David M. Blei
Implicit Causal Models for Genome-wide Association Studies
Oct 30, 2017
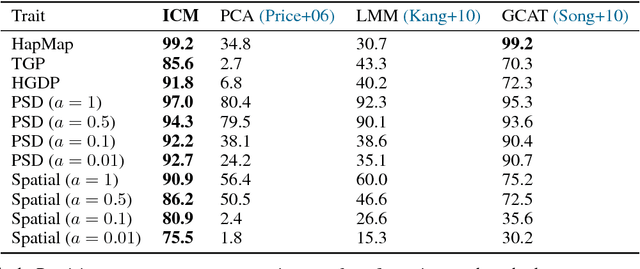
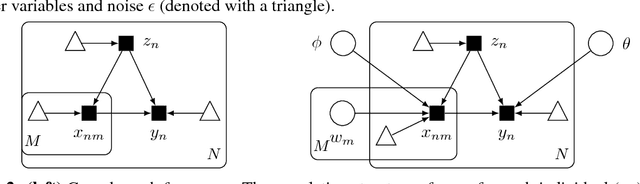
Abstract:Progress in probabilistic generative models has accelerated, developing richer models with neural architectures, implicit densities, and with scalable algorithms for their Bayesian inference. However, there has been limited progress in models that capture causal relationships, for example, how individual genetic factors cause major human diseases. In this work, we focus on two challenges in particular: How do we build richer causal models, which can capture highly nonlinear relationships and interactions between multiple causes? How do we adjust for latent confounders, which are variables influencing both cause and effect and which prevent learning of causal relationships? To address these challenges, we synthesize ideas from causality and modern probabilistic modeling. For the first, we describe implicit causal models, a class of causal models that leverages neural architectures with an implicit density. For the second, we describe an implicit causal model that adjusts for confounders by sharing strength across examples. In experiments, we scale Bayesian inference on up to a billion genetic measurements. We achieve state of the art accuracy for identifying causal factors: we significantly outperform existing genetics methods by an absolute difference of 15-45.3%.
Proximity Variational Inference
May 24, 2017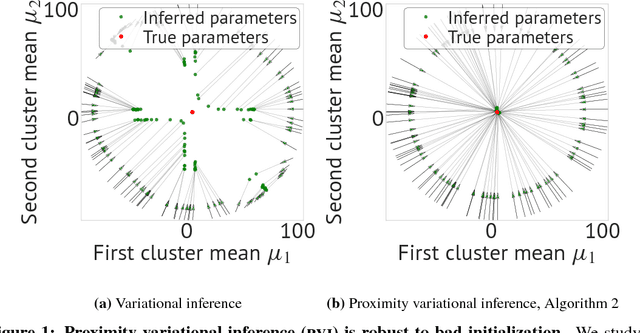
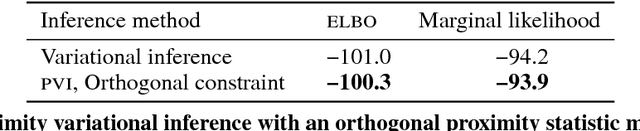

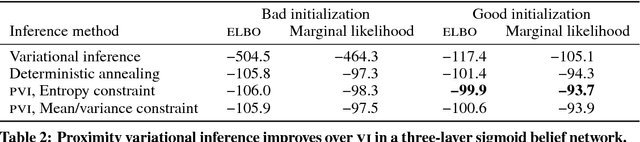

Abstract:Variational inference is a powerful approach for approximate posterior inference. However, it is sensitive to initialization and can be subject to poor local optima. In this paper, we develop proximity variational inference (PVI). PVI is a new method for optimizing the variational objective that constrains subsequent iterates of the variational parameters to robustify the optimization path. Consequently, PVI is less sensitive to initialization and optimization quirks and finds better local optima. We demonstrate our method on three proximity statistics. We study PVI on a Bernoulli factor model and sigmoid belief network with both real and synthetic data and compare to deterministic annealing (Katahira et al., 2008). We highlight the flexibility of PVI by designing a proximity statistic for Bayesian deep learning models such as the variational autoencoder (Kingma and Welling, 2014; Rezende et al., 2014). Empirically, we show that PVI consistently finds better local optima and gives better predictive performance.
Reparameterization Gradients through Acceptance-Rejection Sampling Algorithms
Mar 10, 2017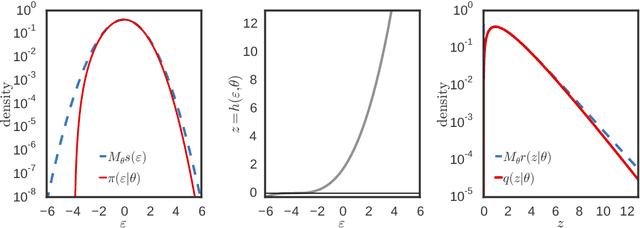
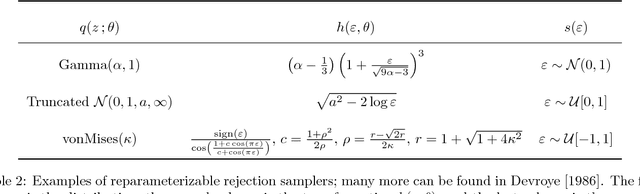

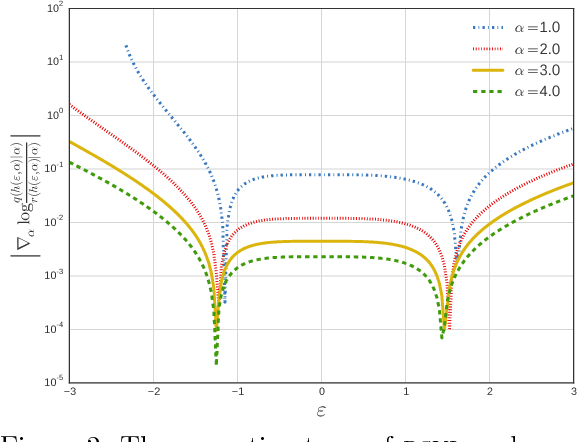

Abstract:Variational inference using the reparameterization trick has enabled large-scale approximate Bayesian inference in complex probabilistic models, leveraging stochastic optimization to sidestep intractable expectations. The reparameterization trick is applicable when we can simulate a random variable by applying a differentiable deterministic function on an auxiliary random variable whose distribution is fixed. For many distributions of interest (such as the gamma or Dirichlet), simulation of random variables relies on acceptance-rejection sampling. The discontinuity introduced by the accept-reject step means that standard reparameterization tricks are not applicable. We propose a new method that lets us leverage reparameterization gradients even when variables are outputs of a acceptance-rejection sampling algorithm. Our approach enables reparameterization on a larger class of variational distributions. In several studies of real and synthetic data, we show that the variance of the estimator of the gradient is significantly lower than other state-of-the-art methods. This leads to faster convergence of stochastic gradient variational inference.
Deep Probabilistic Programming
Mar 07, 2017
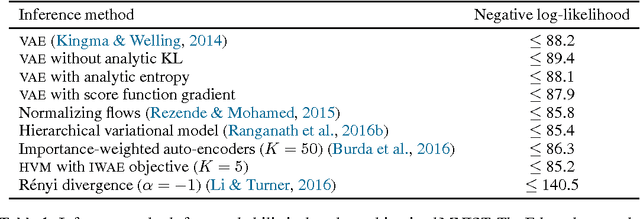
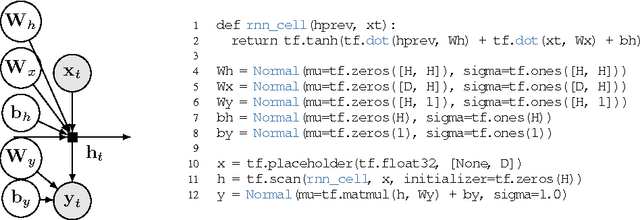



Abstract:We propose Edward, a Turing-complete probabilistic programming language. Edward defines two compositional representations---random variables and inference. By treating inference as a first class citizen, on a par with modeling, we show that probabilistic programming can be as flexible and computationally efficient as traditional deep learning. For flexibility, Edward makes it easy to fit the same model using a variety of composable inference methods, ranging from point estimation to variational inference to MCMC. In addition, Edward can reuse the modeling representation as part of inference, facilitating the design of rich variational models and generative adversarial networks. For efficiency, Edward is integrated into TensorFlow, providing significant speedups over existing probabilistic systems. For example, we show on a benchmark logistic regression task that Edward is at least 35x faster than Stan and 6x faster than PyMC3. Further, Edward incurs no runtime overhead: it is as fast as handwritten TensorFlow.
Edward: A library for probabilistic modeling, inference, and criticism
Feb 01, 2017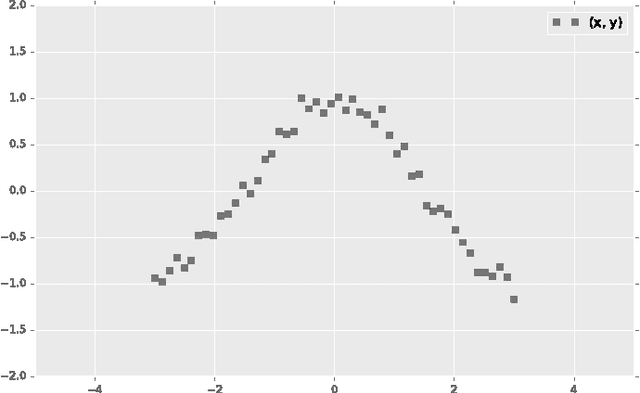
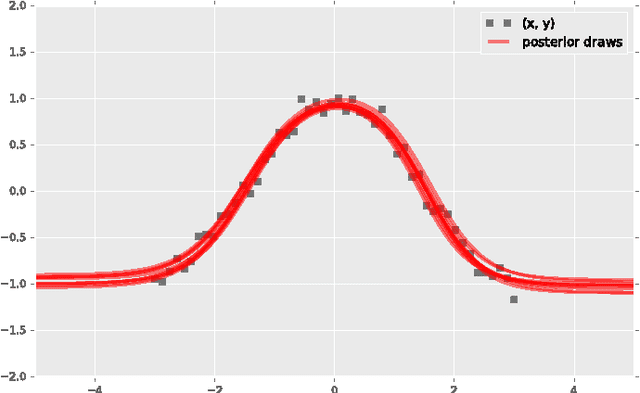
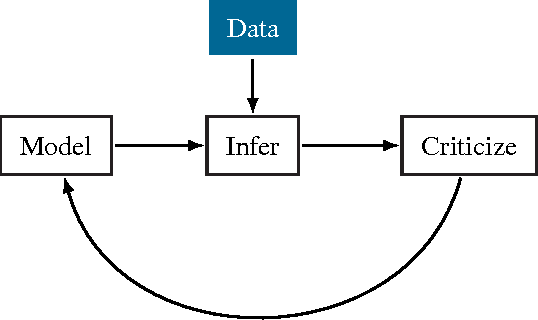
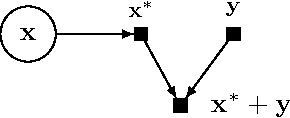
Abstract:Probabilistic modeling is a powerful approach for analyzing empirical information. We describe Edward, a library for probabilistic modeling. Edward's design reflects an iterative process pioneered by George Box: build a model of a phenomenon, make inferences about the model given data, and criticize the model's fit to the data. Edward supports a broad class of probabilistic models, efficient algorithms for inference, and many techniques for model criticism. The library builds on top of TensorFlow to support distributed training and hardware such as GPUs. Edward enables the development of complex probabilistic models and their algorithms at a massive scale.
Exponential Family Embeddings
Nov 21, 2016
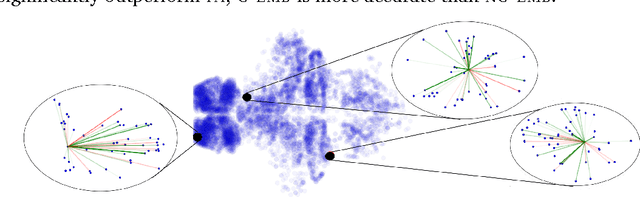

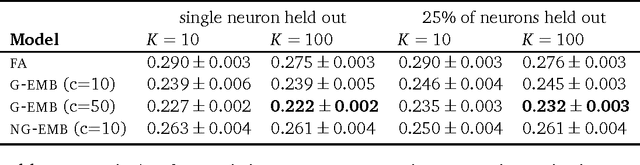
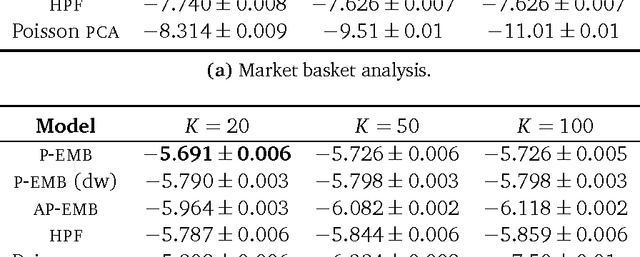
Abstract:Word embeddings are a powerful approach for capturing semantic similarity among terms in a vocabulary. In this paper, we develop exponential family embeddings, a class of methods that extends the idea of word embeddings to other types of high-dimensional data. As examples, we studied neural data with real-valued observations, count data from a market basket analysis, and ratings data from a movie recommendation system. The main idea is to model each observation conditioned on a set of other observations. This set is called the context, and the way the context is defined is a modeling choice that depends on the problem. In language the context is the surrounding words; in neuroscience the context is close-by neurons; in market basket data the context is other items in the shopping cart. Each type of embedding model defines the context, the exponential family of conditional distributions, and how the latent embedding vectors are shared across data. We infer the embeddings with a scalable algorithm based on stochastic gradient descent. On all three applications - neural activity of zebrafish, users' shopping behavior, and movie ratings - we found exponential family embedding models to be more effective than other types of dimension reduction. They better reconstruct held-out data and find interesting qualitative structure.
Recurrent switching linear dynamical systems
Oct 26, 2016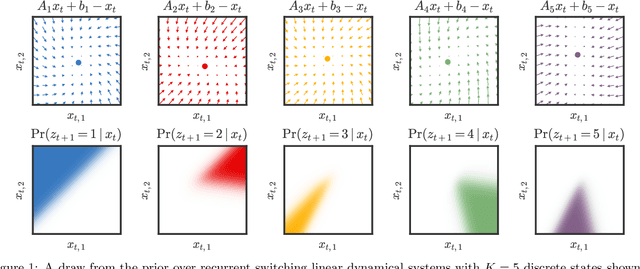
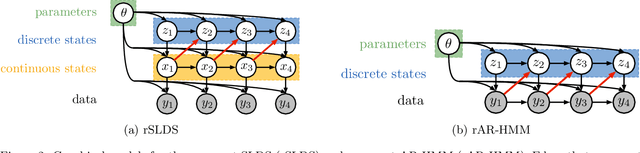
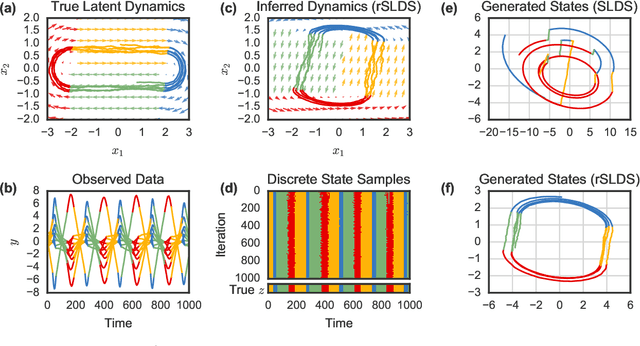
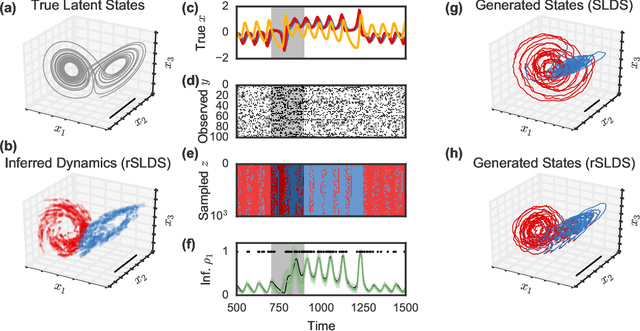
Abstract:Many natural systems, such as neurons firing in the brain or basketball teams traversing a court, give rise to time series data with complex, nonlinear dynamics. We can gain insight into these systems by decomposing the data into segments that are each explained by simpler dynamic units. Building on switching linear dynamical systems (SLDS), we present a new model class that not only discovers these dynamical units, but also explains how their switching behavior depends on observations or continuous latent states. These "recurrent" switching linear dynamical systems provide further insight by discovering the conditions under which each unit is deployed, something that traditional SLDS models fail to do. We leverage recent algorithmic advances in approximate inference to make Bayesian inference in these models easy, fast, and scalable.
The Generalized Reparameterization Gradient
Oct 19, 2016
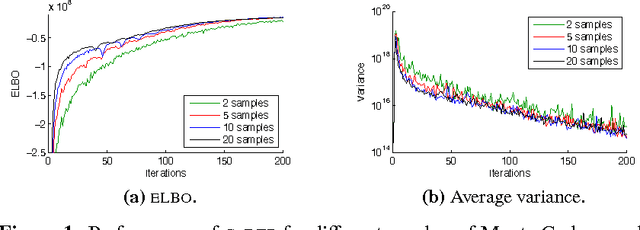
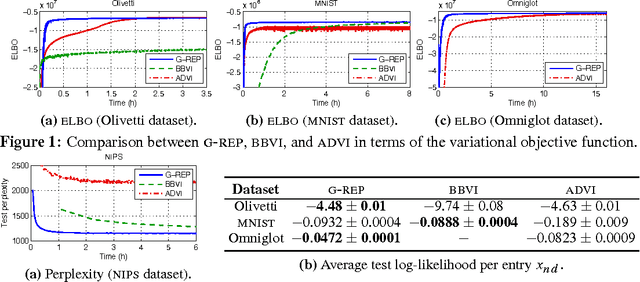

Abstract:The reparameterization gradient has become a widely used method to obtain Monte Carlo gradients to optimize the variational objective. However, this technique does not easily apply to commonly used distributions such as beta or gamma without further approximations, and most practical applications of the reparameterization gradient fit Gaussian distributions. In this paper, we introduce the generalized reparameterization gradient, a method that extends the reparameterization gradient to a wider class of variational distributions. Generalized reparameterizations use invertible transformations of the latent variables which lead to transformed distributions that weakly depend on the variational parameters. This results in new Monte Carlo gradients that combine reparameterization gradients and score function gradients. We demonstrate our approach on variational inference for two complex probabilistic models. The generalized reparameterization is effective: even a single sample from the variational distribution is enough to obtain a low-variance gradient.
A General Method for Robust Bayesian Modeling
Sep 07, 2016
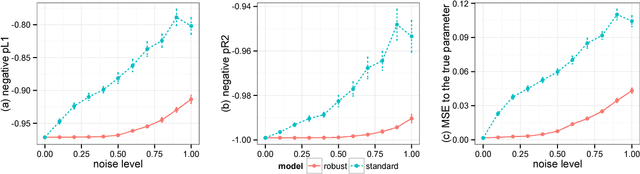
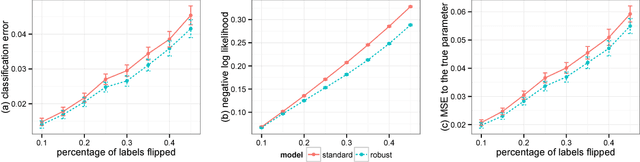
Abstract:Robust Bayesian models are appealing alternatives to standard models, providing protection from data that contains outliers or other departures from the model assumptions. Historically, robust models were mostly developed on a case-by-case basis; examples include robust linear regression, robust mixture models, and bursty topic models. In this paper we develop a general approach to robust Bayesian modeling. We show how to turn an existing Bayesian model into a robust model, and then develop a generic strategy for computing with it. We use our method to study robust variants of several models, including linear regression, Poisson regression, logistic regression, and probabilistic topic models. We discuss the connections between our methods and existing approaches, especially empirical Bayes and James-Stein estimation.
Bayesian Poisson Tucker Decomposition for Learning the Structure of International Relations
Jun 06, 2016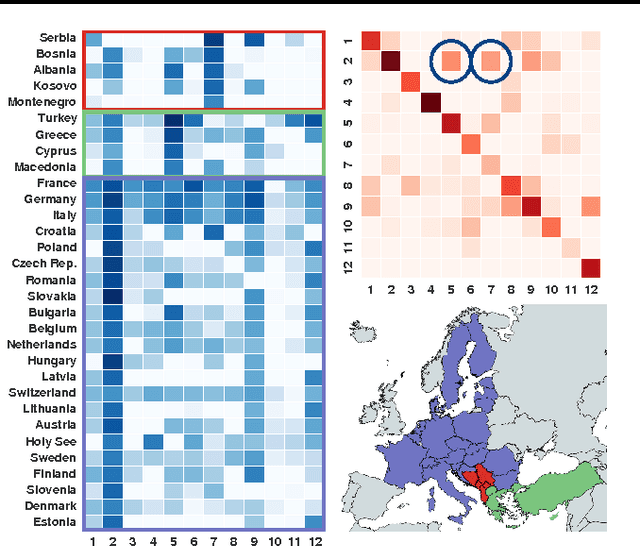
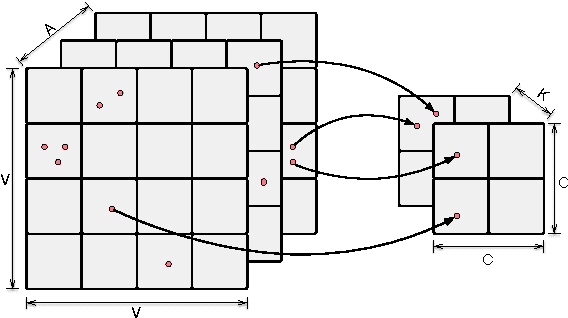
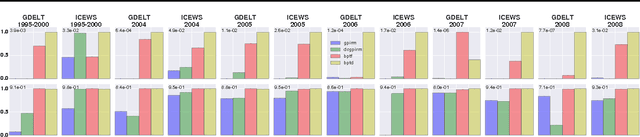

Abstract:We introduce Bayesian Poisson Tucker decomposition (BPTD) for modeling country--country interaction event data. These data consist of interaction events of the form "country $i$ took action $a$ toward country $j$ at time $t$." BPTD discovers overlapping country--community memberships, including the number of latent communities. In addition, it discovers directed community--community interaction networks that are specific to "topics" of action types and temporal "regimes." We show that BPTD yields an efficient MCMC inference algorithm and achieves better predictive performance than related models. We also demonstrate that it discovers interpretable latent structure that agrees with our knowledge of international relations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge