Swarnadip Chatterjee
Needle in a Haystack -- One-Class Representation Learning for Detecting Rare Malignant Cells in Computational Cytology
Apr 09, 2026Abstract:In computational cytology, detecting malignancy on whole-slide images is difficult because malignant cells are morphologically diverse yet vanishingly rare amid a vast background of normal cells. Accurate detection of these extremely rare malignant cells remains challenging due to large class imbalance and limited annotations. Conventional weakly supervised approaches, such as multiple instance learning (MIL), often fail to generalize at the instance level, especially when the fraction of malignant cells (witness rate) is exceedingly low. In this study, we explore the use of one-class representation learning techniques for detecting malignant cells in low-witness-rate scenarios. These methods are trained exclusively on slide-negative patches, without requiring any instance-level supervision. Specifically, we evaluate two OCC approaches, DSVDD and DROC, and compare them with FS-SIL, WS-SIL, and the recent ItS2CLR method. The one-class methods learn compact representations of normality and detect deviations at test time. Experiments on a publicly available bone marrow cytomorphology dataset (TCIA) and an in-house oral cancer cytology dataset show that DSVDD achieves state-of-the-art performance in instance-level abnormality ranking, particularly in ultra-low witness-rate regimes ($\leq 1\%$) and, in some cases, even outperforming fully supervised learning, which is typically not a practical option in whole-slide cytology due to the infeasibility of exhaustive instance-level annotations. DROC is also competitive under extreme rarity, benefiting from distribution-augmented contrastive learning. These findings highlight one-class representation learning as a robust and interpretable superior choice to MIL for malignant cell detection under extreme rarity.
SLAM-AGS: Slide-Label Aware Multi-Task Pretraining Using Adaptive Gradient Surgery in Computational Cytology
Nov 18, 2025Abstract:Computational cytology faces two major challenges: i) instance-level labels are unreliable and prohibitively costly to obtain, ii) witness rates are extremely low. We propose SLAM-AGS, a Slide-Label-Aware Multitask pretraining framework that jointly optimizes (i) a weakly supervised similarity objective on slide-negative patches and (ii) a self-supervised contrastive objective on slide-positive patches, yielding stronger performance on downstream tasks. To stabilize learning, we apply Adaptive Gradient Surgery to tackle conflicting task gradients and prevent model collapse. We integrate the pretrained encoder into an attention-based Multiple Instance Learning aggregator for bag-level prediction and attention-guided retrieval of the most abnormal instances in a bag. On a publicly available bone-marrow cytology dataset, with simulated witness rates from 10% down to 0.5%, SLAM-AGS improves bag-level F1-Score and Top 400 positive cell retrieval over other pretraining methods, with the largest gains at low witness rates, showing that resolving gradient interference enables stable pretraining and better performance on downstream tasks. To facilitate reproducibility, we share our complete implementation and evaluation framework as open source: https://github.com/Ace95/SLAM-AGS.
An Unsupervised Approach for Overlapping Cervical Cell Cytoplasm Segmentation
Feb 17, 2017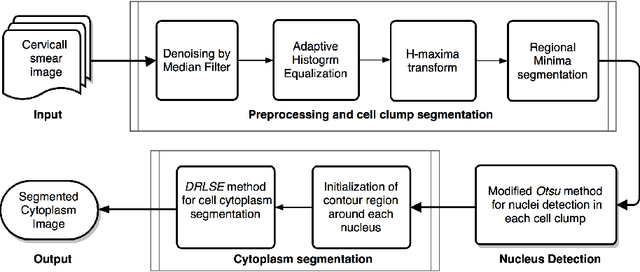
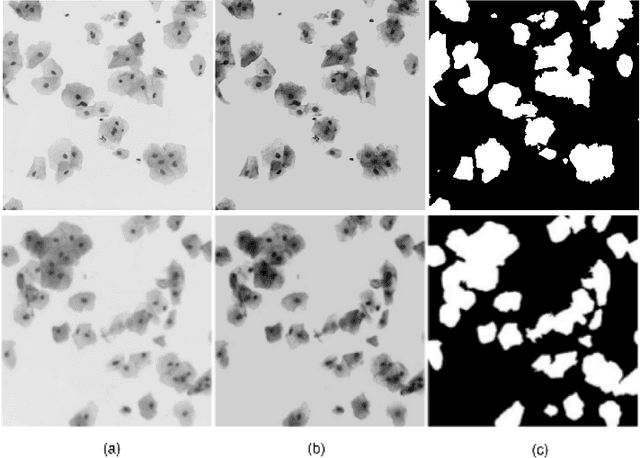
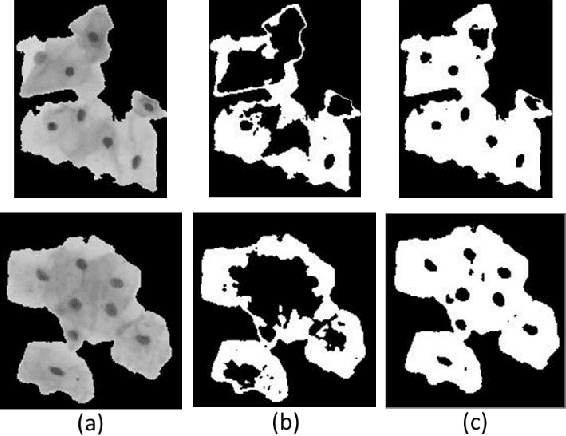


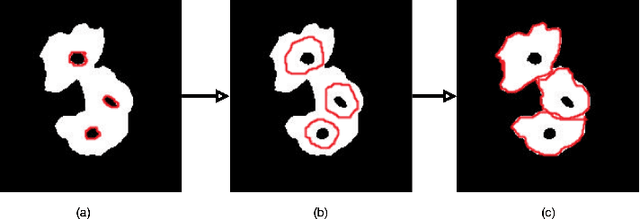
Abstract:The poor contrast and the overlapping of cervical cell cytoplasm are the major issues in the accurate segmentation of cervical cell cytoplasm. This paper presents an automated unsupervised cytoplasm segmentation approach which can effectively find the cytoplasm boundaries in overlapping cells. The proposed approach first segments the cell clumps from the cervical smear image and detects the nuclei in each cell clump. A modified Otsu method with prior class probability is proposed for accurate segmentation of nuclei from the cell clumps. Using distance regularized level set evolution, the contour around each nucleus is evolved until it reaches the cytoplasm boundaries. Promising results were obtained by experimenting on ISBI 2015 challenge dataset.
* 4 pages, 4 figures, Biomedical Engineering and Sciences (IECBES), 2016 IEEE EMBS Conference on. IEEE, 2016
Unsupervised Segmentation of Overlapping Cervical Cell Cytoplasm
May 21, 2015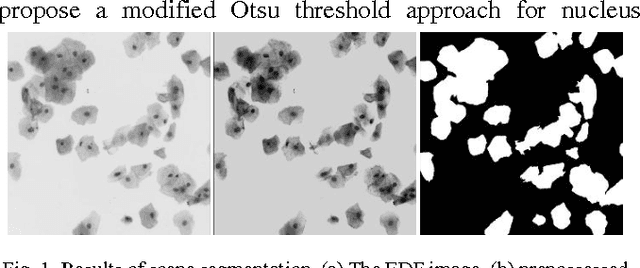
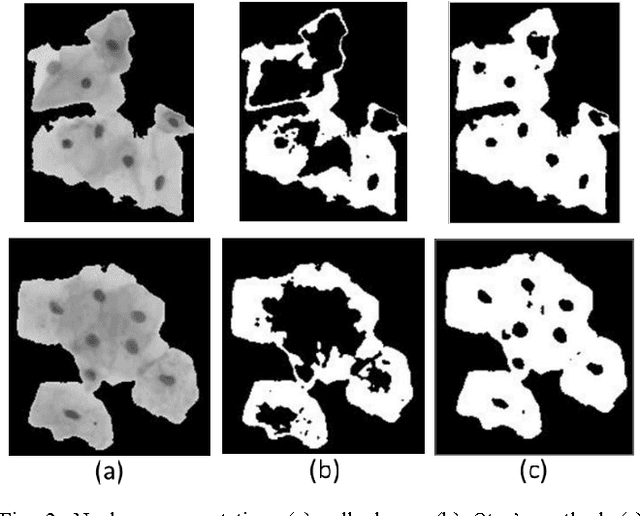

Abstract:Overlapping of cervical cells and poor contrast of cell cytoplasm are the major issues in accurate detection and segmentation of cervical cells. An unsupervised cell segmentation approach is presented here. Cell clump segmentation was carried out using the extended depth of field (EDF) image created from the images of different focal planes. A modified Otsu method with prior class weights is proposed for accurate segmentation of nuclei from the cell clumps. The cell cytoplasm was further segmented from cell clump depending upon the number of nucleus detected in that cell clump. Level set model was used for cytoplasm segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge