Daniel S. Weld
University of Washington, Allen Institute for Artificial Intelligence
Exploring The Role of Local and Global Explanations in Recommender Systems
Sep 27, 2021

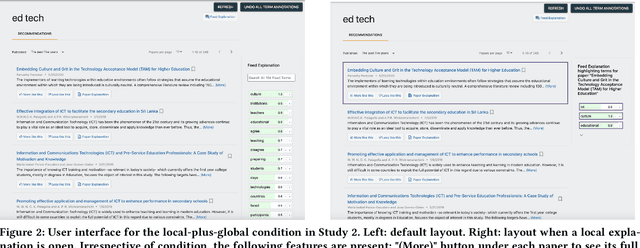

Abstract:Explanations are well-known to improve recommender systems' transparency. These explanations may be local, explaining an individual recommendation, or global, explaining the recommender model in general. Despite their widespread use, there has been little investigation into the relative benefits of these two approaches. Do they provide the same benefits to users, or do they serve different purposes? We conducted a 30-participant exploratory study and a 30-participant controlled user study with a research-paper recommender system to analyze how providing participants local, global, or both explanations influences user understanding of system behavior. Our results provide evidence suggesting that both explanations are more helpful than either alone for explaining how to improve recommendations, yet both appeared less helpful than global alone for efficiency in identifying false positives and negatives. However, we note that the two explanation approaches may be better compared in the context of a higher-stakes or more opaque domain.
A Search Engine for Discovery of Scientific Challenges and Directions
Sep 10, 2021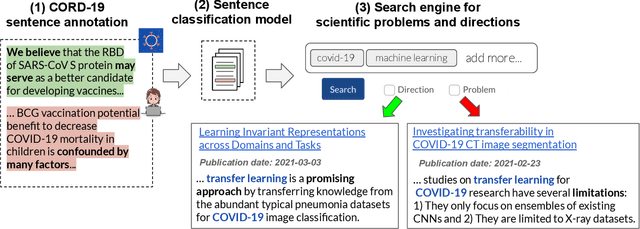


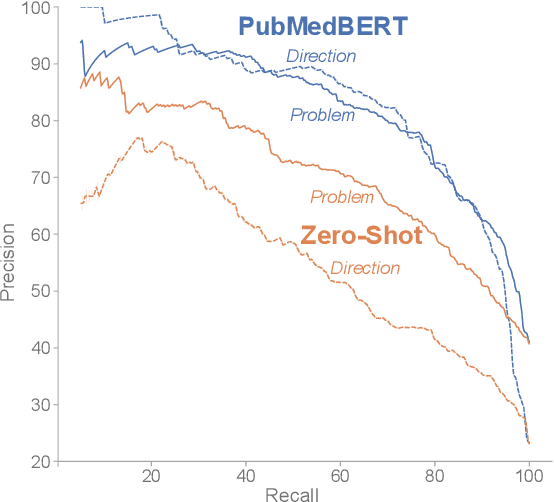
Abstract:Keeping track of scientific challenges, advances and emerging directions is a fundamental part of research. However, researchers face a flood of papers that hinders discovery of important knowledge. In biomedicine, this directly impacts human lives. To address this problem, we present a novel task of extraction and search of scientific challenges and directions, to facilitate rapid knowledge discovery. We construct and release an expert-annotated corpus of texts sampled from full-length papers, labeled with novel semantic categories that generalize across many types of challenges and directions. We focus on a large corpus of interdisciplinary work relating to the COVID-19 pandemic, ranging from biomedicine to areas such as AI and economics. We apply a model trained on our data to identify challenges and directions across the corpus and build a dedicated search engine. In experiments with 19 researchers and clinicians using our system, we outperform a popular scientific search engine in assisting knowledge discovery. Finally, we show that models trained on our resource generalize to the wider biomedical domain and to AI papers, highlighting its broad utility. We make our data, model and search engine publicly available. https://challenges.apps.allenai.org/
Incorporating Visual Layout Structures for Scientific Text Classification
Jun 21, 2021
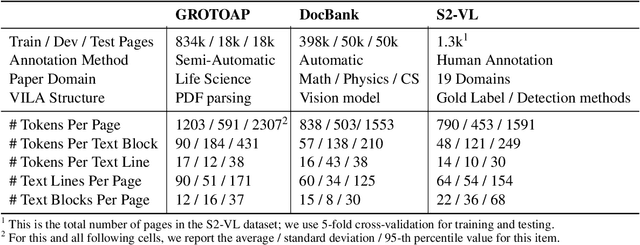
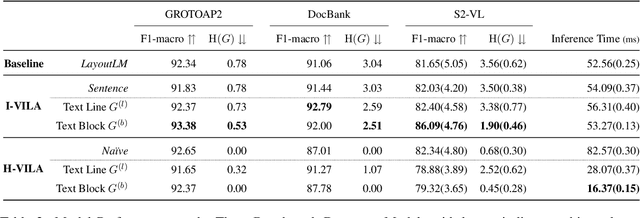
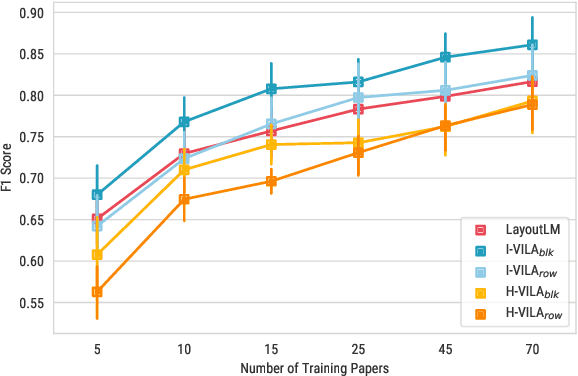
Abstract:Classifying the core textual components of a scientific paper-title, author, body text, etc.-is a critical first step in automated scientific document understanding. Previous work has shown how using elementary layout information, i.e., each token's 2D position on the page, leads to more accurate classification. We introduce new methods for incorporating VIsual LAyout (VILA) structures, e.g., the grouping of page texts into text lines or text blocks, into language models to further improve performance. We show that the I-VILA approach, which simply adds special tokens denoting the boundaries of layout structures into model inputs, can lead to 1.9% Macro F1 improvements for token classification. Moreover, we design a hierarchical model, H-VILA, that encodes the text based on layout structures and record an up-to 47% inference time reduction with less than 1.5% Macro F1 loss for the text classification models. Experiments are conducted on a newly curated evaluation suite, S2-VLUE, with a novel metric measuring classification uniformity within visual groups and a new dataset of gold annotations covering papers from 19 scientific disciplines. Pre-trained weights, benchmark datasets, and source code will be available at https://github.com/allenai/VILA.
GENIE: A Leaderboard for Human-in-the-Loop Evaluation of Text Generation
Jan 17, 2021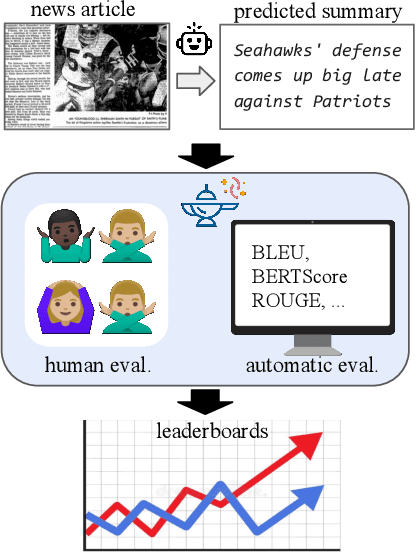
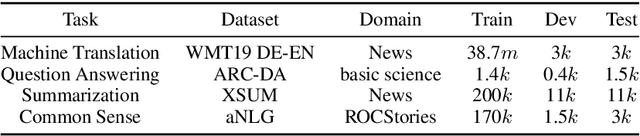

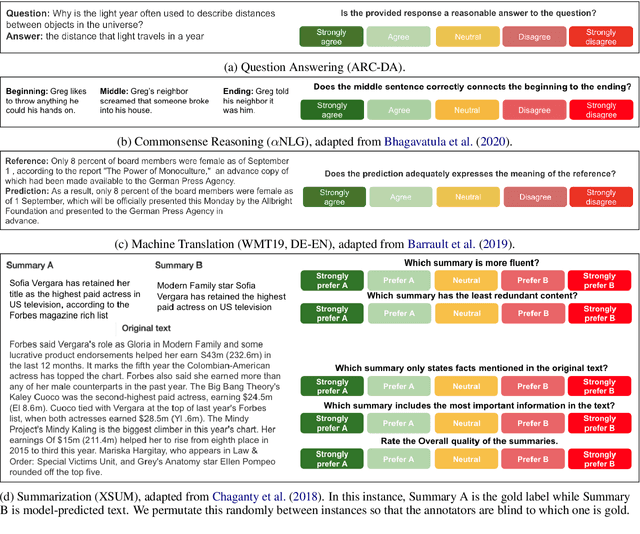
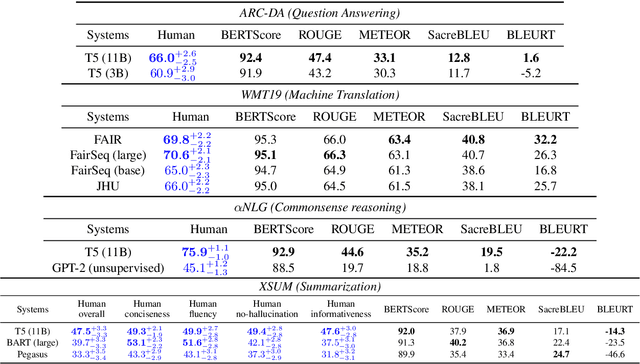
Abstract:Leaderboards have eased model development for many NLP datasets by standardizing their evaluation and delegating it to an independent external repository. Their adoption, however, is so far limited to tasks that can be reliably evaluated in an automatic manner. This work introduces GENIE, an extensible human evaluation leaderboard, which brings the ease of leaderboards to text generation tasks. GENIE automatically posts leaderboard submissions to crowdsourcing platforms asking human annotators to evaluate them on various axes (e.g., correctness, conciseness, fluency) and compares their answers to various automatic metrics. We introduce several datasets in English to GENIE, representing four core challenges in text generation: machine translation, summarization, commonsense reasoning, and machine comprehension. We provide formal granular evaluation metrics and identify areas for future research. We make GENIE publicly available and hope that it will spur progress in language generation models as well as their automatic and manual evaluation.
Polyjuice: Automated, General-purpose Counterfactual Generation
Jan 01, 2021
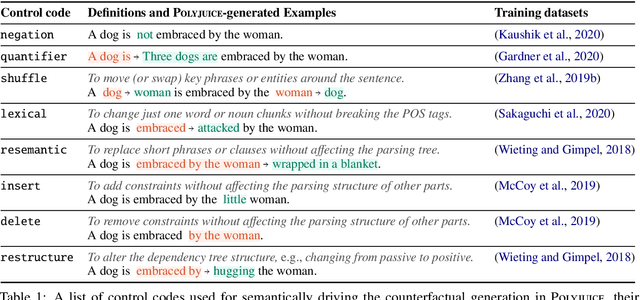

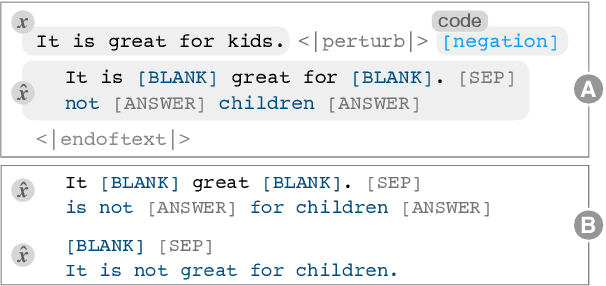
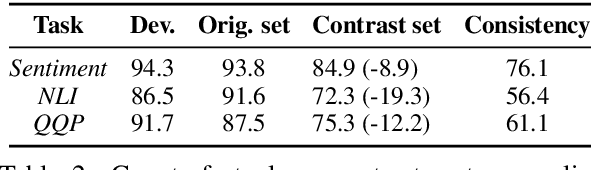
Abstract:Counterfactual examples have been shown to be useful for many applications, including calibrating, evaluating, and explaining model decision boundaries. However, previous methods for generating such counterfactual examples have been tightly tailored to a specific application, used a limited range of linguistic patterns, or are hard to scale. We propose to disentangle counterfactual generation from its use cases, i.e., gather general-purpose counterfactuals first, and then select them for specific applications. We frame the automated counterfactual generation as text generation, and finetune GPT-2 into a generator, Polyjuice, which produces fluent and diverse counterfactuals. Our method also allows control over where perturbations happen and what they do. We show Polyjuice supports multiple use cases: by generating diverse counterfactuals for humans to label, Polyjuice helps produce high-quality datasets for model training and evaluation, requiring 40% less human effort. When used to generate explanations, Polyjuice helps augment feature attribution methods to reveal models' erroneous behaviors.
Document-Level Definition Detection in Scholarly Documents: Existing Models, Error Analyses, and Future Directions
Oct 11, 2020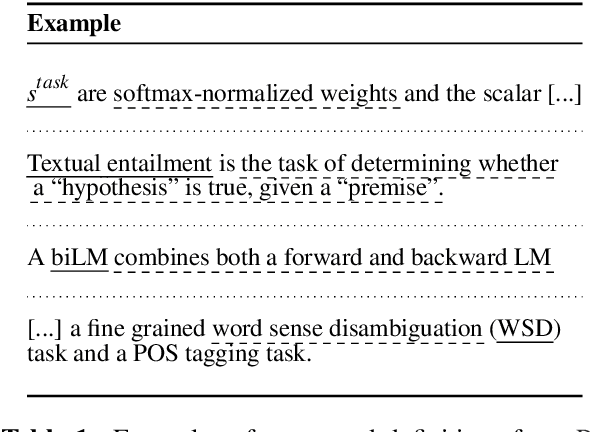
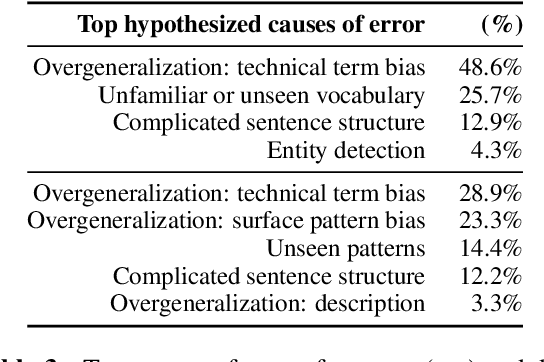
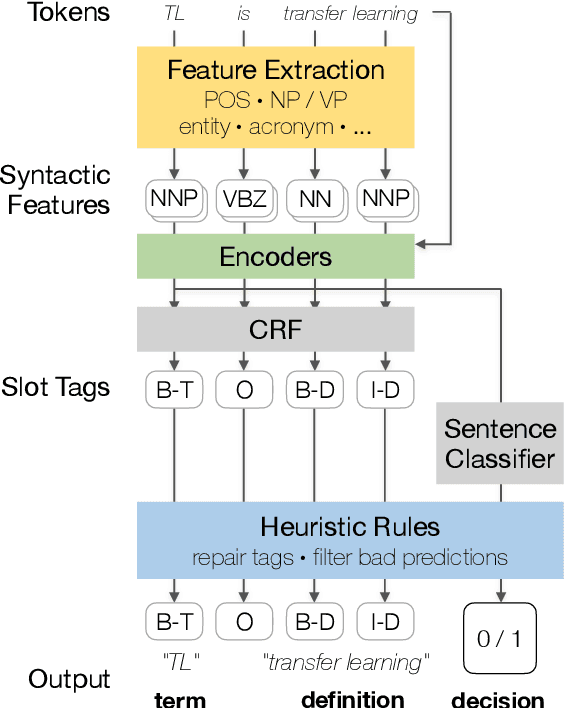


Abstract:The task of definition detection is important for scholarly papers, because papers often make use of technical terminology that may be unfamiliar to readers. Despite prior work on definition detection, current approaches are far from being accurate enough to use in real-world applications. In this paper, we first perform in-depth error analysis of the current best performing definition detection system and discover major causes of errors. Based on this analysis, we develop a new definition detection system, HEDDEx, that utilizes syntactic features, transformer encoders, and heuristic filters, and evaluate it on a standard sentence-level benchmark. Because current benchmarks evaluate randomly sampled sentences, we propose an alternative evaluation that assesses every sentence within a document. This allows for evaluating recall in addition to precision. HEDDEx outperforms the leading system on both the sentence-level and the document-level tasks, by 12.7 F1 points and 14.4 F1 points, respectively. We note that performance on the high-recall document-level task is much lower than in the standard evaluation approach, due to the necessity of incorporation of document structure as features. We discuss remaining challenges in document-level definition detection, ideas for improvements, and potential issues for the development of reading aid applications.
Augmenting Scientific Papers with Just-in-Time, Position-Sensitive Definitions of Terms and Symbols
Sep 29, 2020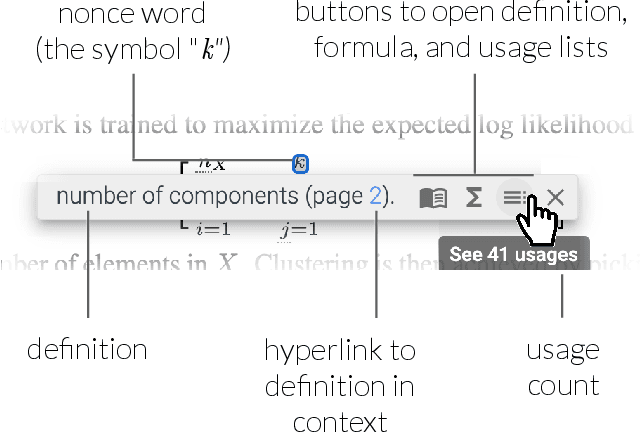
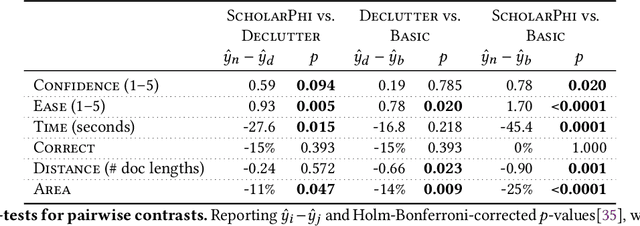
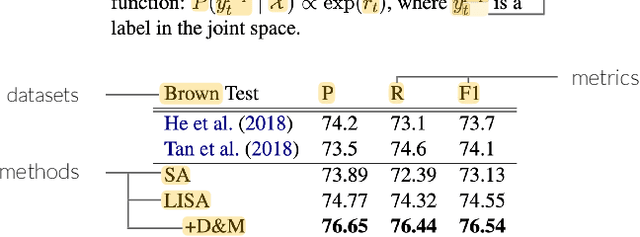

Abstract:Despite the central importance of research papers to scientific progress, they can be difficult to read. Comprehension is often stymied when the information needed to understand a passage resides somewhere else: in another section, or in another paper. In this work, we envision how interfaces can bring definitions of technical terms and symbols to readers when and where they need them most. We introduce ScholarPhi, an augmented reading interface with four novel features: (1) tooltips that surface position-sensitive definitions from elsewhere in a paper, (2) a filter over the paper that "declutters" it to reveal how the term or symbol is used across the paper, (3) automatic equation diagrams that expose multiple definitions in parallel, and (4) an automatically generated glossary of important terms and symbols. A usability study showed that the tool helps researchers of all experience levels read papers. Furthermore, researchers were eager to have ScholarPhi's definitions available to support their everyday reading.
Does the Whole Exceed its Parts? The Effect of AI Explanations on Complementary Team Performance
Jun 30, 2020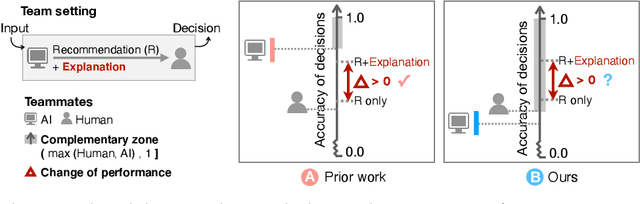
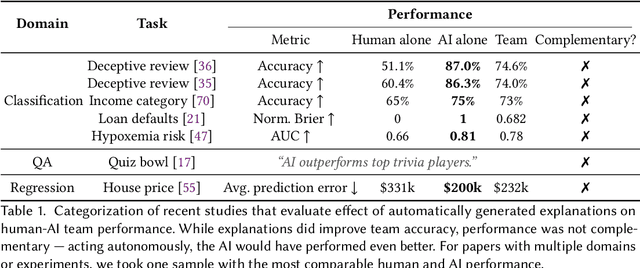
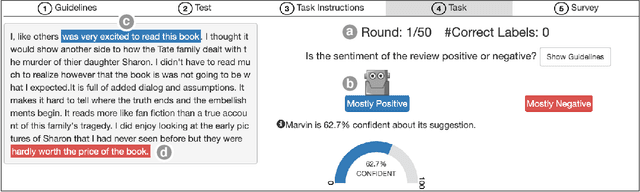
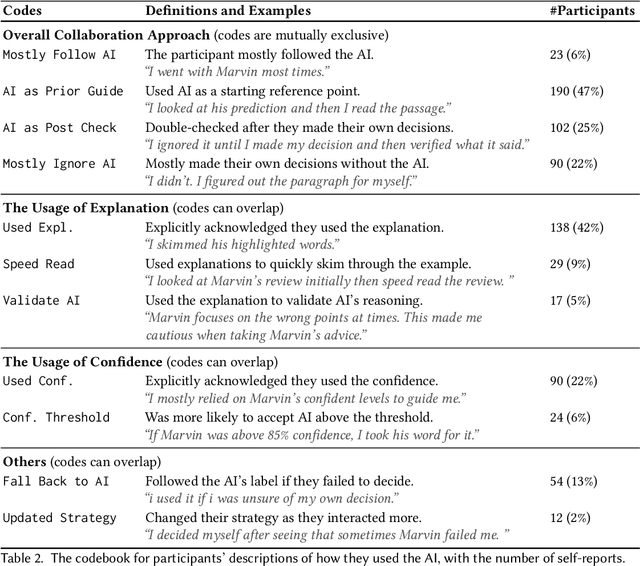
Abstract:Increasingly, organizations are pairing humans with AI systems to improve decision-making and reducing costs. Proponents of human-centered AI argue that team performance can even further improve when the AI model explains its recommendations. However, a careful analysis of existing literature reveals that prior studies observed improvements due to explanations only when the AI, alone, outperformed both the human and the best human-AI team. This raises an important question: can explanations lead to complementary performance, i.e., with accuracy higher than both the human and the AI working alone? We address this question by devising comprehensive studies on human-AI teaming, where participants solve a task with help from an AI system without explanations and from one with varying types of AI explanation support. We carefully controlled to ensure comparable human and AI accuracy across experiments on three NLP datasets (two for sentiment analysis and one for question answering). While we found complementary improvements from AI augmentation, they were not increased by state-of-the-art explanations compared to simpler strategies, such as displaying the AI's confidence. We show that explanations increase the chance that humans will accept the AI's recommendation regardless of whether the AI is correct. While this clarifies the gains in team performance from explanations in prior work, it poses new challenges for human-centered AI: how can we best design systems to produce complementary performance? Can we develop explanatory approaches that help humans decide whether and when to trust AI input?
High-Precision Extraction of Emerging Concepts from Scientific Literature
Jun 11, 2020
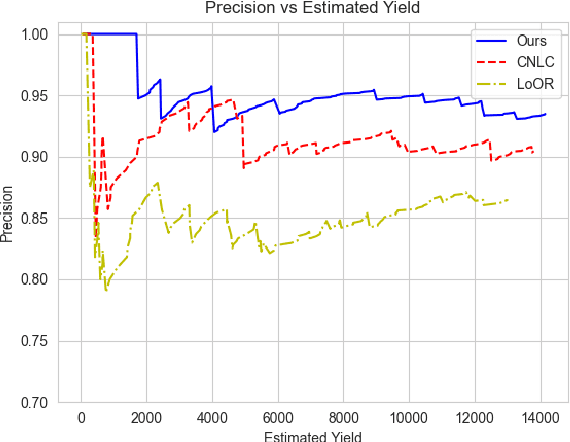
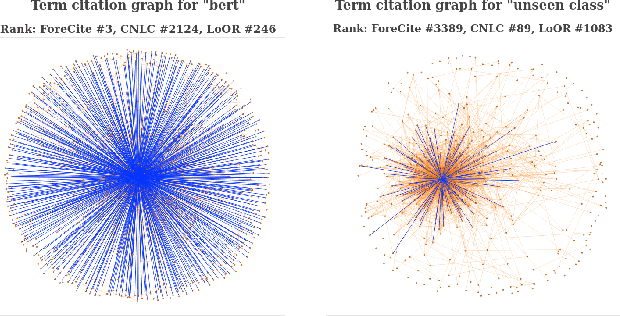
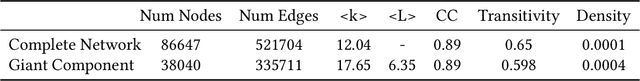
Abstract:Identification of new concepts in scientific literature can help power faceted search, scientific trend analysis, knowledge-base construction, and more, but current methods are lacking. Manual identification cannot keep up with the torrent of new publications, while the precision of existing automatic techniques is too low for many applications. We present an unsupervised concept extraction method for scientific literature that achieves much higher precision than previous work. Our approach relies on a simple but novel intuition: each scientific concept is likely to be introduced or popularized by a single paper that is disproportionately cited by subsequent papers mentioning the concept. From a corpus of computer science papers on arXiv, we find that our method achieves a Precision@1000 of 99%, compared to 86% for prior work, and a substantially better precision-yield trade-off across the top 15,000 extractions. To stimulate research in this area, we release our code and data (https://github.com/allenai/ForeCite).
SciSight: Combining faceted navigation and research group detection for COVID-19 exploratory scientific search
May 27, 2020
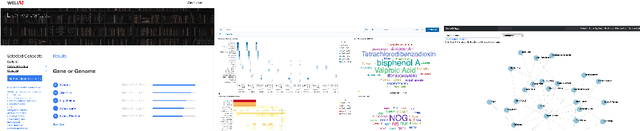
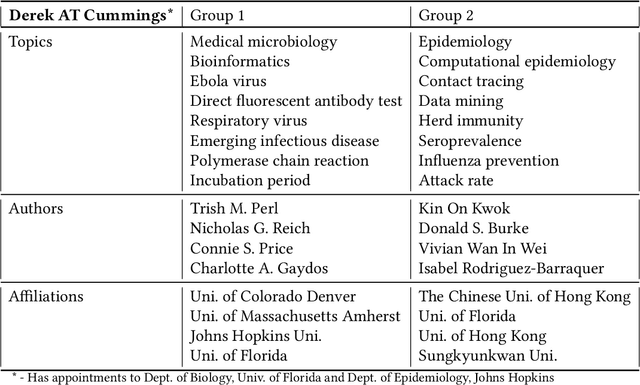
Abstract:The COVID-19 pandemic has sparked unprecedented mobilization of scientists, already generating thousands of new papers that join a litany of previous biomedical work in related areas. This deluge of information makes it hard for researchers to keep track of their own research area, let alone explore new directions. Standard search engines are designed primarily for targeted search and are not geared for discovery or making connections that are not obvious from reading individual papers. In this paper, we present our ongoing work on SciSight, a novel framework for exploratory search of COVID-19 research. Based on formative interviews with scientists and a review of existing tools, we build and integrate two key capabilities: first, exploring interactions between biomedical facets (e.g., proteins, genes, drugs, diseases, patient characteristics); and second, discovering groups of researchers and how they are connected. We extract entities using a language model pre-trained on several biomedical information extraction tasks, and enrich them with data from the Microsoft Academic Graph (MAG). To find research groups automatically, we use hierarchical clustering with overlap to allow authors, as they do, to belong to multiple groups. Finally, we introduce a novel presentation of these groups based on both topical and social affinities, allowing users to drill down from groups to papers to associations between entities, and update query suggestions on the fly with the goal of facilitating exploratory navigation. SciSight has thus far served over 10K users with over 30K page views and 13% returning users. Preliminary user interviews with biomedical researchers suggest that SciSight complements current approaches and helps find new and relevant knowledge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge