David Sontag
Leveraging Time Irreversibility with Order-Contrastive Pre-training
Nov 04, 2021
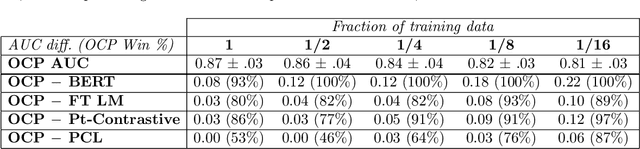
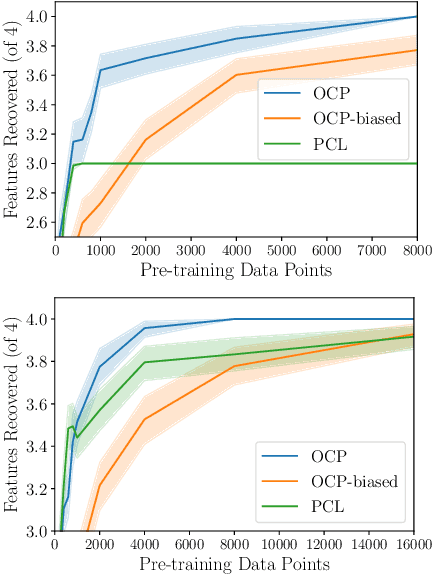
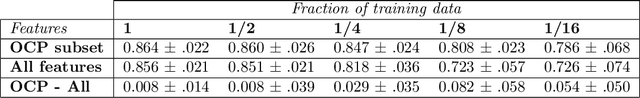
Abstract:Label-scarce, high-dimensional domains such as healthcare present a challenge for modern machine learning techniques. To overcome the difficulties posed by a lack of labeled data, we explore an "order-contrastive" method for self-supervised pre-training on longitudinal data. We sample pairs of time segments, switch the order for half of them, and train a model to predict whether a given pair is in the correct order. Intuitively, the ordering task allows the model to attend to the least time-reversible features (for example, features that indicate progression of a chronic disease). The same features are often useful for downstream tasks of interest. To quantify this, we study a simple theoretical setting where we prove a finite-sample guarantee for the downstream error of a representation learned with order-contrastive pre-training. Empirically, in synthetic and longitudinal healthcare settings, we demonstrate the effectiveness of order-contrastive pre-training in the small-data regime over supervised learning and other self-supervised pre-training baselines. Our results indicate that pre-training methods designed for particular classes of distributions and downstream tasks can improve the performance of self-supervised learning.
Using Time-Series Privileged Information for Provably Efficient Learning of Prediction Models
Oct 28, 2021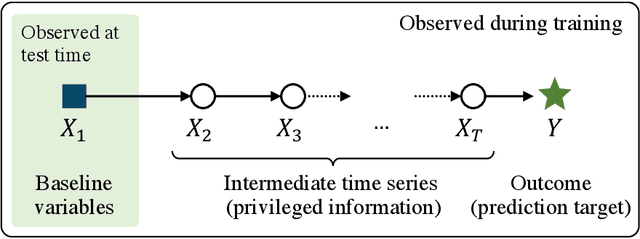
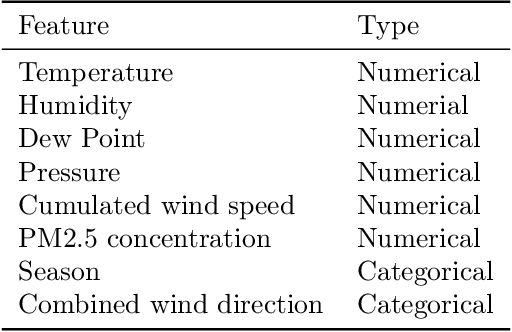
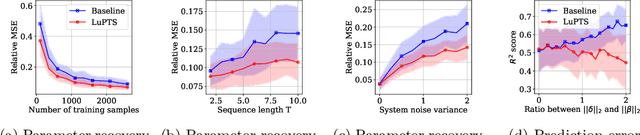
Abstract:We study prediction of future outcomes with supervised models that use privileged information during learning. The privileged information comprises samples of time series observed between the baseline time of prediction and the future outcome; this information is only available at training time which differs from the traditional supervised learning. Our question is when using this privileged data leads to more sample-efficient learning of models that use only baseline data for predictions at test time. We give an algorithm for this setting and prove that when the time series are drawn from a non-stationary Gaussian-linear dynamical system of fixed horizon, learning with privileged information is more efficient than learning without it. On synthetic data, we test the limits of our algorithm and theory, both when our assumptions hold and when they are violated. On three diverse real-world datasets, we show that our approach is generally preferable to classical learning, particularly when data is scarce. Finally, we relate our estimator to a distillation approach both theoretically and empirically.
Finding Regions of Heterogeneity in Decision-Making via Expected Conditional Covariance
Oct 27, 2021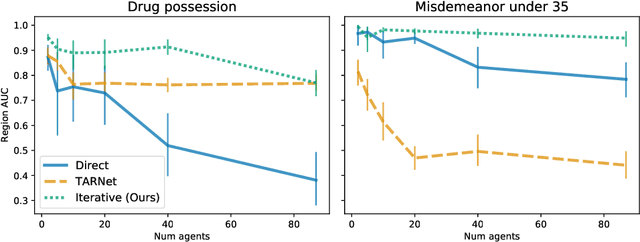
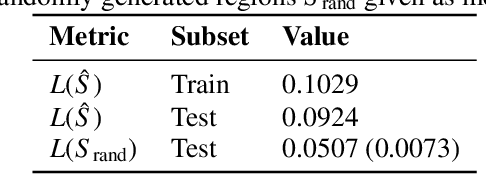
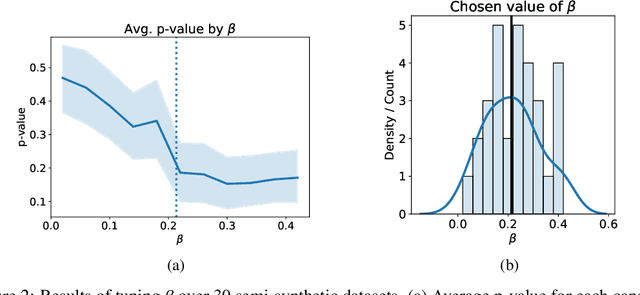

Abstract:Individuals often make different decisions when faced with the same context, due to personal preferences and background. For instance, judges may vary in their leniency towards certain drug-related offenses, and doctors may vary in their preference for how to start treatment for certain types of patients. With these examples in mind, we present an algorithm for identifying types of contexts (e.g., types of cases or patients) with high inter-decision-maker disagreement. We formalize this as a causal inference problem, seeking a region where the assignment of decision-maker has a large causal effect on the decision. Our algorithm finds such a region by maximizing an empirical objective, and we give a generalization bound for its performance. In a semi-synthetic experiment, we show that our algorithm recovers the correct region of heterogeneity accurately compared to baselines. Finally, we apply our algorithm to real-world healthcare datasets, recovering variation that aligns with existing clinical knowledge.
CLIP: A Dataset for Extracting Action Items for Physicians from Hospital Discharge Notes
Jun 04, 2021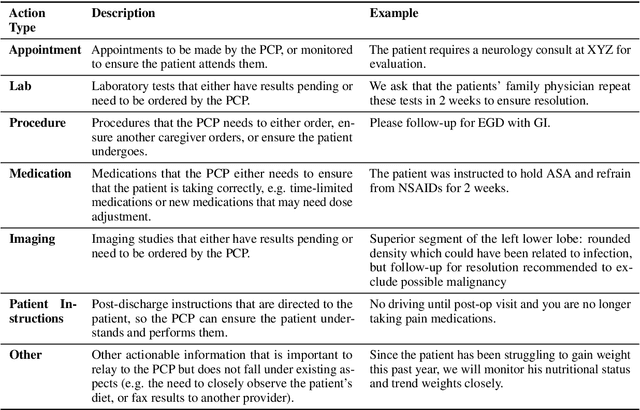
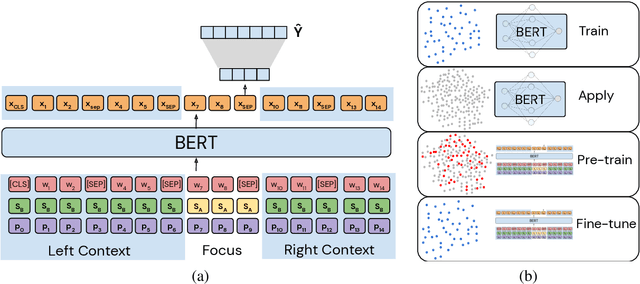

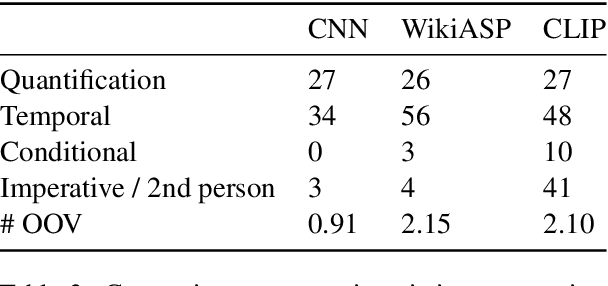
Abstract:Continuity of care is crucial to ensuring positive health outcomes for patients discharged from an inpatient hospital setting, and improved information sharing can help. To share information, caregivers write discharge notes containing action items to share with patients and their future caregivers, but these action items are easily lost due to the lengthiness of the documents. In this work, we describe our creation of a dataset of clinical action items annotated over MIMIC-III, the largest publicly available dataset of real clinical notes. This dataset, which we call CLIP, is annotated by physicians and covers 718 documents representing 100K sentences. We describe the task of extracting the action items from these documents as multi-aspect extractive summarization, with each aspect representing a type of action to be taken. We evaluate several machine learning models on this task, and show that the best models exploit in-domain language model pre-training on 59K unannotated documents, and incorporate context from neighboring sentences. We also propose an approach to pre-training data selection that allows us to explore the trade-off between size and domain-specificity of pre-training datasets for this task.
Regularizing towards Causal Invariance: Linear Models with Proxies
Mar 03, 2021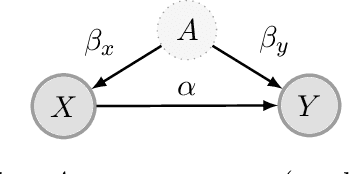
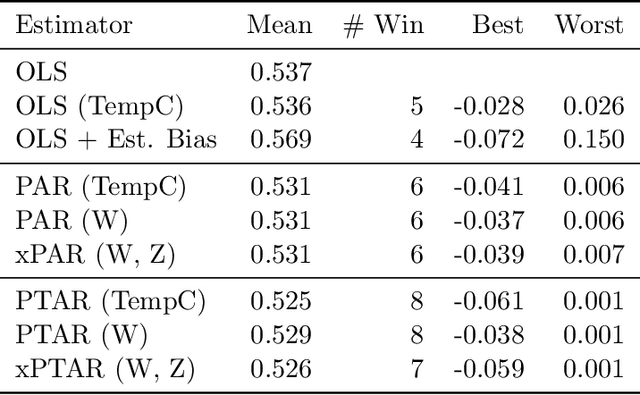
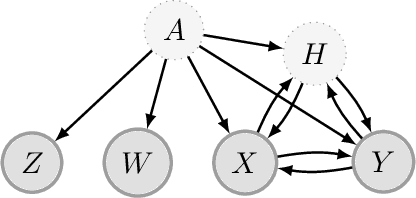
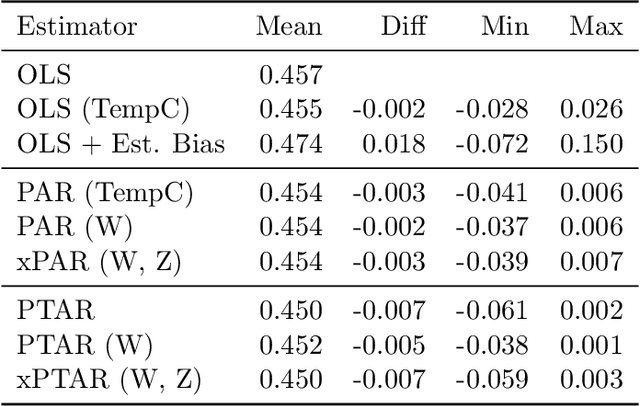
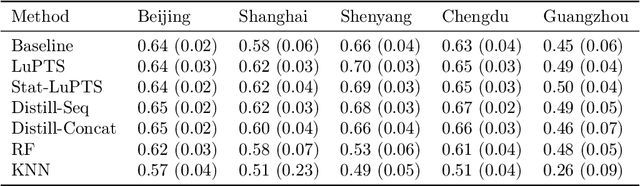
Abstract:We propose a method for learning linear models whose predictive performance is robust to causal interventions on unobserved variables, when noisy proxies of those variables are available. Our approach takes the form of a regularization term that trades off between in-distribution performance and robustness to interventions. Under the assumption of a linear structural causal model, we show that a single proxy can be used to create estimators that are prediction optimal under interventions of bounded strength. This strength depends on the magnitude of the measurement noise in the proxy, which is, in general, not identifiable. In the case of two proxy variables, we propose a modified estimator that is prediction optimal under interventions up to a known strength. We further show how to extend these estimators to scenarios where additional information about the "test time" intervention is available during training. We evaluate our theoretical findings in synthetic experiments and using real data of hourly pollution levels across several cities in China.
Beyond Perturbation Stability: LP Recovery Guarantees for MAP Inference on Noisy Stable Instances
Feb 26, 2021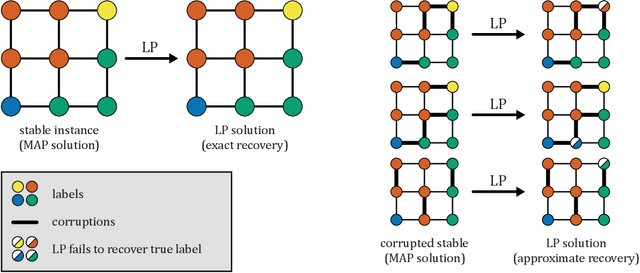

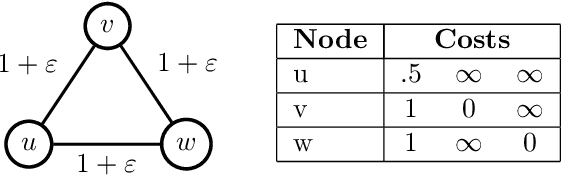

Abstract:Several works have shown that perturbation stable instances of the MAP inference problem in Potts models can be solved exactly using a natural linear programming (LP) relaxation. However, most of these works give few (or no) guarantees for the LP solutions on instances that do not satisfy the relatively strict perturbation stability definitions. In this work, we go beyond these stability results by showing that the LP approximately recovers the MAP solution of a stable instance even after the instance is corrupted by noise. This "noisy stable" model realistically fits with practical MAP inference problems: we design an algorithm for finding "close" stable instances, and show that several real-world instances from computer vision have nearby instances that are perturbation stable. These results suggest a new theoretical explanation for the excellent performance of this LP relaxation in practice.
Neural Pharmacodynamic State Space Modeling
Feb 22, 2021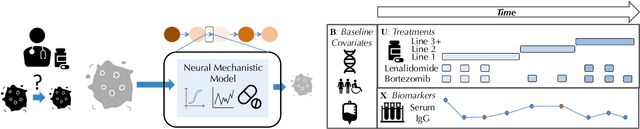
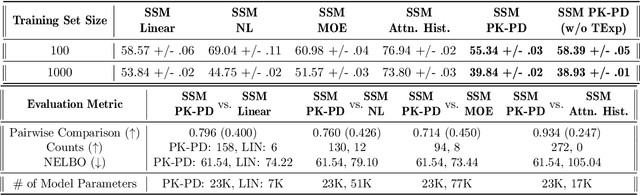
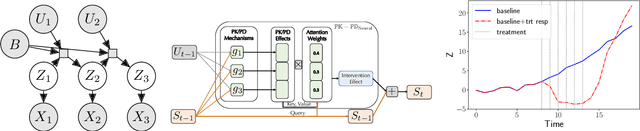
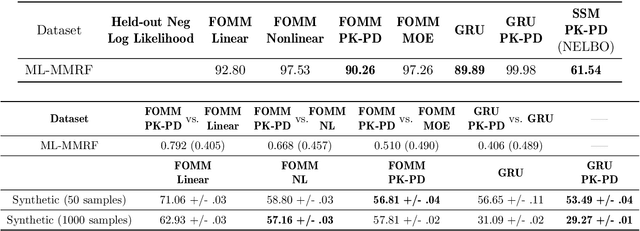
Abstract:Modeling the time-series of high-dimensional, longitudinal data is important for predicting patient disease progression. However, existing neural network based approaches that learn representations of patient state, while very flexible, are susceptible to overfitting. We propose a deep generative model that makes use of a novel attention-based neural architecture inspired by the physics of how treatments affect disease state. The result is a scalable and accurate model of high-dimensional patient biomarkers as they vary over time. Our proposed model yields significant improvements in generalization and, on real-world clinical data, provides interpretable insights into the dynamics of cancer progression.
Clustering Left-Censored Multivariate Time-Series
Feb 19, 2021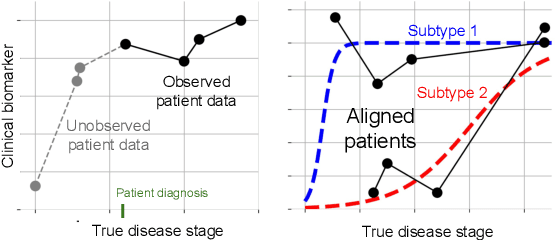
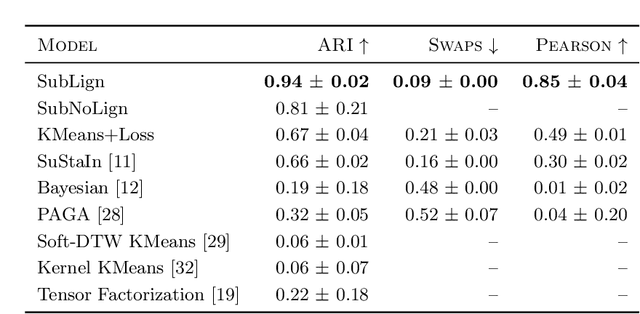
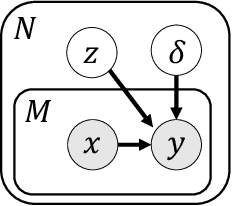
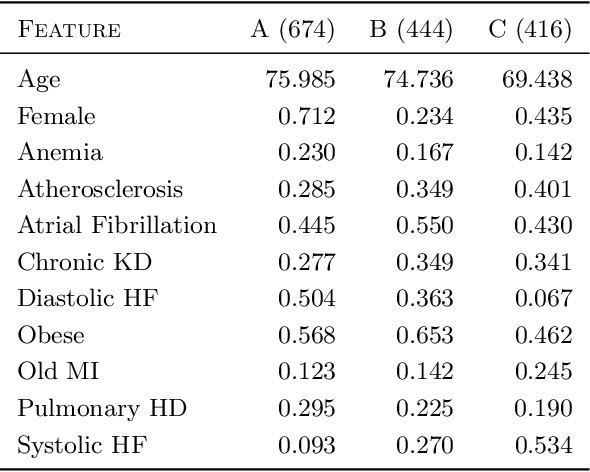
Abstract:Unsupervised learning seeks to uncover patterns in data. However, different kinds of noise may impede the discovery of useful substructure from real-world time-series data. In this work, we focus on mitigating the interference of left-censorship in the task of clustering. We provide conditions under which clusters and left-censorship may be identified; motivated by this result, we develop a deep generative, continuous-time model of time-series data that clusters while correcting for censorship time. We demonstrate accurate, stable, and interpretable results on synthetic data that outperform several benchmarks. To showcase the utility of our framework on real-world problems, we study how left-censorship can adversely affect the task of disease phenotyping, resulting in the often incorrect assumption that longitudinal patient data are aligned by disease stage. In reality, patients at the time of diagnosis are at different stages of the disease -- both late and early due to differences in when patients seek medical care and such discrepancy can confound unsupervised learning algorithms. On two clinical datasets, our model corrects for this form of censorship and recovers known clinical subtypes.
Graph cuts always find a global optimum
Nov 07, 2020
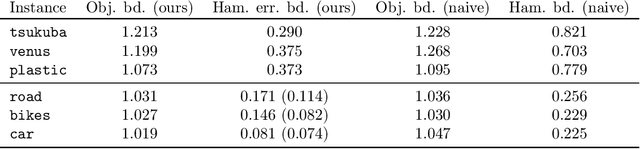

Abstract:We prove that the alpha-expansion algorithm for MAP inference always returns a globally optimal assignment for Markov Random Fields with Potts pairwise potentials, with a catch: the returned assignment is only guaranteed to be optimal in a small perturbation of the original problem instance. In other words, all local minima with respect to expansion moves are global minima to slightly perturbed versions of the problem. On "real-world" instances, MAP assignments of small perturbations of the problem should be very similar to the MAP assignment(s) of the original problem instance. We design an algorithm that can certify whether this is the case in practice. On several MAP inference problem instances from computer vision, this algorithm certifies that MAP solutions to all of these perturbations are very close to solutions of the original instance. These results taken together give a cohesive explanation for the good performance of "graph cuts" algorithms in practice. Every local expansion minimum is a global minimum in a small perturbation of the problem, and all of these global minima are close to the original solution.
Trajectory Inspection: A Method for Iterative Clinician-Driven Design of Reinforcement Learning Studies
Oct 08, 2020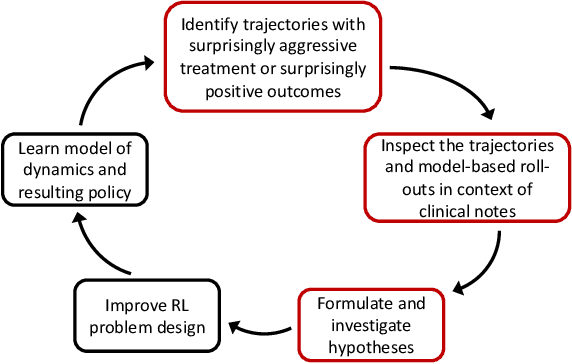
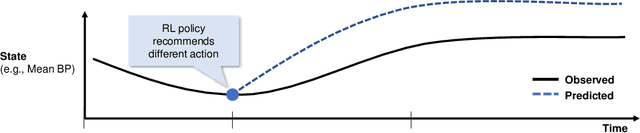
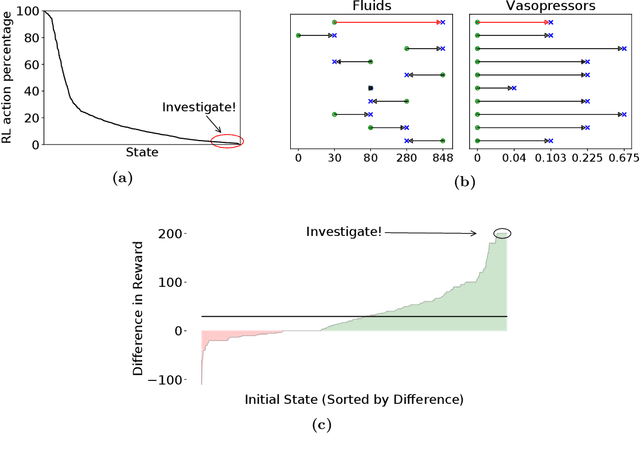
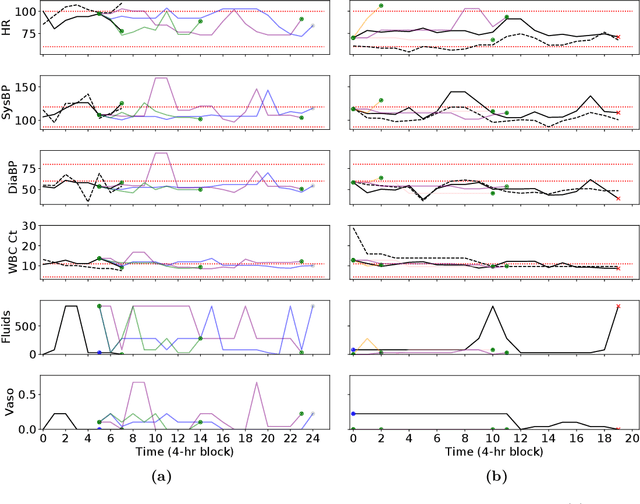
Abstract:Treatment policies learned via reinforcement learning (RL) from observational health data are sensitive to subtle choices in study design. We highlight a simple approach, trajectory inspection, to bring clinicians into an iterative design process for model-based RL studies. We inspect trajectories where the model recommends unexpectedly aggressive treatments or believes its recommendations would lead to much more positive outcomes. Then, we examine clinical trajectories simulated with the learned model and policy alongside the actual hospital course to uncover possible modeling issues. To demonstrate that this approach yields insights, we apply it to recent work on RL for inpatient sepsis management. We find that a design choice around maximum trajectory length leads to a model bias towards discharge, that the RL policy preference for high vasopressor doses may be linked to small sample sizes, and that the model has a clinically implausible expectation of discharge without weaning off vasopressors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge