly apply to ultrasound data, which exhibit heterogeneous anatomical structures and diverse diagnostic attributes. To bridge this gap, we construct US-365K, a large-scale ultrasound image-text dataset containing 365k paired samples across 52 anatomical categories. We establish Ultrasonographic Diagnostic Taxonomy (UDT) containing two hierarchical knowledge frameworks. Ultrasonographic Hierarchical Anatomical Taxonomy standardizes anatomical organization, and Ultrasonographic Diagnostic Attribute Framework formalizes nine diagnostic dimensions, including body system, organ, diagnosis, shape, margins, echogenicity, internal characteristics, posterior acoustic phenomena, and vascularity. Building upon these foundations, we propose Ultrasound-CLIP, a semantic-aware contrastive learning framework that introduces semantic soft labels and semantic loss to refine sample discrimination. Moreover, we construct a heterogeneous graph modality derived from UDAF's textual representations, enabling structured reasoning over lesion-attribute relations. Extensive experiments with patient-level data splitting demonstrate that our approach achieves state-of-the-art performance on classification and retrieval benchmarks, while also delivering strong generalization to zero-shot, linear probing, and fine-tuning tasks.
Auxiliary Discrminator Sequence Generative Adversarial Networks (ADSeqGAN) for Few Sample Molecule Generation
Feb 23, 2025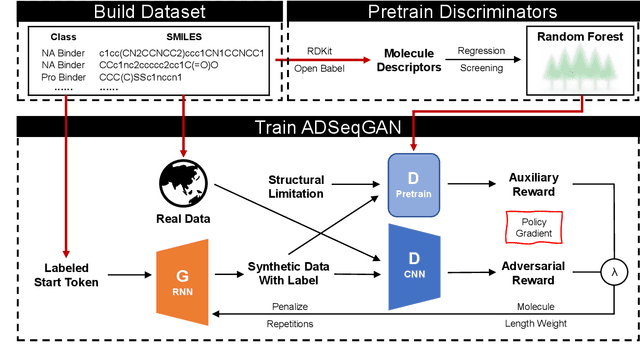



Abstract:In this work, we introduce Auxiliary Discriminator Sequence Generative Adversarial Networks (ADSeqGAN), a novel approach for molecular generation in small-sample datasets. Traditional generative models often struggle with limited training data, particularly in drug discovery, where molecular datasets for specific therapeutic targets, such as nucleic acids binders and central nervous system (CNS) drugs, are scarce. ADSeqGAN addresses this challenge by integrating an auxiliary random forest classifier as an additional discriminator into the GAN framework, significantly improves molecular generation quality and class specificity. Our method incorporates pretrained generator and Wasserstein distance to enhance training stability and diversity. We evaluate ADSeqGAN on a dataset comprising nucleic acid-targeting and protein-targeting small molecules, demonstrating its superior ability to generate nucleic acid binders compared to baseline models such as SeqGAN, ORGAN, and MolGPT. Through an oversampling strategy, ADSeqGAN also significantly improves CNS drug generation, achieving a higher yield than traditional de novo models. Critical assessments, including docking simulations and molecular property analysis, confirm that ADSeqGAN-generated molecules exhibit strong binding affinities, enhanced chemical diversity, and improved synthetic feasibility. Overall, ADSeqGAN presents a novel framework for generative molecular design in data-scarce scenarios, offering potential applications in computational drug discovery. We have demonstrated the successful applications of ADSeqGAN in generating synthetic nucleic acid-targeting and CNS drugs in this work.
Emerging Opportunities of Using Large Language Models for Translation Between Drug Molecules and Indications
Feb 16, 2024Abstract:A drug molecule is a substance that changes the organism's mental or physical state. Every approved drug has an indication, which refers to the therapeutic use of that drug for treating a particular medical condition. While the Large Language Model (LLM), a generative Artificial Intelligence (AI) technique, has recently demonstrated effectiveness in translating between molecules and their textual descriptions, there remains a gap in research regarding their application in facilitating the translation between drug molecules and indications, or vice versa, which could greatly benefit the drug discovery process. The capability of generating a drug from a given indication would allow for the discovery of drugs targeting specific diseases or targets and ultimately provide patients with better treatments. In this paper, we first propose a new task, which is the translation between drug molecules and corresponding indications, and then test existing LLMs on this new task. Specifically, we consider nine variations of the T5 LLM and evaluate them on two public datasets obtained from ChEMBL and DrugBank. Our experiments show the early results of using LLMs for this task and provide a perspective on the state-of-the-art. We also emphasize the current limitations and discuss future work that has the potential to improve the performance on this task. The creation of molecules from indications, or vice versa, will allow for more efficient targeting of diseases and significantly reduce the cost of drug discovery, with the potential to revolutionize the field of drug discovery in the era of generative AI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge