Ching-Ping Wang
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
Airway Tree Modeling Using Dual-channel 3D UNet 3+ with Vesselness Prior
Aug 30, 2022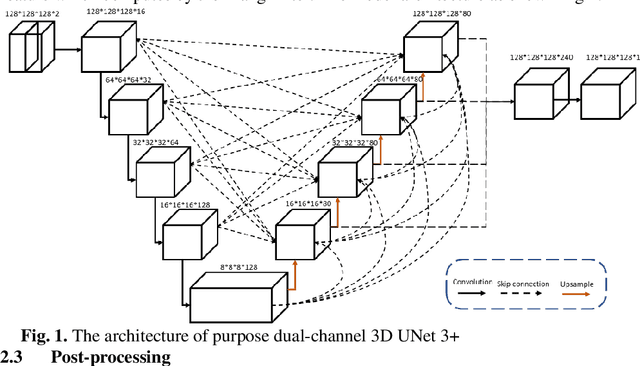
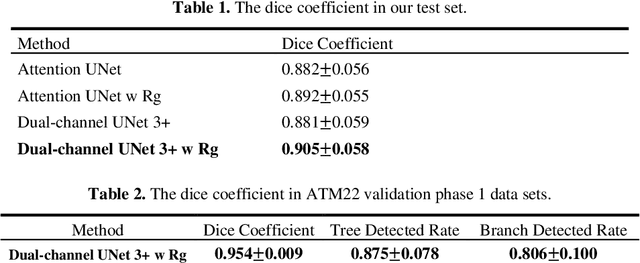
Abstract:The lung airway tree modeling is essential to work for the diagnosis of pulmonary diseases, especially for X-Ray computed tomography (CT). The airway tree modeling on CT images can provide the experts with 3-dimension measurements like wall thickness, etc. This information can tremendously aid the diagnosis of pulmonary diseases like chronic obstructive pulmonary disease [1-4]. Many scholars have attempted various ways to model the lung airway tree, which can be split into two major categories based on its nature. Namely, the model-based approach and the deep learning approach. The performance of a typical model-based approach usually depends on the manual tuning of the model parameter, which can be its advantages and disadvantages. The advantage is its don't require a large amount of training data which can be beneficial for a small dataset like medical imaging. On the other hand, the performance of model-based may be a misconcep-tion [5,6]. In recent years, deep learning has achieved good results in the field of medical image processing, and many scholars have used UNet-based methods in medical image segmentation [7-11]. Among all the variation of UNet, the UNet 3+ [11] have relatively good result compare to the rest of the variation of UNet. Therefor to further improve the accuracy of lung airway tree modeling, this study combines the Frangi filter [5] with UNet 3+ [11] to develop a dual-channel 3D UNet 3+. The Frangi filter is used to extracting vessel-like feature. The vessel-like feature then used as input to guide the dual-channel UNet 3+ training and testing procedures.
Using Multi-scale SwinTransformer-HTC with Data augmentation in CoNIC Challenge
Mar 02, 2022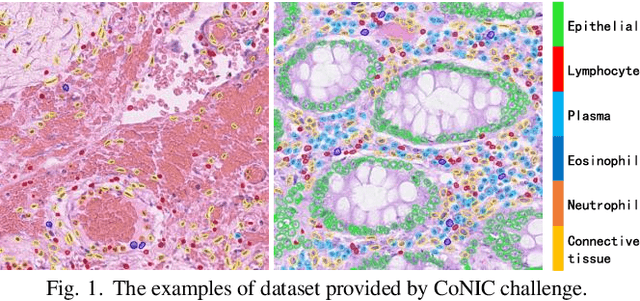
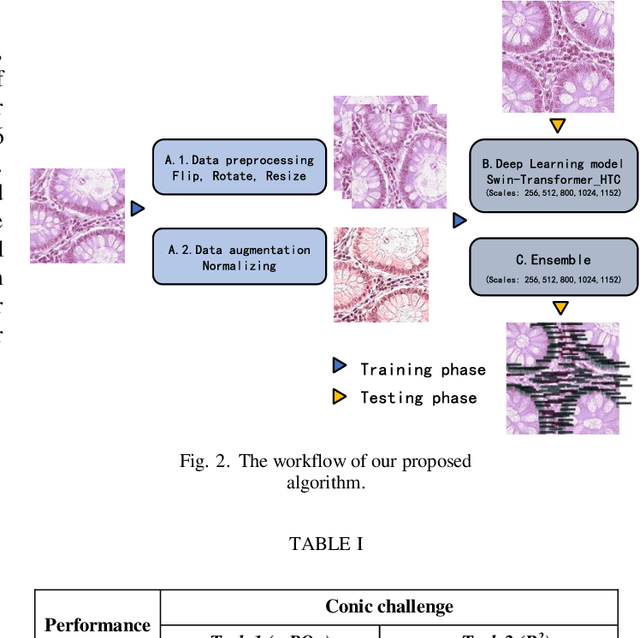
Abstract:Colorectal cancer is one of the most common cancers worldwide, so early pathological examination is very important. However, it is time-consuming and labor-intensive to identify the number and type of cells on H&E images in clinical. Therefore, automatic segmentation and classification task and counting the cellular composition of H&E images from pathological sections is proposed by CoNIC Challenge 2022. We proposed a multi-scale Swin transformer with HTC for this challenge, and also applied the known normalization methods to generate more augmentation data. Finally, our strategy showed that the multi-scale played a crucial role to identify different scale features and the augmentation arose the recognition of model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge