Debdoot Sheet
UltraCompression: Framework for High Density Compression of Ultrasound Volumes using Physics Modeling Deep Neural Networks
Jan 17, 2019
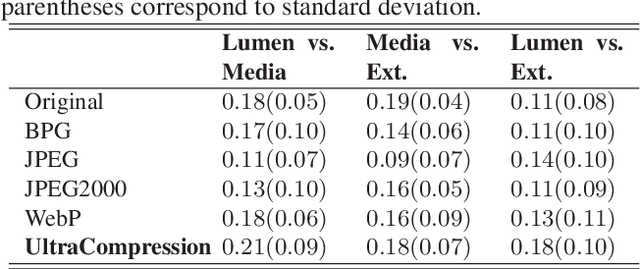
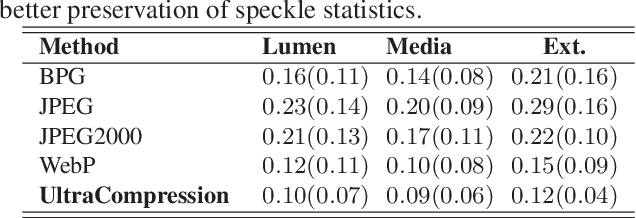
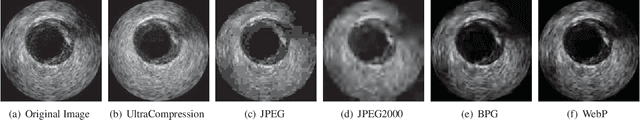
Abstract:Ultrasound image compression by preserving speckle-based key information is a challenging task. In this paper, we introduce an ultrasound image compression framework with the ability to retain realism of speckle appearance despite achieving very high-density compression factors. The compressor employs a tissue segmentation method, transmitting segments along with transducer frequency, number of samples and image size as essential information required for decompression. The decompressor is based on a convolutional network trained to generate patho-realistic ultrasound images which convey essential information pertinent to tissue pathology visible in the images. We demonstrate generalizability of the building blocks using two variants to build the compressor. We have evaluated the quality of decompressed images using distortion losses as well as perception loss and compared it with other off the shelf solutions. The proposed method achieves a compression ratio of $725:1$ while preserving the statistical distribution of speckles. This enables image segmentation on decompressed images to achieve dice score of $0.89 \pm 0.11$, which evidently is not so accurately achievable when images are compressed with current standards like JPEG, JPEG 2000, WebP and BPG. We envision this frame work to serve as a roadmap for speckle image compression standards.
Fully Convolutional Model for Variable Bit Length and Lossy High Density Compression of Mammograms
May 17, 2018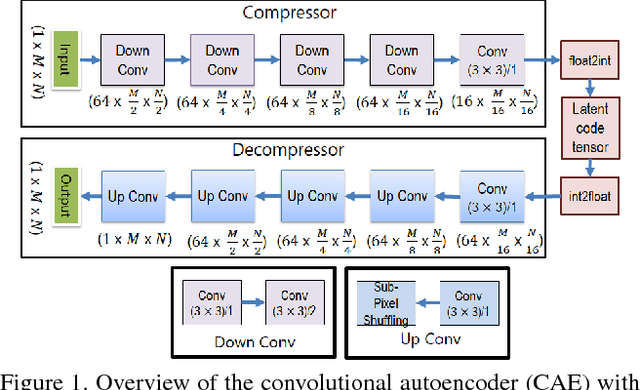
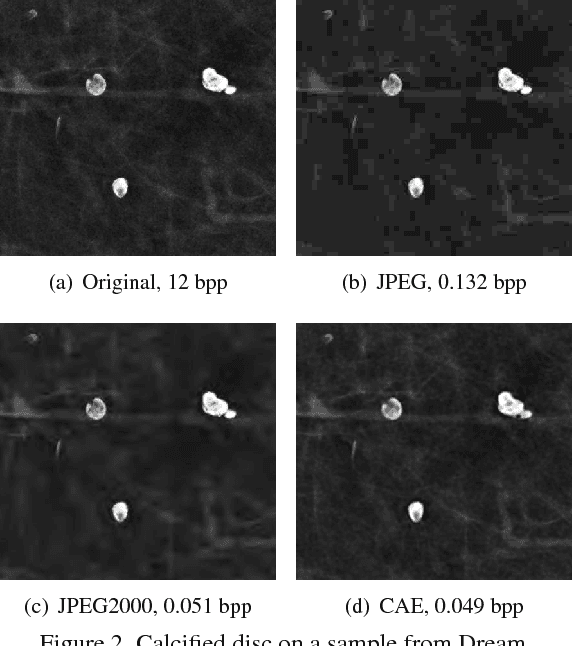
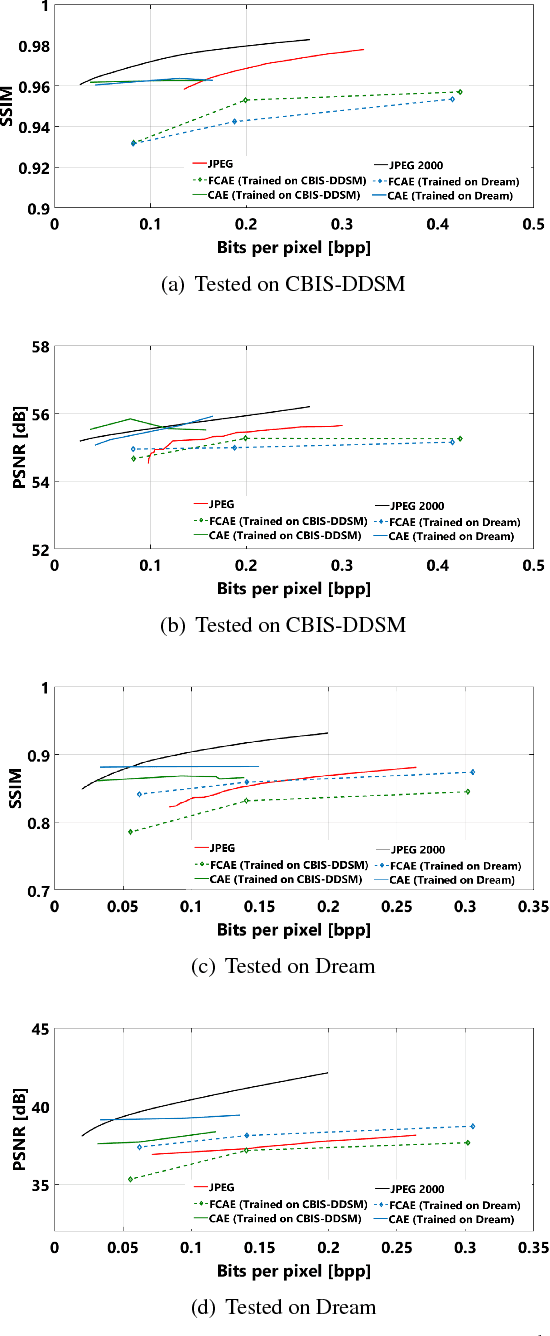
Abstract:Early works on medical image compression date to the 1980's with the impetus on deployment of teleradiology systems for high-resolution digital X-ray detectors. Commercially deployed systems during the period could compress 4,096 x 4,096 sized images at 12 bpp to 2 bpp using lossless arithmetic coding, and over the years JPEG and JPEG2000 were imbibed reaching upto 0.1 bpp. Inspired by the reprise of deep learning based compression for natural images over the last two years, we propose a fully convolutional autoencoder for diagnostically relevant feature preserving lossy compression. This is followed by leveraging arithmetic coding for encapsulating high redundancy of features for further high-density code packing leading to variable bit length. We demonstrate performance on two different publicly available digital mammography datasets using peak signal-to-noise ratio (pSNR), structural similarity (SSIM) index and domain adaptability tests between datasets. At high density compression factors of >300x (~0.04 bpp), our approach rivals JPEG and JPEG2000 as evaluated through a Radiologist's visual Turing test.
Simulating Patho-realistic Ultrasound Images using Deep Generative Networks with Adversarial Learning
Jan 08, 2018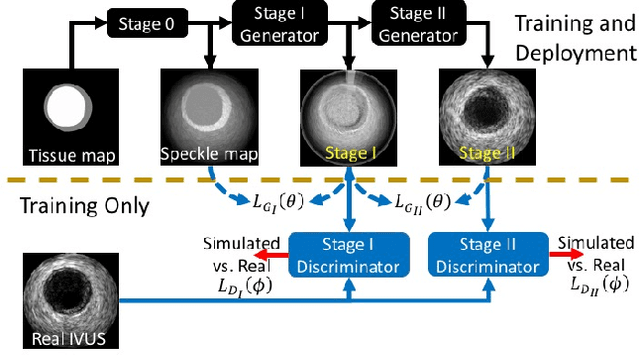
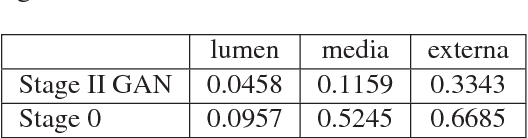
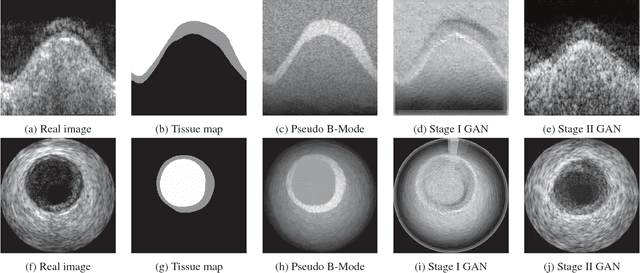

Abstract:Ultrasound imaging makes use of backscattering of waves during their interaction with scatterers present in biological tissues. Simulation of synthetic ultrasound images is a challenging problem on account of inability to accurately model various factors of which some include intra-/inter scanline interference, transducer to surface coupling, artifacts on transducer elements, inhomogeneous shadowing and nonlinear attenuation. Current approaches typically solve wave space equations making them computationally expensive and slow to operate. We propose a generative adversarial network (GAN) inspired approach for fast simulation of patho-realistic ultrasound images. We apply the framework to intravascular ultrasound (IVUS) simulation. A stage 0 simulation performed using pseudo B-mode ultrasound image simulator yields speckle mapping of a digitally defined phantom. The stage I GAN subsequently refines them to preserve tissue specific speckle intensities. The stage II GAN further refines them to generate high resolution images with patho-realistic speckle profiles. We evaluate patho-realism of simulated images with a visual Turing test indicating an equivocal confusion in discriminating simulated from real. We also quantify the shift in tissue specific intensity distributions of the real and simulated images to prove their similarity.
ReLayNet: Retinal Layer and Fluid Segmentation of Macular Optical Coherence Tomography using Fully Convolutional Network
Jul 07, 2017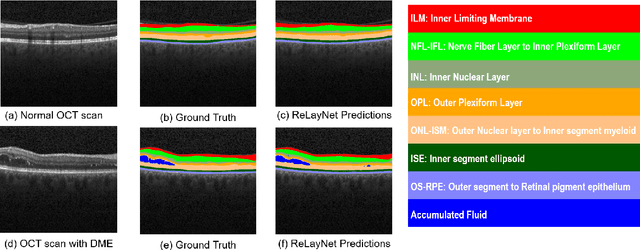
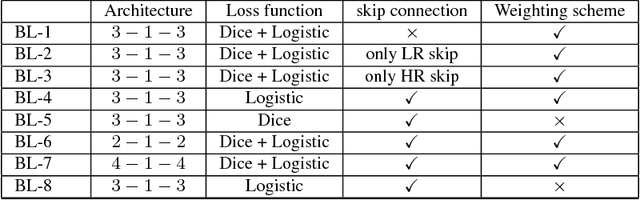

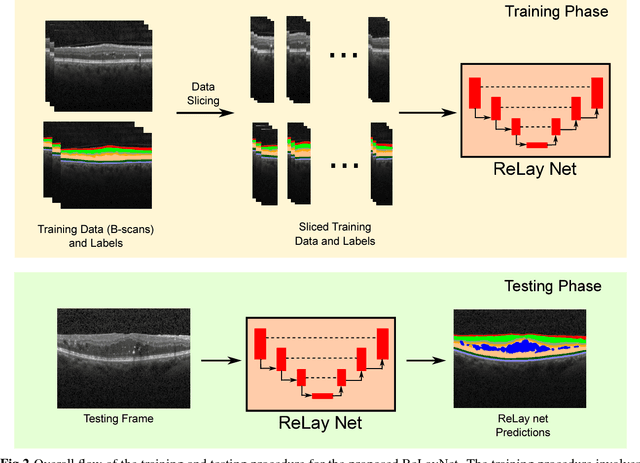
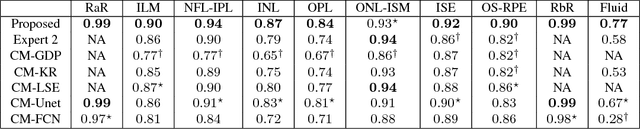
Abstract:Optical coherence tomography (OCT) is used for non-invasive diagnosis of diabetic macular edema assessing the retinal layers. In this paper, we propose a new fully convolutional deep architecture, termed ReLayNet, for end-to-end segmentation of retinal layers and fluid masses in eye OCT scans. ReLayNet uses a contracting path of convolutional blocks (encoders) to learn a hierarchy of contextual features, followed by an expansive path of convolutional blocks (decoders) for semantic segmentation. ReLayNet is trained to optimize a joint loss function comprising of weighted logistic regression and Dice overlap loss. The framework is validated on a publicly available benchmark dataset with comparisons against five state-of-the-art segmentation methods including two deep learning based approaches to substantiate its effectiveness.
Error Corrective Boosting for Learning Fully Convolutional Networks with Limited Data
Jul 02, 2017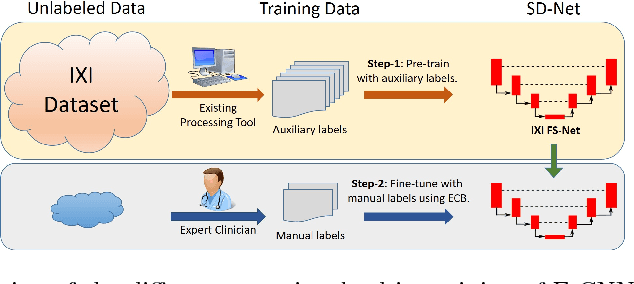
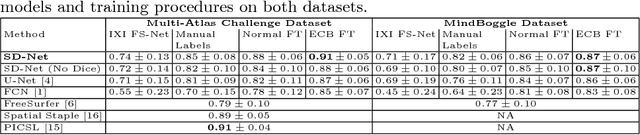
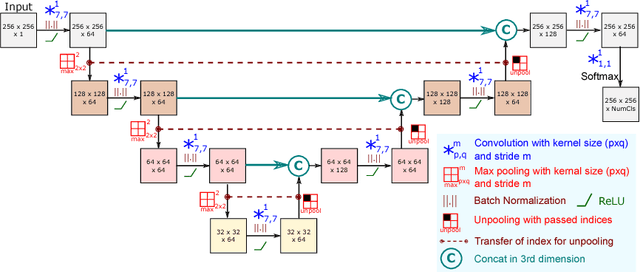
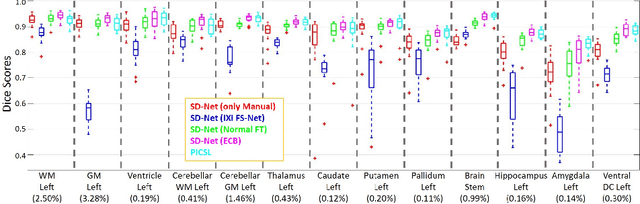
Abstract:Training deep fully convolutional neural networks (F-CNNs) for semantic image segmentation requires access to abundant labeled data. While large datasets of unlabeled image data are available in medical applications, access to manually labeled data is very limited. We propose to automatically create auxiliary labels on initially unlabeled data with existing tools and to use them for pre-training. For the subsequent fine-tuning of the network with manually labeled data, we introduce error corrective boosting (ECB), which emphasizes parameter updates on classes with lower accuracy. Furthermore, we introduce SkipDeconv-Net (SD-Net), a new F-CNN architecture for brain segmentation that combines skip connections with the unpooling strategy for upsampling. The SD-Net addresses challenges of severe class imbalance and errors along boundaries. With application to whole-brain MRI T1 scan segmentation, we generate auxiliary labels on a large dataset with FreeSurfer and fine-tune on two datasets with manual annotations. Our results show that the inclusion of auxiliary labels and ECB yields significant improvements. SD-Net segments a 3D scan in 7 secs in comparison to 30 hours for the closest multi-atlas segmentation method, while reaching similar performance. It also outperforms the latest state-of-the-art F-CNN models.
An Unsupervised Approach for Overlapping Cervical Cell Cytoplasm Segmentation
Feb 17, 2017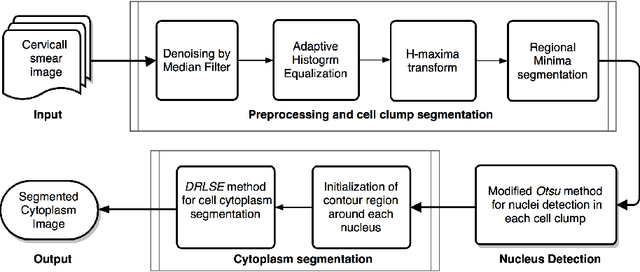
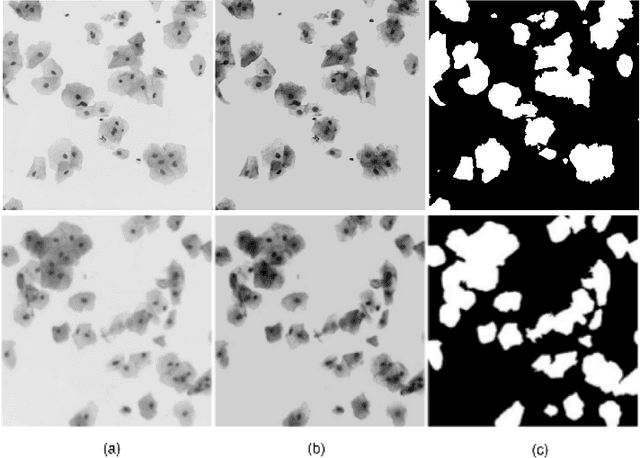
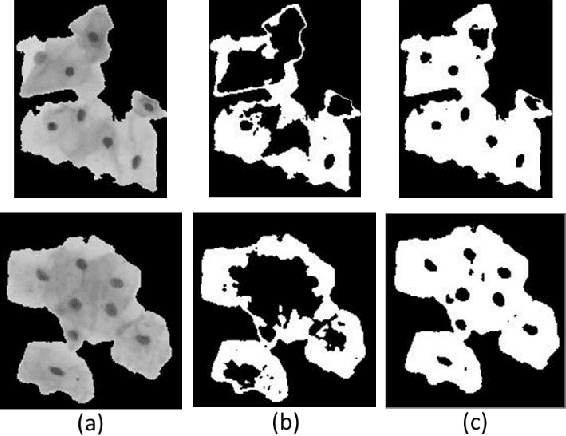


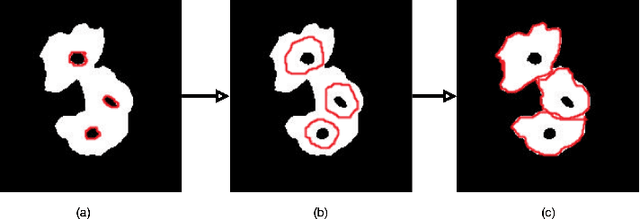
Abstract:The poor contrast and the overlapping of cervical cell cytoplasm are the major issues in the accurate segmentation of cervical cell cytoplasm. This paper presents an automated unsupervised cytoplasm segmentation approach which can effectively find the cytoplasm boundaries in overlapping cells. The proposed approach first segments the cell clumps from the cervical smear image and detects the nuclei in each cell clump. A modified Otsu method with prior class probability is proposed for accurate segmentation of nuclei from the cell clumps. Using distance regularized level set evolution, the contour around each nucleus is evolved until it reaches the cytoplasm boundaries. Promising results were obtained by experimenting on ISBI 2015 challenge dataset.
* 4 pages, 4 figures, Biomedical Engineering and Sciences (IECBES), 2016 IEEE EMBS Conference on. IEEE, 2016
Deep Neural Ensemble for Retinal Vessel Segmentation in Fundus Images towards Achieving Label-free Angiography
Sep 19, 2016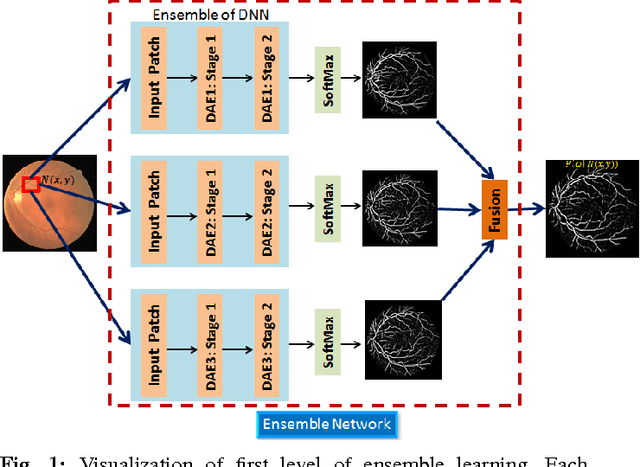
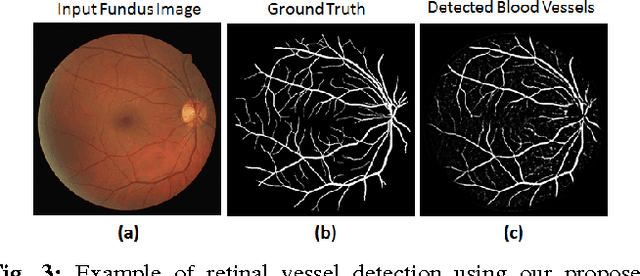
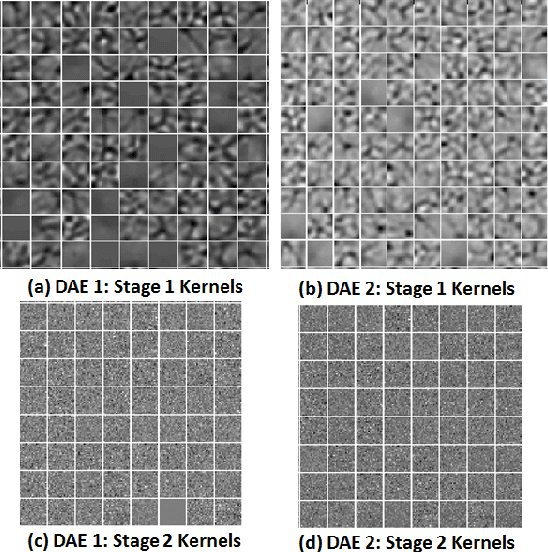
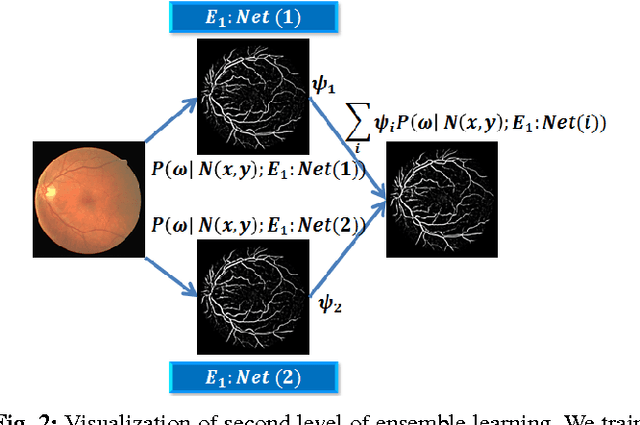


Abstract:Automated segmentation of retinal blood vessels in label-free fundus images entails a pivotal role in computed aided diagnosis of ophthalmic pathologies, viz., diabetic retinopathy, hypertensive disorders and cardiovascular diseases. The challenge remains active in medical image analysis research due to varied distribution of blood vessels, which manifest variations in their dimensions of physical appearance against a noisy background. In this paper we formulate the segmentation challenge as a classification task. Specifically, we employ unsupervised hierarchical feature learning using ensemble of two level of sparsely trained denoised stacked autoencoder. First level training with bootstrap samples ensures decoupling and second level ensemble formed by different network architectures ensures architectural revision. We show that ensemble training of auto-encoders fosters diversity in learning dictionary of visual kernels for vessel segmentation. SoftMax classifier is used for fine tuning each member auto-encoder and multiple strategies are explored for 2-level fusion of ensemble members. On DRIVE dataset, we achieve maximum average accuracy of 95.33\% with an impressively low standard deviation of 0.003 and Kappa agreement coefficient of 0.708 . Comparison with other major algorithms substantiates the high efficacy of our model.
DASA: Domain Adaptation in Stacked Autoencoders using Systematic Dropout
Mar 19, 2016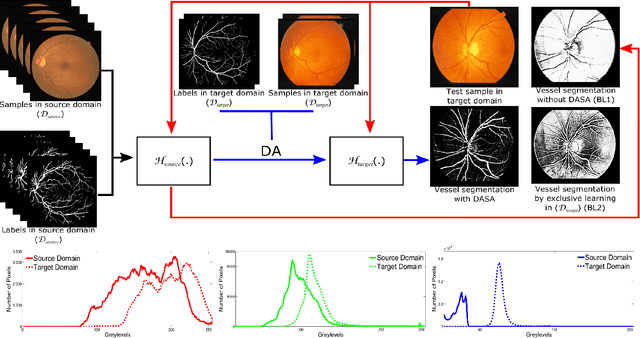
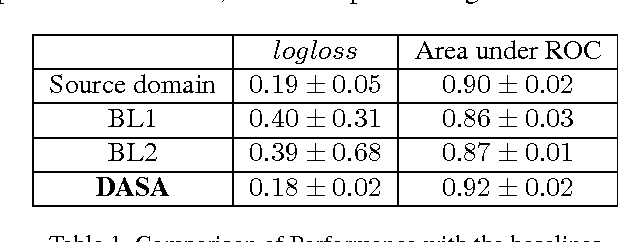
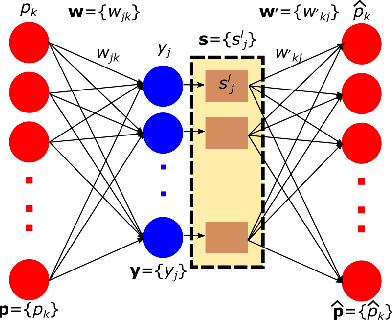
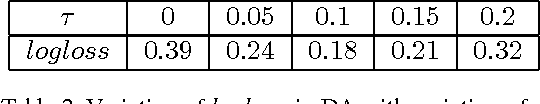
Abstract:Domain adaptation deals with adapting behaviour of machine learning based systems trained using samples in source domain to their deployment in target domain where the statistics of samples in both domains are dissimilar. The task of directly training or adapting a learner in the target domain is challenged by lack of abundant labeled samples. In this paper we propose a technique for domain adaptation in stacked autoencoder (SAE) based deep neural networks (DNN) performed in two stages: (i) unsupervised weight adaptation using systematic dropouts in mini-batch training, (ii) supervised fine-tuning with limited number of labeled samples in target domain. We experimentally evaluate performance in the problem of retinal vessel segmentation where the SAE-DNN is trained using large number of labeled samples in the source domain (DRIVE dataset) and adapted using less number of labeled samples in target domain (STARE dataset). The performance of SAE-DNN measured using $logloss$ in source domain is $0.19$, without and with adaptation are $0.40$ and $0.18$, and $0.39$ when trained exclusively with limited samples in target domain. The area under ROC curve is observed respectively as $0.90$, $0.86$, $0.92$ and $0.87$. The high efficiency of vessel segmentation with DASA strongly substantiates our claim.
Ensemble of Deep Convolutional Neural Networks for Learning to Detect Retinal Vessels in Fundus Images
Mar 15, 2016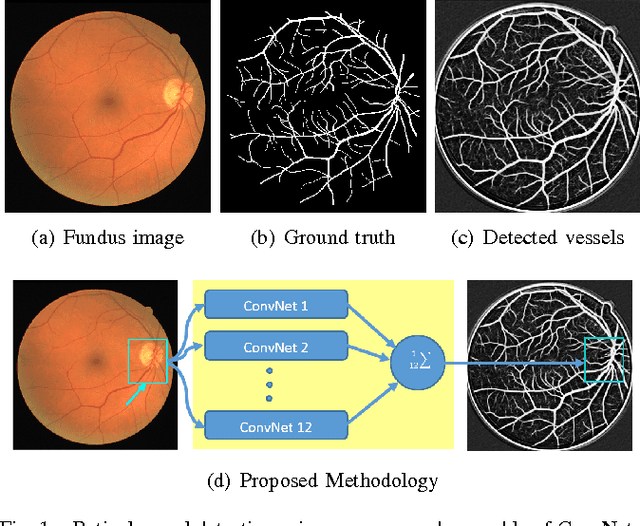
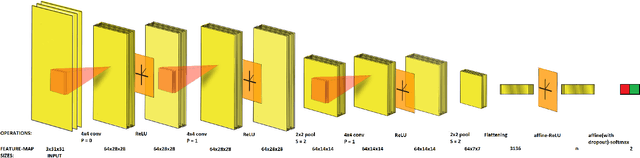
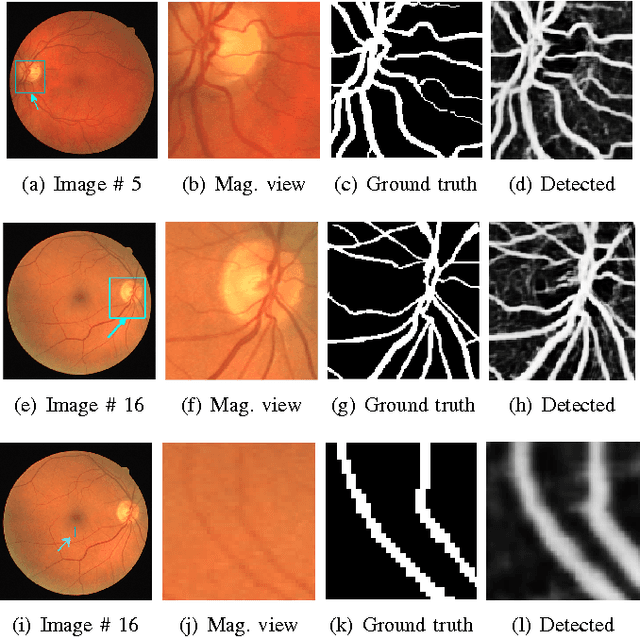
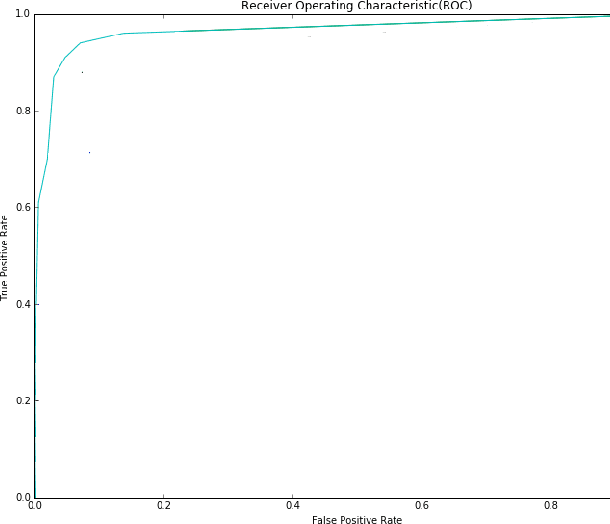
Abstract:Vision impairment due to pathological damage of the retina can largely be prevented through periodic screening using fundus color imaging. However the challenge with large scale screening is the inability to exhaustively detect fine blood vessels crucial to disease diagnosis. In this work we present a computational imaging framework using deep and ensemble learning for reliable detection of blood vessels in fundus color images. An ensemble of deep convolutional neural networks is trained to segment vessel and non-vessel areas of a color fundus image. During inference, the responses of the individual ConvNets of the ensemble are averaged to form the final segmentation. In experimental evaluation with the DRIVE database, we achieve the objective of vessel detection with maximum average accuracy of 94.7\% and area under ROC curve of 0.9283.
Unsupervised Segmentation of Overlapping Cervical Cell Cytoplasm
May 21, 2015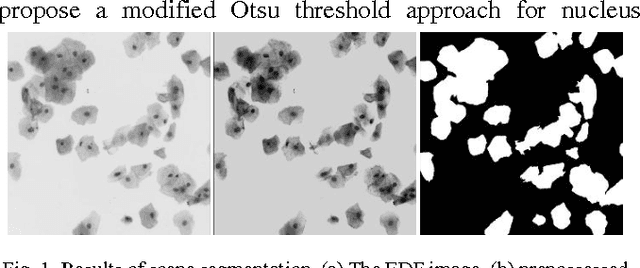
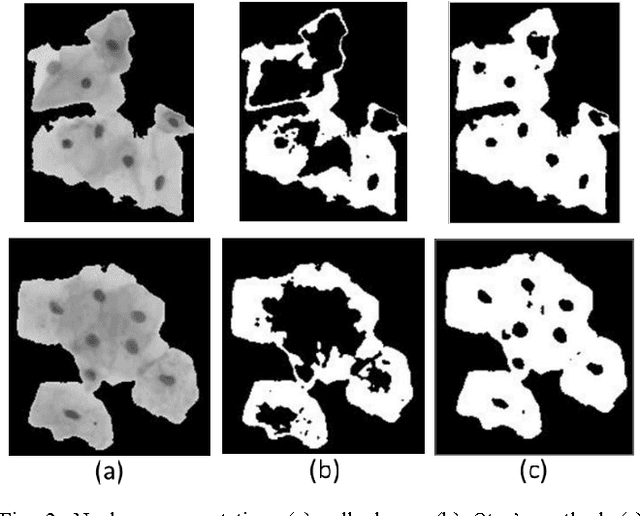

Abstract:Overlapping of cervical cells and poor contrast of cell cytoplasm are the major issues in accurate detection and segmentation of cervical cells. An unsupervised cell segmentation approach is presented here. Cell clump segmentation was carried out using the extended depth of field (EDF) image created from the images of different focal planes. A modified Otsu method with prior class weights is proposed for accurate segmentation of nuclei from the cell clumps. The cell cytoplasm was further segmented from cell clump depending upon the number of nucleus detected in that cell clump. Level set model was used for cytoplasm segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge