Davood Karimi
Atlas-Assisted Segment Anything Model for Fetal Brain MRI (FeTal-SAM)
Jan 22, 2026Abstract:This paper presents FeTal-SAM, a novel adaptation of the Segment Anything Model (SAM) tailored for fetal brain MRI segmentation. Traditional deep learning methods often require large annotated datasets for a fixed set of labels, making them inflexible when clinical or research needs change. By integrating atlas-based prompts and foundation-model principles, FeTal-SAM addresses two key limitations in fetal brain MRI segmentation: (1) the need to retrain models for varying label definitions, and (2) the lack of insight into whether segmentations are driven by genuine image contrast or by learned spatial priors. We leverage multi-atlas registration to generate spatially aligned label templates that serve as dense prompts, alongside a bounding-box prompt, for SAM's segmentation decoder. This strategy enables binary segmentation on a per-structure basis, which is subsequently fused to reconstruct the full 3D segmentation volumes. Evaluations on two datasets, the dHCP dataset and an in-house dataset demonstrate FeTal-SAM's robust performance across gestational ages. Notably, it achieves Dice scores comparable to state-of-the-art baselines which were trained for each dataset and label definition for well-contrasted structures like cortical plate and cerebellum, while maintaining the flexibility to segment any user-specified anatomy. Although slightly lower accuracy is observed for subtle, low-contrast structures (e.g., hippocampus, amygdala), our results highlight FeTal-SAM's potential to serve as a general-purpose segmentation model without exhaustive retraining. This method thus constitutes a promising step toward clinically adaptable fetal brain MRI analysis tools.
USFetal: Tools for Fetal Brain Ultrasound Compounding
Jan 11, 2026Abstract:Ultrasound offers a safe, cost-effective, and widely accessible technology for fetal brain imaging, making it especially suitable for routine clinical use. However, it suffers from view-dependent artifacts, operator variability, and a limited field of view, which make interpretation and quantitative evaluation challenging. Ultrasound compounding aims to overcome these limitations by integrating complementary information from multiple 3D acquisitions into a single, coherent volumetric representation. This work provides four main contributions: (1) We present the first systematic categorization of computational strategies for fetal brain ultrasound compounding, including both classical techniques and modern learning-based frameworks. (2) We implement and compare representative methods across four key categories - multi-scale, transformation-based, variational, and deep learning approaches - emphasizing their core principles and practical advantages. (3) Motivated by the lack of full-view, artifact-free ground truth required for supervised learning, we focus on unsupervised and self-supervised strategies and introduce two new deep learning based approaches: a self-supervised compounding framework and an adaptation of unsupervised deep plug-and-play priors for compounding. (4) We conduct a comprehensive evaluation on ten multi-view fetal brain ultrasound datasets, using both expert radiologist scoring and standard quantitative image-quality metrics. We also release the USFetal Compounding Toolbox, publicly available to support benchmarking and future research. Keywords: Ultrasound compounding, fetal brain, deep learning, self-supervised, unsupervised.
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025


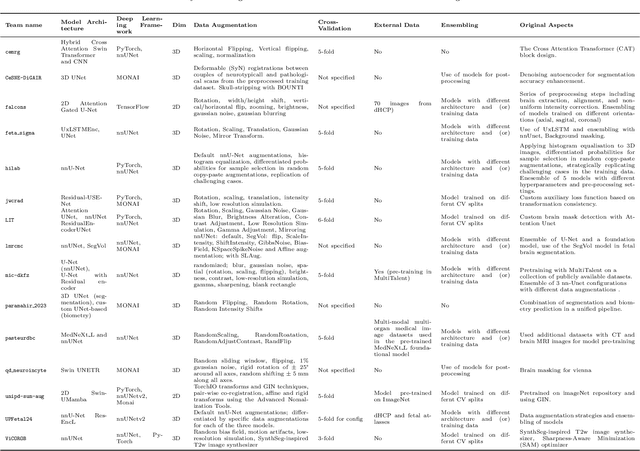
Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
Equivariant Spherical CNNs for Accurate Fiber Orientation Distribution Estimation in Neonatal Diffusion MRI with Reduced Acquisition Time
Apr 02, 2025Abstract:Early and accurate assessment of brain microstructure using diffusion Magnetic Resonance Imaging (dMRI) is crucial for identifying neurodevelopmental disorders in neonates, but remains challenging due to low signal-to-noise ratio (SNR), motion artifacts, and ongoing myelination. In this study, we propose a rotationally equivariant Spherical Convolutional Neural Network (sCNN) framework tailored for neonatal dMRI. We predict the Fiber Orientation Distribution (FOD) from multi-shell dMRI signals acquired with a reduced set of gradient directions (30% of the full protocol), enabling faster and more cost-effective acquisitions. We train and evaluate the performance of our sCNN using real data from 43 neonatal dMRI datasets provided by the Developing Human Connectome Project (dHCP). Our results demonstrate that the sCNN achieves significantly lower mean squared error (MSE) and higher angular correlation coefficient (ACC) compared to a Multi-Layer Perceptron (MLP) baseline, indicating improved accuracy in FOD estimation. Furthermore, tractography results based on the sCNN-predicted FODs show improved anatomical plausibility, coverage, and coherence compared to those from the MLP. These findings highlight that sCNNs, with their inherent rotational equivariance, offer a promising approach for accurate and clinically efficient dMRI analysis, paving the way for improved diagnostic capabilities and characterization of early brain development.
FetDTIAlign: A Deep Learning Framework for Affine and Deformable Registration of Fetal Brain dMRI
Feb 03, 2025Abstract:Diffusion MRI (dMRI) provides unique insights into fetal brain microstructure in utero. Longitudinal and cross-sectional fetal dMRI studies can reveal crucial neurodevelopmental changes but require precise spatial alignment across scans and subjects. This is challenging due to low data quality, rapid brain development, and limited anatomical landmarks. Existing registration methods, designed for high-quality adult data, struggle with these complexities. To address this, we introduce FetDTIAlign, a deep learning approach for fetal brain dMRI registration, enabling accurate affine and deformable alignment. FetDTIAlign features a dual-encoder architecture and iterative feature-based inference, reducing the impact of noise and low resolution. It optimizes network configurations and domain-specific features at each registration stage, enhancing both robustness and accuracy. We validated FetDTIAlign on data from 23 to 36 weeks gestation, covering 60 white matter tracts. It consistently outperformed two classical optimization-based methods and a deep learning pipeline, achieving superior anatomical correspondence. Further validation on external data from the Developing Human Connectome Project confirmed its generalizability across acquisition protocols. Our results demonstrate the feasibility of deep learning for fetal brain dMRI registration, providing a more accurate and reliable alternative to classical techniques. By enabling precise cross-subject and tract-specific analyses, FetDTIAlign supports new discoveries in early brain development.
Ground-truth effects in learning-based fiber orientation distribution estimation in neonatal brains
Sep 02, 2024



Abstract:Diffusion Magnetic Resonance Imaging (dMRI) is a non-invasive method for depicting brain microstructure in vivo. Fiber orientation distributions (FODs) are mathematical representations extensively used to map white matter fiber configurations. Recently, FOD estimation with deep neural networks has seen growing success, in particular, those of neonates estimated with fewer diffusion measurements. These methods are mostly trained on target FODs reconstructed with multi-shell multi-tissue constrained spherical deconvolution (MSMT-CSD), which might not be the ideal ground truth for developing brains. Here, we investigate this hypothesis by training a state-of-the-art model based on the U-Net architecture on both MSMT-CSD and single-shell three-tissue constrained spherical deconvolution (SS3T-CSD). Our results suggest that SS3T-CSD might be more suited for neonatal brains, given that the ratio between single and multiple fiber-estimated voxels with SS3T-CSD is more realistic compared to MSMT-CSD. Additionally, increasing the number of input gradient directions significantly improves performance with SS3T-CSD over MSMT-CSD. Finally, in an age domain-shift setting, SS3T-CSD maintains robust performance across age groups, indicating its potential for more accurate neonatal brain imaging.
Streamline tractography of the fetal brain in utero with machine learning
Aug 26, 2024



Abstract:Diffusion-weighted magnetic resonance imaging (dMRI) is the only non-invasive tool for studying white matter tracts and structural connectivity of the brain. These assessments rely heavily on tractography techniques, which reconstruct virtual streamlines representing white matter fibers. Much effort has been devoted to improving tractography methodology for adult brains, while tractography of the fetal brain has been largely neglected. Fetal tractography faces unique difficulties due to low dMRI signal quality, immature and rapidly developing brain structures, and paucity of reference data. This work presents the first machine learning model for fetal tractography. The model input consists of five sources of information: (1) Fiber orientation, inferred from a diffusion tensor fit to the dMRI signal; (2) Directions of recent propagation steps; (3) Global spatial information, encoded as distances to keypoints in the brain cortex; (4) Tissue segmentation information; and (5) Prior information about the expected local fiber orientations supplied with an atlas. In order to mitigate the local tensor estimation error, a large spatial context around the current point in the diffusion tensor image is encoded using convolutional and attention neural network modules. Moreover, the diffusion tensor information at a hypothetical next point is included in the model input. Filtering rules based on anatomically constrained tractography are applied to prune implausible streamlines. We trained the model on manually-refined whole-brain fetal tractograms and validated the trained model on an independent set of 11 test scans with gestational ages between 23 and 36 weeks. Results show that our proposed method achieves superior performance across all evaluated tracts. The new method can significantly advance the capabilities of dMRI for studying normal and abnormal brain development in utero.
HAITCH: A Framework for Distortion and Motion Correction in Fetal Multi-Shell Diffusion-Weighted MRI
Jun 28, 2024



Abstract:Diffusion magnetic resonance imaging (dMRI) is pivotal for probing the microstructure of the rapidly-developing fetal brain. However, fetal motion during scans and its interaction with magnetic field inhomogeneities result in artifacts and data scattering across spatial and angular domains. The effects of those artifacts are more pronounced in high-angular resolution fetal dMRI, where signal-to-noise ratio is very low. Those effects lead to biased estimates and compromise the consistency and reliability of dMRI analysis. This work presents HAITCH, the first and the only publicly available tool to correct and reconstruct multi-shell high-angular resolution fetal dMRI data. HAITCH offers several technical advances that include a blip-reversed dual-echo acquisition for dynamic distortion correction, advanced motion correction for model-free and robust reconstruction, optimized multi-shell design for enhanced information capture and increased tolerance to motion, and outlier detection for improved reconstruction fidelity. The framework is open-source, flexible, and can be used to process any type of fetal dMRI data including single-echo or single-shell acquisitions, but is most effective when used with multi-shell multi-echo fetal dMRI data that cannot be processed with any of the existing tools. Validation experiments on real fetal dMRI scans demonstrate significant improvements and accurate correction across diverse fetal ages and motion levels. HAITCH successfully removes artifacts and reconstructs high-fidelity fetal dMRI data suitable for advanced diffusion modeling, including fiber orientation distribution function estimation. These advancements pave the way for more reliable analysis of the fetal brain microstructure and tractography under challenging imaging conditions.
Anatomically Constrained Tractography of the Fetal Brain
Mar 04, 2024Abstract:Diffusion-weighted Magnetic Resonance Imaging (dMRI) is increasingly used to study the fetal brain in utero. An important computation enabled by dMRI is streamline tractography, which has unique applications such as tract-specific analysis of the brain white matter and structural connectivity assessment. However, due to the low fetal dMRI data quality and the challenging nature of tractography, existing methods tend to produce highly inaccurate results. They generate many false streamlines while failing to reconstruct streamlines that constitute the major white matter tracts. In this paper, we advocate for anatomically constrained tractography based on an accurate segmentation of the fetal brain tissue directly in the dMRI space. We develop a deep learning method to compute the segmentation automatically. Experiments on independent test data show that this method can accurately segment the fetal brain tissue and drastically improve tractography results. It enables the reconstruction of highly curved tracts such as optic radiations. Importantly, our method infers the tissue segmentation and streamline propagation direction from a diffusion tensor fit to the dMRI data, making it applicable to routine fetal dMRI scans. The proposed method can lead to significant improvements in the accuracy and reproducibility of quantitative assessment of the fetal brain with dMRI.
Cross-Age and Cross-Site Domain Shift Impacts on Deep Learning-Based White Matter Fiber Estimation in Newborn and Baby Brains
Dec 22, 2023Abstract:Deep learning models have shown great promise in estimating tissue microstructure from limited diffusion magnetic resonance imaging data. However, these models face domain shift challenges when test and train data are from different scanners and protocols, or when the models are applied to data with inherent variations such as the developing brains of infants and children scanned at various ages. Several techniques have been proposed to address some of these challenges, such as data harmonization or domain adaptation in the adult brain. However, those techniques remain unexplored for the estimation of fiber orientation distribution functions in the rapidly developing brains of infants. In this work, we extensively investigate the age effect and domain shift within and across two different cohorts of 201 newborns and 165 babies using the Method of Moments and fine-tuning strategies. Our results show that reduced variations in the microstructural development of babies in comparison to newborns directly impact the deep learning models' cross-age performance. We also demonstrate that a small number of target domain samples can significantly mitigate domain shift problems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge