Dorin Comaniciu
Machine Learning Automatically Detects COVID-19 using Chest CTs in a Large Multicenter Cohort
Jun 11, 2020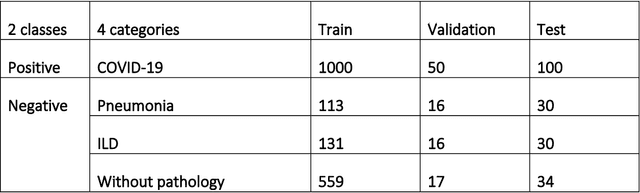
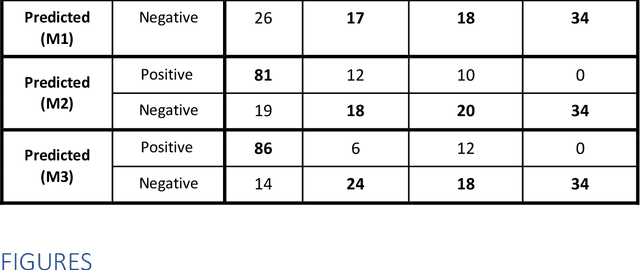
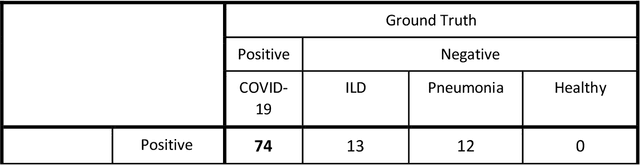

Abstract:Purpose: To investigate if AI-based classifiers can distinguish COVID-19 from other pulmonary diseases and normal groups, using chest CT images. To study the interpretability of discriminative features for COVID19 detection. Materials and Methods: Our database consists of 2096 CT exams that include CTs from 1150 COVID-19 patients. Training was performed on 1000 COVID-19, 131 ILD, 113 other pneumonias, 559 normal CTs, and testing on 100 COVID-19, 30 ILD, 30 other pneumonias, and 34 normal CTs. A metric-based approach for classification of COVID-19 used interpretable features, relying on logistic regression and random forests. A deep learning-based classifier differentiated COVID-19 based on 3D features extracted directly from CT intensities and from the probability distribution of airspace opacities. Results: Most discriminative features of COVID-19 are percentage of airspace opacity, ground glass opacities, consolidations, and peripheral and basal opacities, which coincide with the typical characterization of COVID-19 in the literature. Unsupervised hierarchical clustering compares the distribution of these features across COVID-19 and control cohorts. The metrics-based classifier achieved AUC, sensitivity, and specificity of respectively 0.85, 0.81, and 0.77. The DL-based classifier achieved AUC, sensitivity, and specificity of respectively 0.90, 0.86, and 0.81. Most of ambiguity comes from non-COVID-19 pneumonia with manifestations that overlap with COVID-19, as well as COVID-19 cases in early stages. Conclusion: A new method discriminates COVID-19 from other types of pneumonia, ILD, and normal, using quantitative patterns from chest CT. Our models balance interpretability of results and classification performance, and therefore may be useful to expedite and improve diagnosis of COVID-19.
3D Tomographic Pattern Synthesis for Enhancing the Quantification of COVID-19
May 05, 2020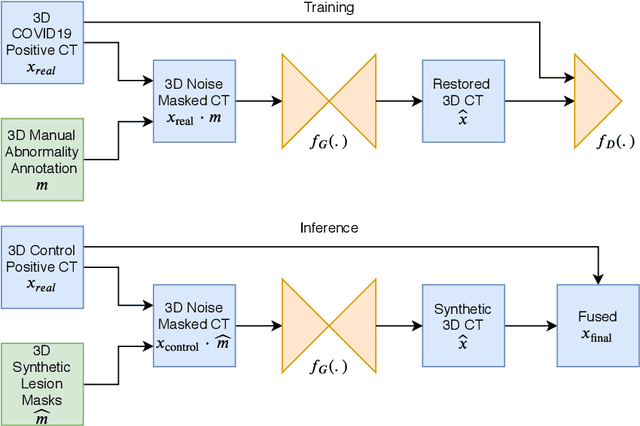
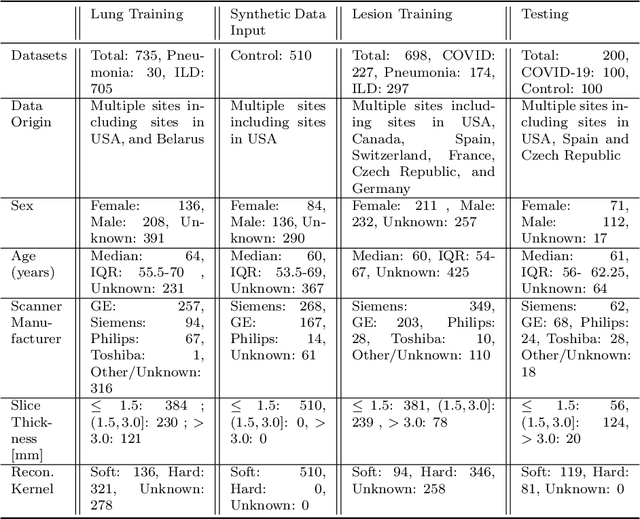
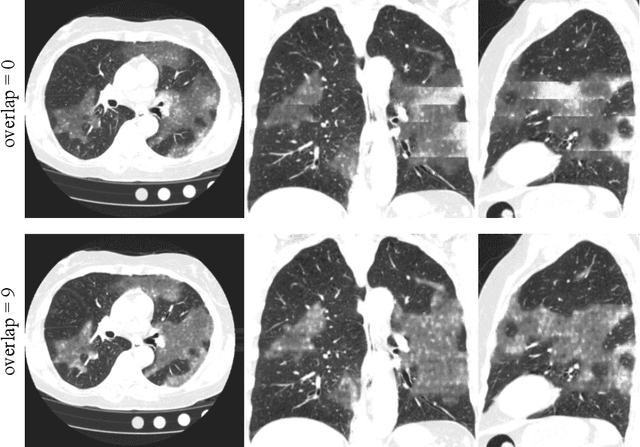
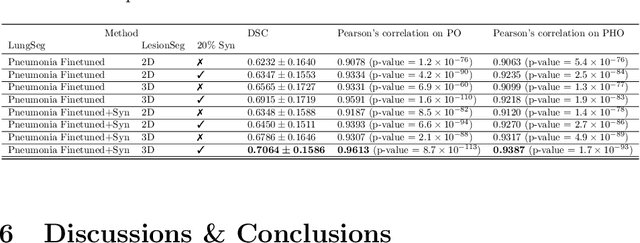
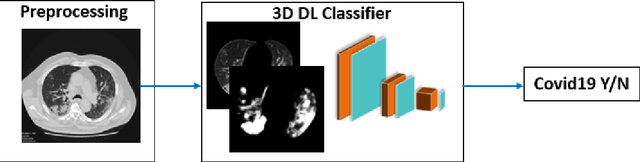
Abstract:The Coronavirus Disease (COVID-19) has affected 1.8 million people and resulted in more than 110,000 deaths as of April 12, 2020. Several studies have shown that tomographic patterns seen on chest Computed Tomography (CT), such as ground-glass opacities, consolidations, and crazy paving pattern, are correlated with the disease severity and progression. CT imaging can thus emerge as an important modality for the management of COVID-19 patients. AI-based solutions can be used to support CT based quantitative reporting and make reading efficient and reproducible if quantitative biomarkers, such as the Percentage of Opacity (PO), can be automatically computed. However, COVID-19 has posed unique challenges to the development of AI, specifically concerning the availability of appropriate image data and annotations at scale. In this paper, we propose to use synthetic datasets to augment an existing COVID-19 database to tackle these challenges. We train a Generative Adversarial Network (GAN) to inpaint COVID-19 related tomographic patterns on chest CTs from patients without infectious diseases. Additionally, we leverage location priors derived from manually labeled COVID-19 chest CTs patients to generate appropriate abnormality distributions. Synthetic data are used to improve both lung segmentation and segmentation of COVID-19 patterns by adding 20% of synthetic data to the real COVID-19 training data. We collected 2143 chest CTs, containing 327 COVID-19 positive cases, acquired from 12 sites across 7 countries. By testing on 100 COVID-19 positive and 100 control cases, we show that synthetic data can help improve both lung segmentation (+6.02% lesion inclusion rate) and abnormality segmentation (+2.78% dice coefficient), leading to an overall more accurate PO computation (+2.82% Pearson coefficient).
Quantification of Tomographic Patterns associated with COVID-19 from Chest CT
Apr 28, 2020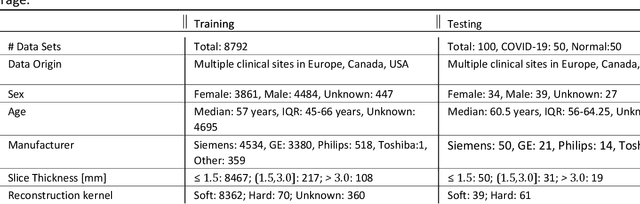
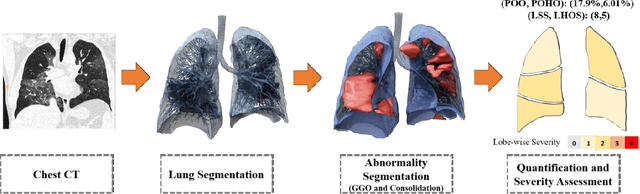
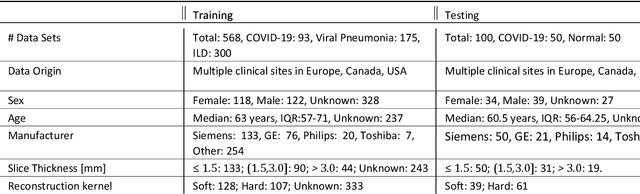
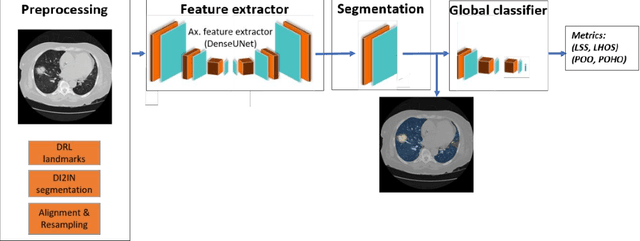
Abstract:Purpose: To present a method that automatically detects and quantifies abnormal tomographic patterns commonly present in COVID-19, namely Ground Glass Opacities (GGO) and consolidations. Given that high opacity abnormalities (i.e., consolidations) were shown to correlate with severe disease, the paper introduces two combined severity measures (Percentage of Opacity, Percentage of High Opacity) and (Lung Severity Score, Lung High Opacity Score). They quantify the extent of overall COVID-19 abnormalities and the presence of high opacity abnormalities, global and lobe-wise, respectively, being computed based on 3D segmentations of lesions, lungs, and lobes. Materials and Methods: The proposed method takes as input a non-contrasted Chest CT and segments the lesions, lungs, and lobes in 3D. It outputs two combined measures of the severity of lung/lobe involvement, quantifying both the extent of COVID-19 abnormalities and presence of high opacities, based on deep learning and deep reinforcement learning. The first measure (POO, POHO) is global, while the second (LSS, LHOS) is lobe-wise. Evaluation is reported on CTs of 100 subjects (50 COVID-19 confirmed and 50 controls) from institutions from Canada, Europe and US. Ground truth is established by manual annotations of lesions, lungs, and lobes. Results: Pearson Correlation Coefficient between method prediction and ground truth is 0.97 (POO), 0.98 (POHO), 0.96 (LSS), 0.96 (LHOS). Automated processing time to compute the severity scores is 10 seconds/case vs 30 mins needed for manual annotations. Conclusion: A new method identifies regions of abnormalities seen in COVID-19 non-contrasted Chest CT and computes (POO, POHO) and (LSS, LHOS) severity scores.
Graph Attention Network based Pruning for Reconstructing 3D Liver Vessel Morphology from Contrasted CT Images
Mar 18, 2020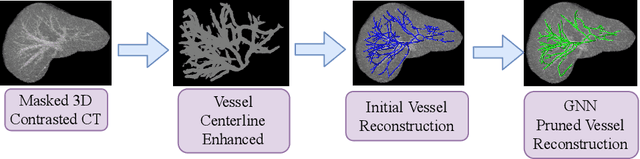
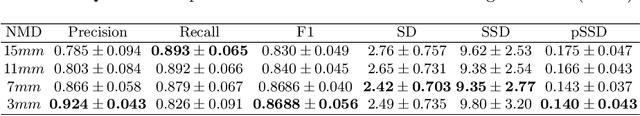
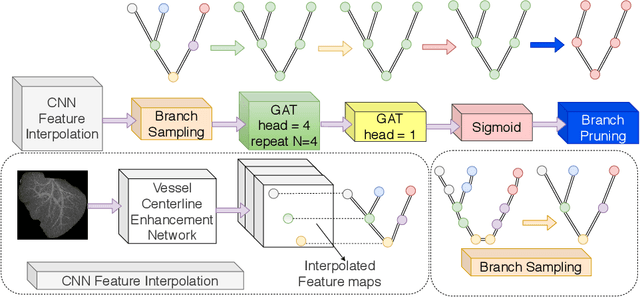
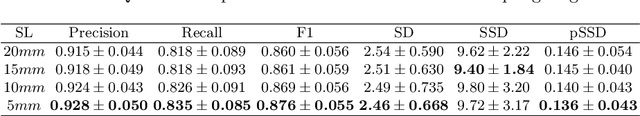
Abstract:With the injection of contrast material into blood vessels, multi-phase contrasted CT images can enhance the visibility of vessel networks in the human body. Reconstructing the 3D geometric morphology of liver vessels from the contrasted CT images can enable multiple liver preoperative surgical planning applications. Automatic reconstruction of liver vessel morphology remains a challenging problem due to the morphological complexity of liver vessels and the inconsistent vessel intensities among different multi-phase contrasted CT images. On the other side, high integrity is required for the 3D reconstruction to avoid decision making biases. In this paper, we propose a framework for liver vessel morphology reconstruction using both a fully convolutional neural network and a graph attention network. A fully convolutional neural network is first trained to produce the liver vessel centerline heatmap. An over-reconstructed liver vessel graph model is then traced based on the heatmap using an image processing based algorithm. We use a graph attention network to prune the false-positive branches by predicting the presence probability of each segmented branch in the initial reconstruction using the aggregated CNN features. We evaluated the proposed framework on an in-house dataset consisting of 418 multi-phase abdomen CT images with contrast. The proposed graph network pruning improves the overall reconstruction F1 score by 6.4% over the baseline. It also outperformed the other state-of-the-art curvilinear structure reconstruction algorithms.
No Surprises: Training Robust Lung Nodule Detection for Low-Dose CT Scans by Augmenting with Adversarial Attacks
Mar 08, 2020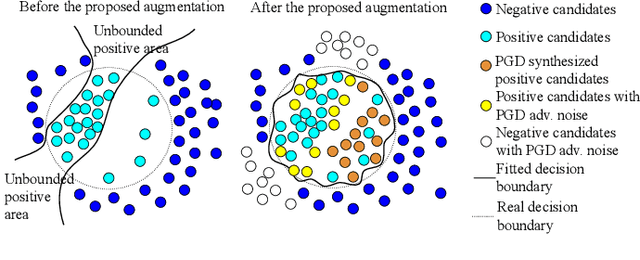
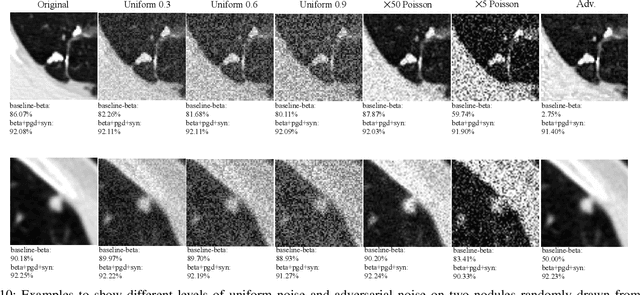

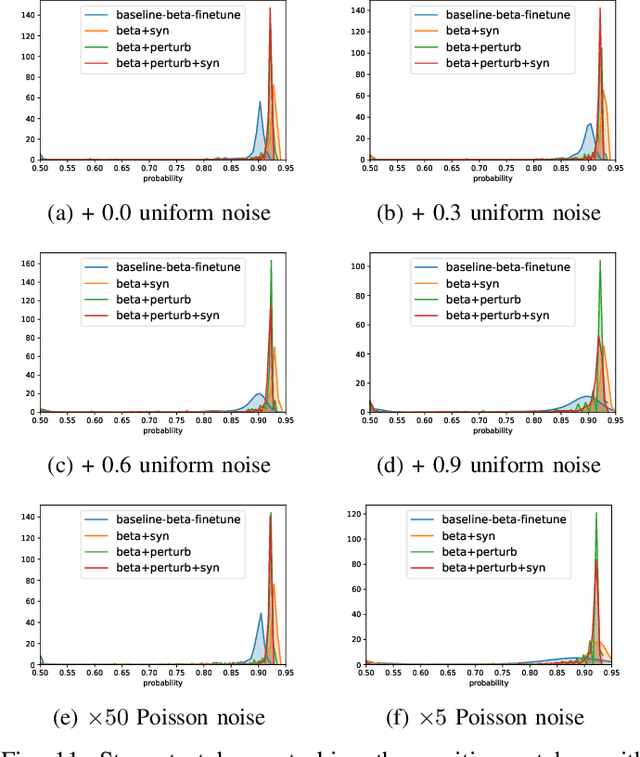
Abstract:Detecting malignant pulmonary nodules at an early stage can allow medical interventions which increases the survival rate of lung cancer patients. Using computer vision techniques to detect nodules can improve the sensitivity and the speed of interpreting chest CT for lung cancer screening. Many studies have used CNNs to detect nodule candidates. Though such approaches have been shown to outperform the conventional image processing based methods regarding the detection accuracy, CNNs are also known to be limited to generalize on under-represented samples in the training set and prone to imperceptible noise perturbations. Such limitations can not be easily addressed by scaling up the dataset or the models. In this work, we propose to add adversarial synthetic nodules and adversarial attack samples to the training data to improve the generalization and the robustness of the lung nodule detection systems. In order to generate hard examples of nodules from a differentiable nodule synthesizer, we use projected gradient descent (PGD) to search the latent code within a bounded neighbourhood that would generate nodules to decrease the detector response. To make the network more robust to unanticipated noise perturbations, we use PGD to search for noise patterns that can trigger the network to give over-confident mistakes. By evaluating on two different benchmark datasets containing consensus annotations from three radiologists, we show that the proposed techniques can improve the detection performance on real CT data. To understand the limitations of both the conventional networks and the proposed augmented networks, we also perform stress-tests on the false positive reduction networks by feeding different types of artificially produced patches. We show that the augmented networks are more robust to both under-represented nodules as well as resistant to noise perturbations.
Quantifying and Leveraging Classification Uncertainty for Chest Radiograph Assessment
Jun 18, 2019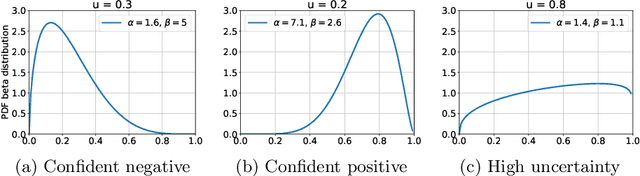
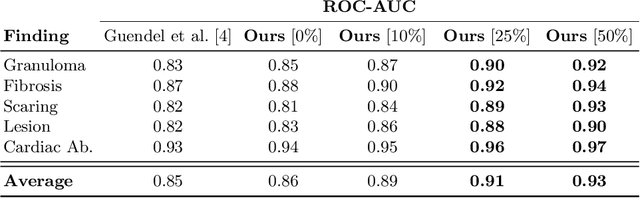
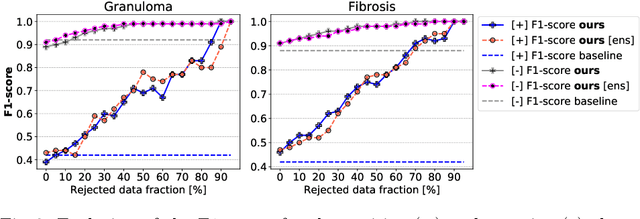
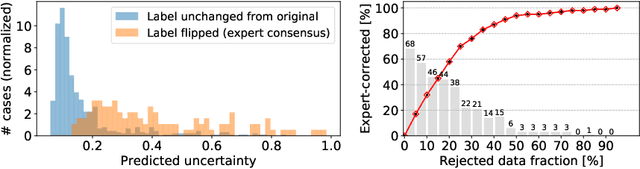
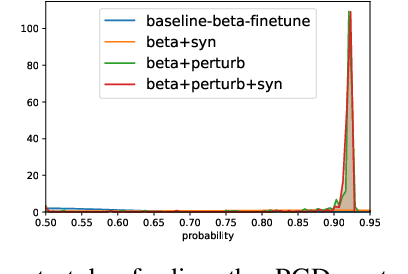
Abstract:The interpretation of chest radiographs is an essential task for the detection of thoracic diseases and abnormalities. However, it is a challenging problem with high inter-rater variability and inherent ambiguity due to inconclusive evidence in the data, limited data quality or subjective definitions of disease appearance. Current deep learning solutions for chest radiograph abnormality classification are typically limited to providing probabilistic predictions, relying on the capacity of learning models to adapt to the high degree of label noise and become robust to the enumerated causal factors. In practice, however, this leads to overconfident systems with poor generalization on unseen data. To account for this, we propose an automatic system that learns not only the probabilistic estimate on the presence of an abnormality, but also an explicit uncertainty measure which captures the confidence of the system in the predicted output. We argue that explicitly learning the classification uncertainty as an orthogonal measure to the predicted output, is essential to account for the inherent variability characteristic of this data. Experiments were conducted on two datasets of chest radiographs of over 85,000 patients. Sample rejection based on the predicted uncertainty can significantly improve the ROC-AUC, e.g., by 8% to 0.91 with an expected rejection rate of under 25%. Eliminating training samples using uncertainty-driven bootstrapping, enables a significant increase in robustness and accuracy. In addition, we present a multi-reader study showing that the predictive uncertainty is indicative of reader errors.
Multi-task Learning for Chest X-ray Abnormality Classification on Noisy Labels
May 15, 2019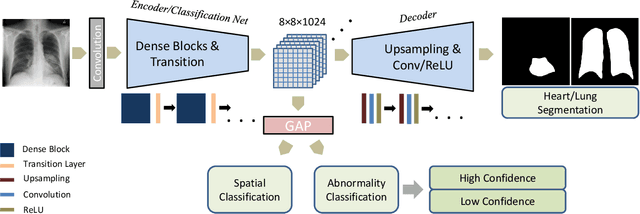
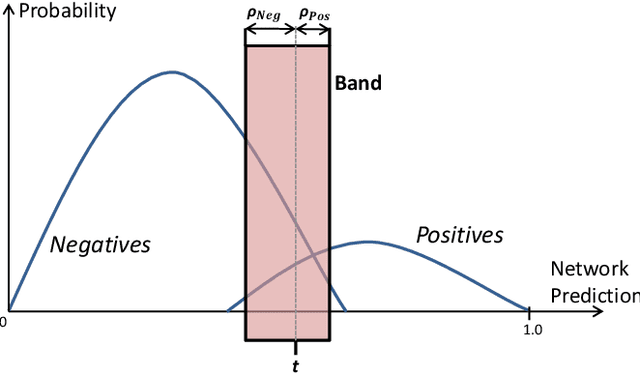
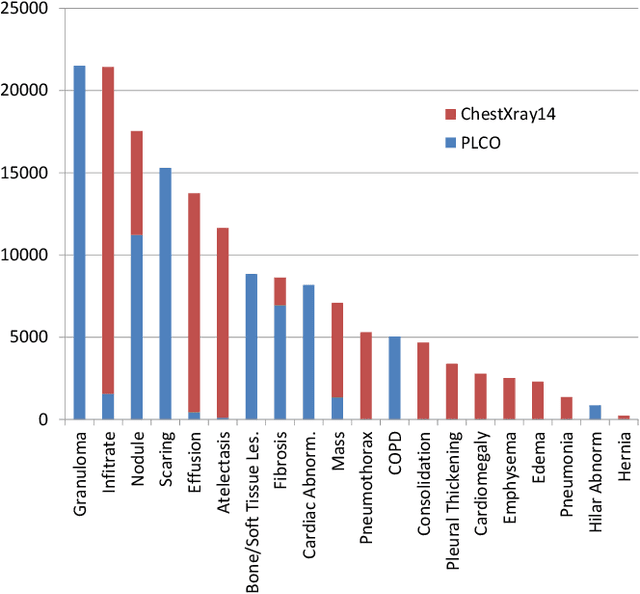
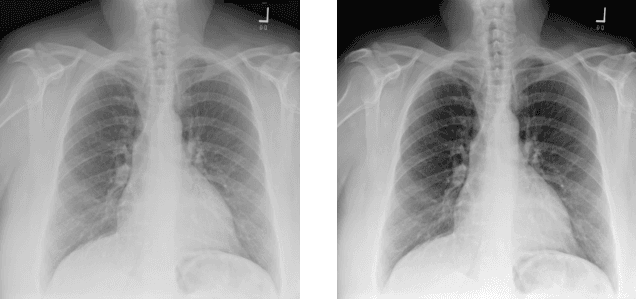
Abstract:Chest X-ray (CXR) is the most common X-ray examination performed in daily clinical practice for the diagnosis of various heart and lung abnormalities. The large amount of data to be read and reported, with 100+ studies per day for a single radiologist, poses a challenge in maintaining consistently high interpretation accuracy. In this work, we propose a method for the classification of different abnormalities based on CXR scans of the human body. The system is based on a novel multi-task deep learning architecture that in addition to the abnormality classification, supports the segmentation of the lungs and heart and classification of regions where the abnormality is located. We demonstrate that by training these tasks concurrently, one can increase the classification performance of the model. Experiments were performed on an extensive collection of 297,541 chest X-ray images from 86,876 patients, leading to a state-of-the-art performance level of 0.883 AUC on average for 12 different abnormalities. We also conducted a detailed performance analysis and compared the accuracy of our system with 3 board-certified radiologists. In this context, we highlight the high level of label noise inherent to this problem. On a reduced subset containing only cases with high confidence reference labels based on the consensus of the 3 radiologists, our system reached an average AUC of 0.945.
Class-Aware Adversarial Lung Nodule Synthesis in CT Images
Dec 28, 2018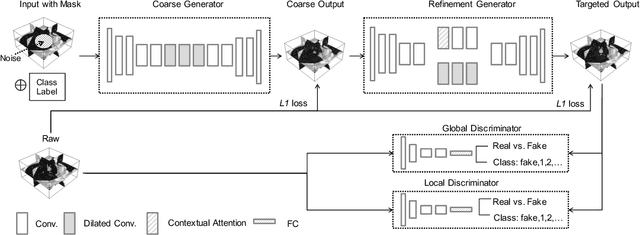
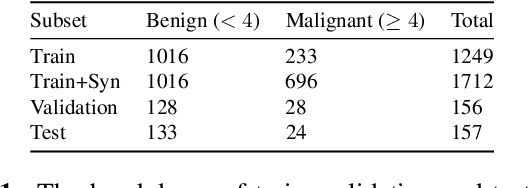

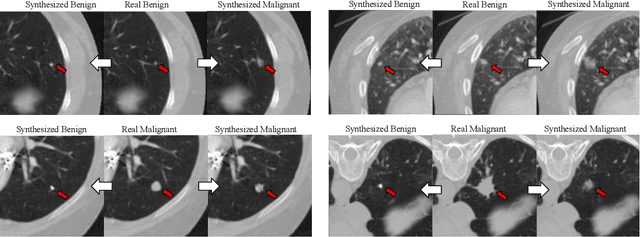
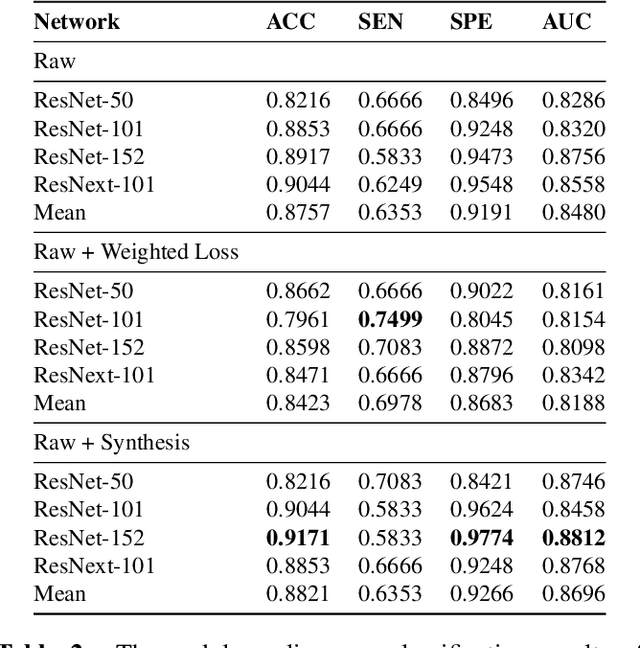
Abstract:Though large-scale datasets are essential for training deep learning systems, it is expensive to scale up the collection of medical imaging datasets. Synthesizing the objects of interests, such as lung nodules, in medical images based on the distribution of annotated datasets can be helpful for improving the supervised learning tasks, especially when the datasets are limited by size and class balance. In this paper, we propose the class-aware adversarial synthesis framework to synthesize lung nodules in CT images. The framework is built with a coarse-to-fine patch in-painter (generator) and two class-aware discriminators. By conditioning on the random latent variables and the target nodule labels, the trained networks are able to generate diverse nodules given the same context. By evaluating on the public LIDC-IDRI dataset, we demonstrate an example application of the proposed framework for improving the accuracy of the lung nodule malignancy estimation as a binary classification problem, which is important in the lung screening scenario. We show that combining the real image patches and the synthetic lung nodules in the training set can improve the mean AUC classification score across different network architectures by 2%.
Decompose to manipulate: Manipulable Object Synthesis in 3D Medical Images with Structured Image Decomposition
Dec 04, 2018
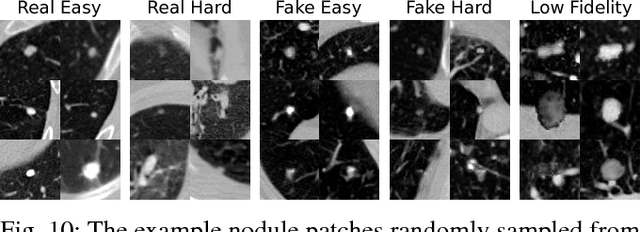

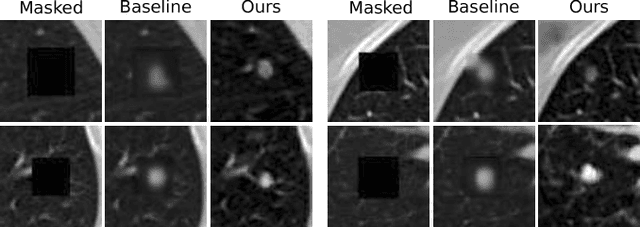
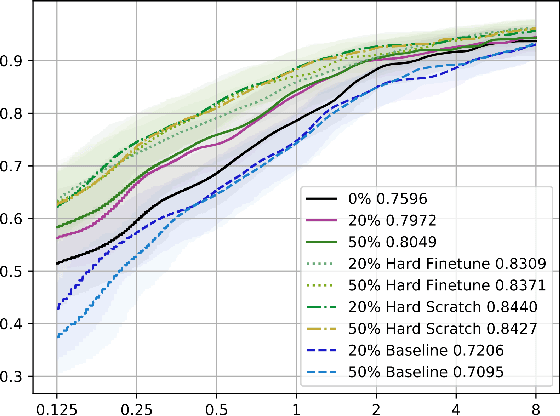
Abstract:The performance of medical image analysis systems is constrained by the quantity of high-quality image annotations. Such systems require data to be annotated by experts with years of training, especially when diagnostic decisions are involved. Such datasets are thus hard to scale up. In this context, it is hard for supervised learning systems to generalize to the cases that are rare in the training set but would be present in real-world clinical practices. We believe that the synthetic image samples generated by a system trained on the real data can be useful for improving the supervised learning tasks in the medical image analysis applications. Allowing the image synthesis to be manipulable could help synthetic images provide complementary information to the training data rather than simply duplicating the real-data manifold. In this paper, we propose a framework for synthesizing 3D objects, such as pulmonary nodules, in 3D medical images with manipulable properties. The manipulation is enabled by decomposing of the object of interests into its segmentation mask and a 1D vector containing the residual information. The synthetic object is refined and blended into the image context with two adversarial discriminators. We evaluate the proposed framework on lung nodules in 3D chest CT images and show that the proposed framework could generate realistic nodules with manipulable shapes, textures and locations, etc. By sampling from both the synthetic nodules and the real nodules from 2800 3D CT volumes during the classifier training, we show the synthetic patches could improve the overall nodule detection performance by average 8.44% competition performance metric (CPM) score.
Combo Loss: Handling Input and Output Imbalance in Multi-Organ Segmentation
Oct 22, 2018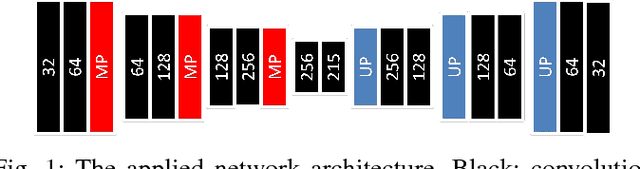
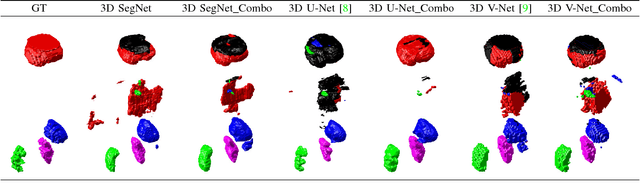
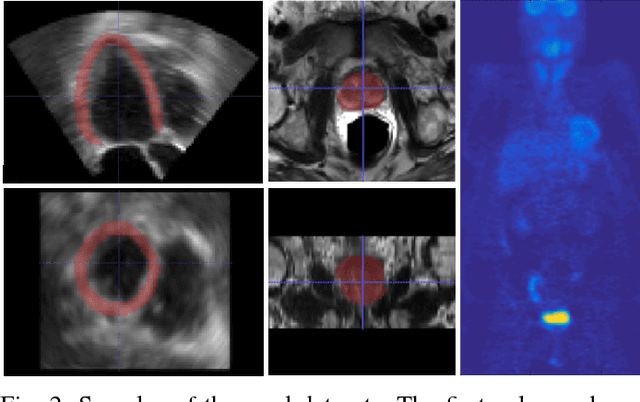
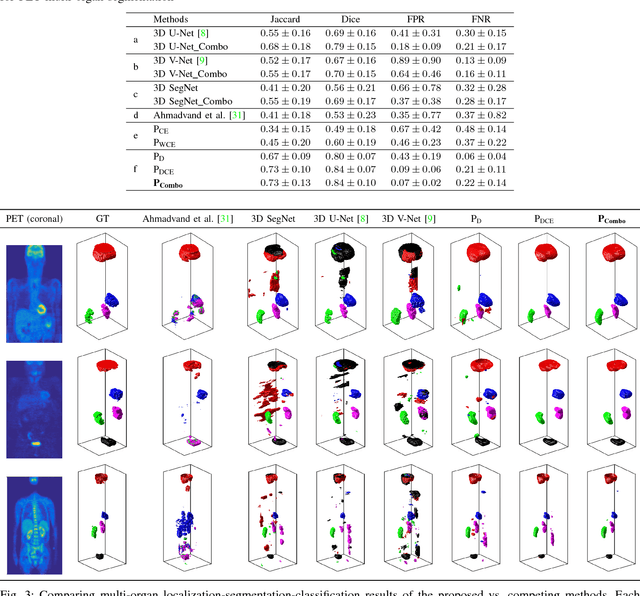

Abstract:Simultaneous segmentation of multiple organs from different medical imaging modalities is a crucial task as it can be utilized for computer-aided diagnosis, computer-assisted surgery, and therapy planning. Thanks to the recent advances in deep learning, several deep neural networks for medical image segmentation have been introduced successfully for this purpose. In this paper, we focus on learning a deep multi-organ segmentation network that labels voxels. In particular, we examine the critical choice of a loss function in order to handle the notorious imbalance problem that plagues both the input and output of a learning model. The input imbalance refers to the class-imbalance in the input training samples (i.e., small foreground objects embedded in an abundance of background voxels, as well as organs of varying sizes). The output imbalance refers to the imbalance between the false positives and false negatives of the inference model. In order to tackle both types of imbalance during training and inference, we introduce a new curriculum learning based loss function. Specifically, we leverage Dice similarity coefficient to deter model parameters from being held at bad local minima and at the same time gradually learn better model parameters by penalizing for false positives/negatives using a cross entropy term. We evaluated the proposed loss function on three datasets: whole body positron emission tomography (PET) scans with 5 target organs, magnetic resonance imaging (MRI) prostate scans, and ultrasound echocardigraphy images with a single target organ i.e., left ventricular. We show that a simple network architecture with the proposed integrative loss function can outperform state-of-the-art methods and results of the competing methods can be improved when our proposed loss is used.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge