ChengXiang Zhai
oBERTa: Improving Sparse Transfer Learning via improved initialization, distillation, and pruning regimes
Apr 04, 2023



Abstract:In this paper, we introduce the range of oBERTa language models, an easy-to-use set of language models which allows Natural Language Processing (NLP) practitioners to obtain between 3.8 and 24.3 times faster models without expertise in model compression. Specifically, oBERTa extends existing work on pruning, knowledge distillation, and quantization and leverages frozen embeddings improves distillation and model initialization to deliver higher accuracy on a broad range of transfer tasks. In generating oBERTa, we explore how the highly optimized RoBERTa differs from the BERT for pruning during pre-training and finetuning. We find it less amenable to compression during fine-tuning. We explore the use of oBERTa on seven representative NLP tasks and find that the improved compression techniques allow a pruned oBERTa model to match the performance of BERTbase and exceed the performance of Prune OFA Large on the SQUAD V1.1 Question Answering dataset, despite being 8x and 2x, respectively faster in inference. We release our code, training regimes, and associated model for broad usage to encourage usage and experimentation
Dense Sparse Retrieval: Using Sparse Language Models for Inference Efficient Dense Retrieval
Mar 31, 2023



Abstract:Vector-based retrieval systems have become a common staple for academic and industrial search applications because they provide a simple and scalable way of extending the search to leverage contextual representations for documents and queries. As these vector-based systems rely on contextual language models, their usage commonly requires GPUs, which can be expensive and difficult to manage. Given recent advances in introducing sparsity into language models for improved inference efficiency, in this paper, we study how sparse language models can be used for dense retrieval to improve inference efficiency. Using the popular retrieval library Tevatron and the MSMARCO, NQ, and TriviaQA datasets, we find that sparse language models can be used as direct replacements with little to no drop in accuracy and up to 4.3x improved inference speeds
Competence-Based Analysis of Language Models
Mar 01, 2023



Abstract:Despite the recent success of large pretrained language models (LMs) on a variety of prompting tasks, these models can be alarmingly brittle to small changes in inputs or application contexts. To better understand such behavior and motivate the design of more robust LMs, we propose a general experimental framework, CALM (Competence-based Analysis of Language Models), where targeted causal interventions are utilized to damage an LM's internal representation of various linguistic properties in order to evaluate its use of each representation in performing a given task. We implement these interventions as gradient-based adversarial attacks, which (in contrast to prior causal probing methodologies) are able to target arbitrarily-encoded representations of relational properties, and carry out a case study of this approach to analyze how BERT-like LMs use representations of several relational properties in performing associated relation prompting tasks. We find that, while the representations LMs leverage in performing each task are highly entangled, they may be meaningfully interpreted in terms of the tasks where they are most utilized; and more broadly, that CALM enables an expanded scope of inquiry in LM analysis that may be useful in predicting and explaining weaknesses of existing LMs.
Entity Set Co-Expansion in StackOverflow
Dec 05, 2022


Abstract:Given a few seed entities of a certain type (e.g., Software or Programming Language), entity set expansion aims to discover an extensive set of entities that share the same type as the seeds. Entity set expansion in software-related domains such as StackOverflow can benefit many downstream tasks (e.g., software knowledge graph construction) and facilitate better IT operations and service management. Meanwhile, existing approaches are less concerned with two problems: (1) How to deal with multiple types of seed entities simultaneously? (2) How to leverage the power of pre-trained language models (PLMs)? Being aware of these two problems, in this paper, we study the entity set co-expansion task in StackOverflow, which extracts Library, OS, Application, and Language entities from StackOverflow question-answer threads. During the co-expansion process, we use PLMs to derive embeddings of candidate entities for calculating similarities between entities. Experimental results show that our proposed SECoExpan framework outperforms previous approaches significantly.
CONCRETE: Improving Cross-lingual Fact-checking with Cross-lingual Retrieval
Sep 05, 2022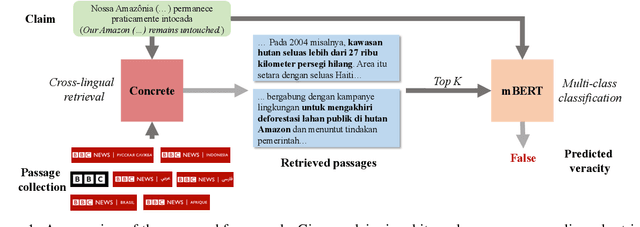
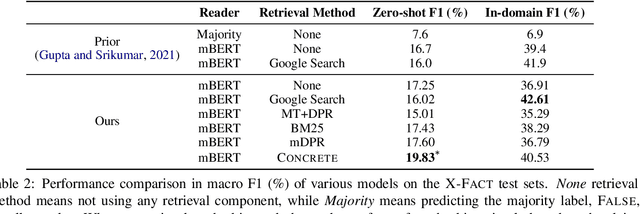
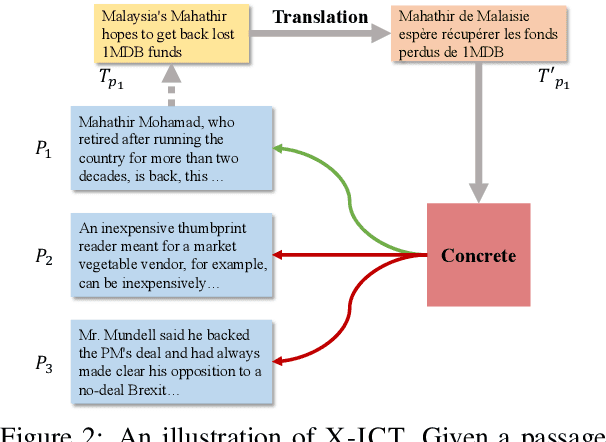

Abstract:Fact-checking has gained increasing attention due to the widespread of falsified information. Most fact-checking approaches focus on claims made in English only due to the data scarcity issue in other languages. The lack of fact-checking datasets in low-resource languages calls for an effective cross-lingual transfer technique for fact-checking. Additionally, trustworthy information in different languages can be complementary and helpful in verifying facts. To this end, we present the first fact-checking framework augmented with cross-lingual retrieval that aggregates evidence retrieved from multiple languages through a cross-lingual retriever. Given the absence of cross-lingual information retrieval datasets with claim-like queries, we train the retriever with our proposed Cross-lingual Inverse Cloze Task (X-ICT), a self-supervised algorithm that creates training instances by translating the title of a passage. The goal for X-ICT is to learn cross-lingual retrieval in which the model learns to identify the passage corresponding to a given translated title. On the X-Fact dataset, our approach achieves 2.23% absolute F1 improvement in the zero-shot cross-lingual setup over prior systems. The source code and data are publicly available at https://github.com/khuangaf/CONCRETE.
Sparse*BERT: Sparse Models are Robust
May 25, 2022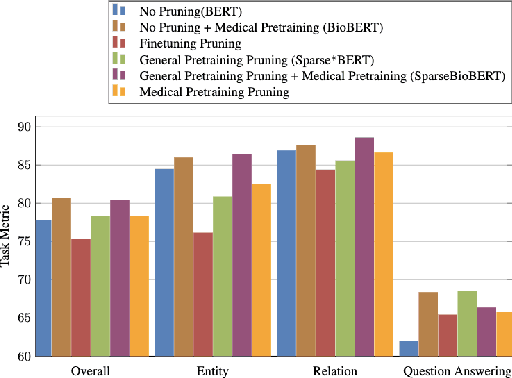
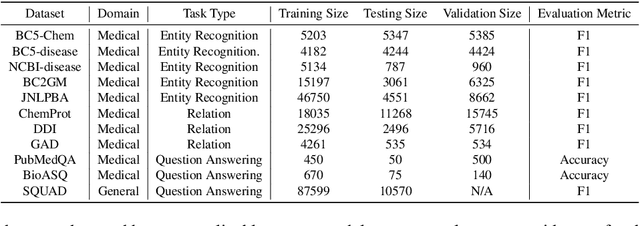
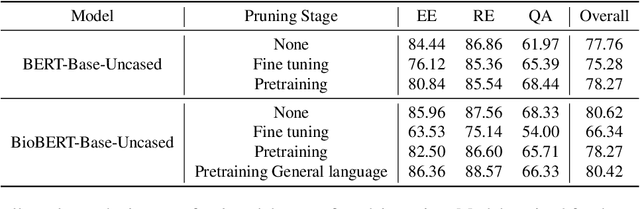
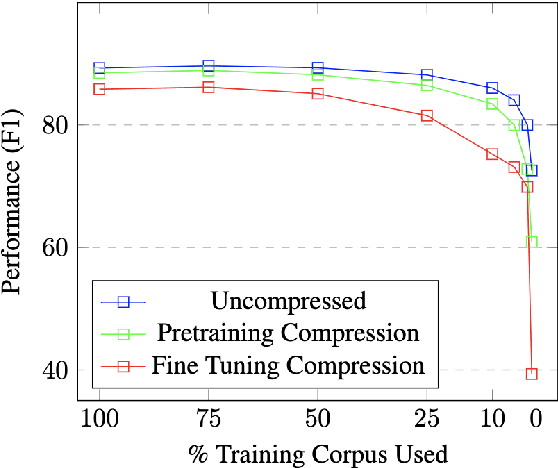
Abstract:Large Language Models have become the core architecture upon which most modern natural language processing (NLP) systems build. These models can consistently deliver impressive accuracy and robustness across tasks and domains, but their high computational overhead can make inference difficult and expensive. To make the usage of these models less costly recent work has explored leveraging structured and unstructured pruning, quantization, and distillation as ways to improve inference speed and decrease size. This paper studies how models pruned using Gradual Unstructured Magnitude Pruning can transfer between domains and tasks. Our experimentation shows that models that are pruned during pretraining using general domain masked language models can transfer to novel domains and tasks without extensive hyperparameter exploration or specialized approaches. We demonstrate that our general sparse model Sparse*BERT can become SparseBioBERT simply by pretraining the compressed architecture on unstructured biomedical text. Moreover, we show that SparseBioBERT can match the quality of BioBERT with only 10\% of the parameters.
Domain Representative Keywords Selection: A Probabilistic Approach
Mar 19, 2022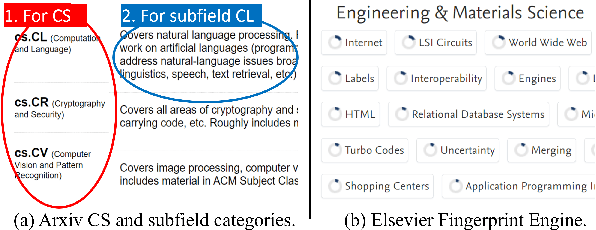
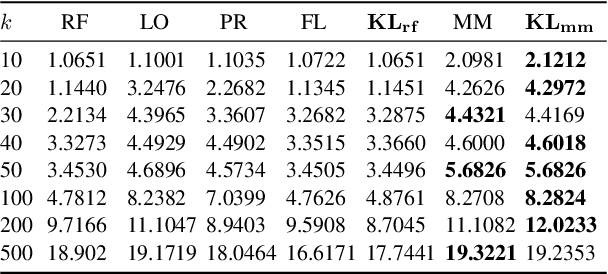
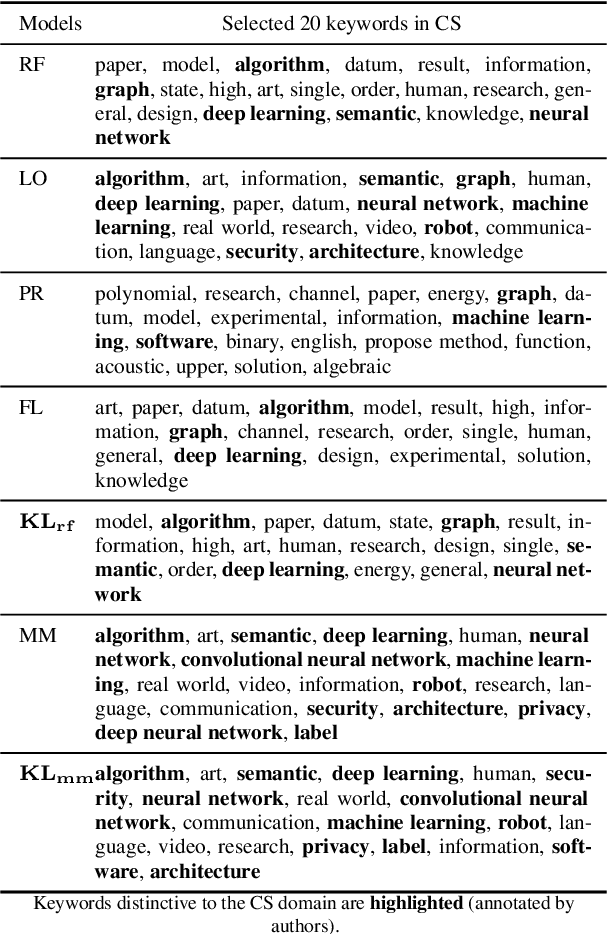
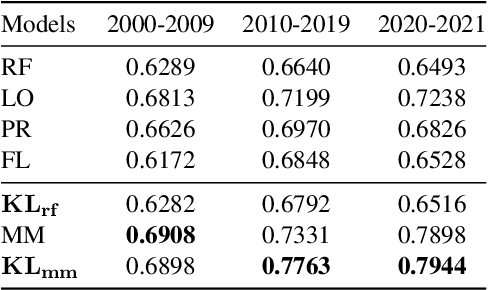
Abstract:We propose a probabilistic approach to select a subset of a \textit{target domain representative keywords} from a candidate set, contrasting with a context domain. Such a task is crucial for many downstream tasks in natural language processing. To contrast the target domain and the context domain, we adapt the \textit{two-component mixture model} concept to generate a distribution of candidate keywords. It provides more importance to the \textit{distinctive} keywords of the target domain than common keywords contrasting with the context domain. To support the \textit{representativeness} of the selected keywords towards the target domain, we introduce an \textit{optimization algorithm} for selecting the subset from the generated candidate distribution. We have shown that the optimization algorithm can be efficiently implemented with a near-optimal approximation guarantee. Finally, extensive experiments on multiple domains demonstrate the superiority of our approach over other baselines for the tasks of keyword summary generation and trending keywords selection.
Improving Candidate Retrieval with Entity Profile Generation for Wikidata Entity Linking
Mar 14, 2022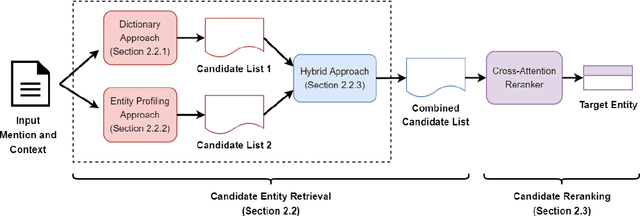
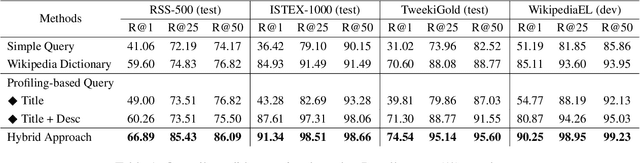
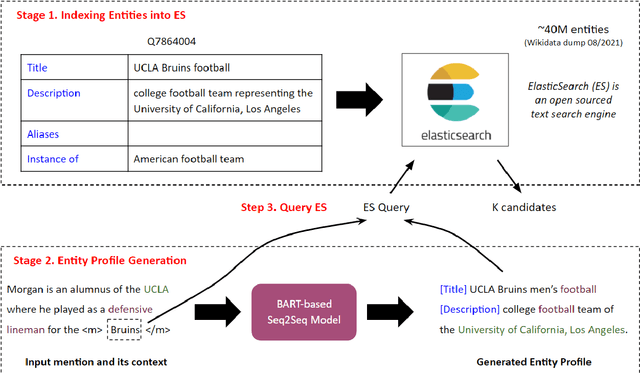
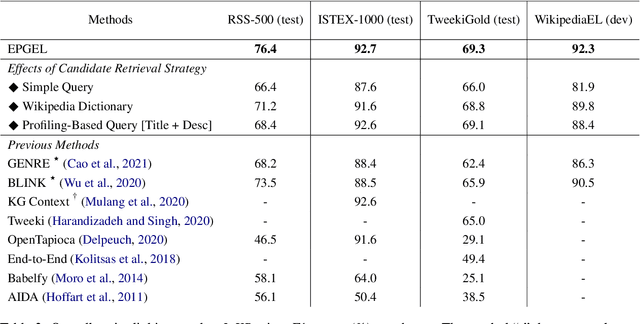
Abstract:Entity linking (EL) is the task of linking entity mentions in a document to referent entities in a knowledge base (KB). Many previous studies focus on Wikipedia-derived KBs. There is little work on EL over Wikidata, even though it is the most extensive crowdsourced KB. The scale of Wikidata can open up many new real-world applications, but its massive number of entities also makes EL challenging. To effectively narrow down the search space, we propose a novel candidate retrieval paradigm based on entity profiling. Wikidata entities and their textual fields are first indexed into a text search engine (e.g., Elasticsearch). During inference, given a mention and its context, we use a sequence-to-sequence (seq2seq) model to generate the profile of the target entity, which consists of its title and description. We use the profile to query the indexed search engine to retrieve candidate entities. Our approach complements the traditional approach of using a Wikipedia anchor-text dictionary, enabling us to further design a highly effective hybrid method for candidate retrieval. Combined with a simple cross-attention reranker, our complete EL framework achieves state-of-the-art results on three Wikidata-based datasets and strong performance on TACKBP-2010.
BERT might be Overkill: A Tiny but Effective Biomedical Entity Linker based on Residual Convolutional Neural Networks
Sep 06, 2021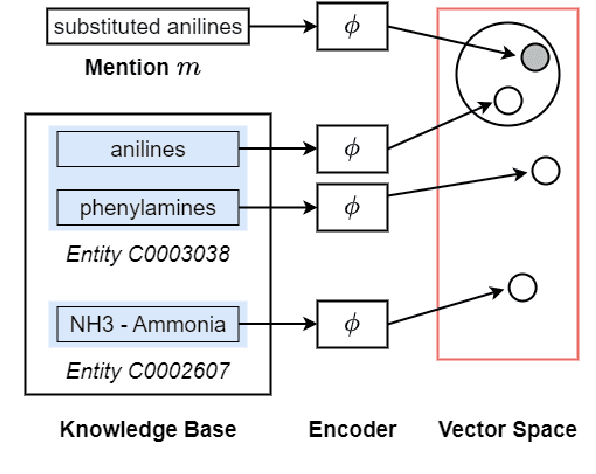
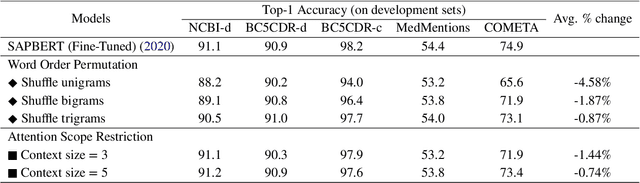
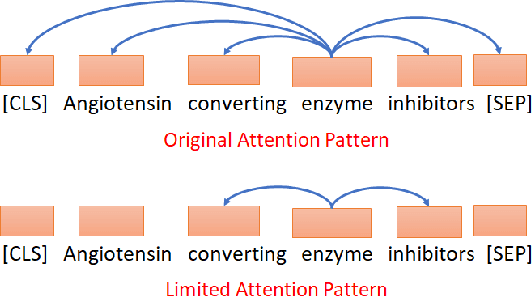
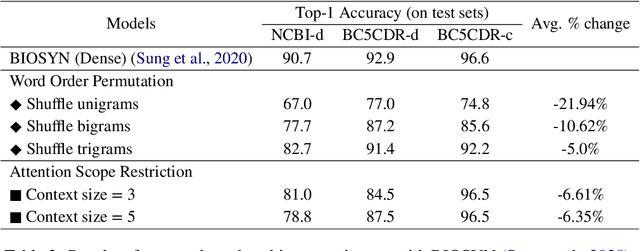
Abstract:Biomedical entity linking is the task of linking entity mentions in a biomedical document to referent entities in a knowledge base. Recently, many BERT-based models have been introduced for the task. While these models have achieved competitive results on many datasets, they are computationally expensive and contain about 110M parameters. Little is known about the factors contributing to their impressive performance and whether the over-parameterization is needed. In this work, we shed some light on the inner working mechanisms of these large BERT-based models. Through a set of probing experiments, we have found that the entity linking performance only changes slightly when the input word order is shuffled or when the attention scope is limited to a fixed window size. From these observations, we propose an efficient convolutional neural network with residual connections for biomedical entity linking. Because of the sparse connectivity and weight sharing properties, our model has a small number of parameters and is highly efficient. On five public datasets, our model achieves comparable or even better linking accuracy than the state-of-the-art BERT-based models while having about 60 times fewer parameters.
Fine-Grained Chemical Entity Typing with Multimodal Knowledge Representation
Aug 29, 2021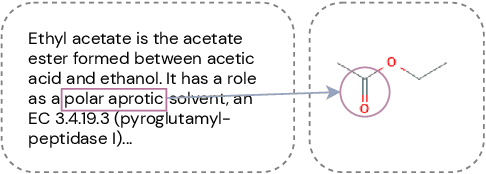
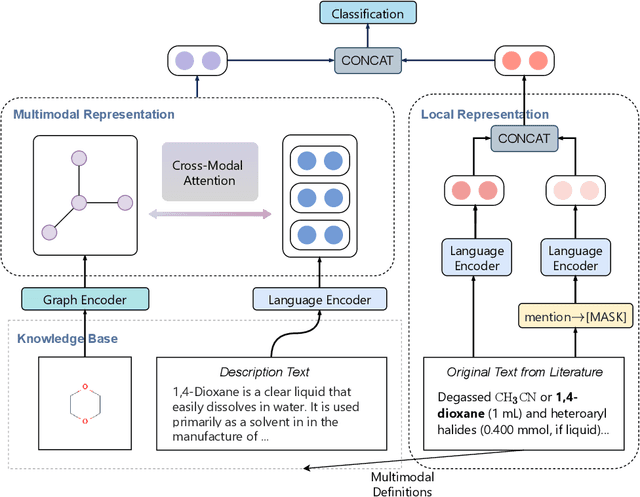
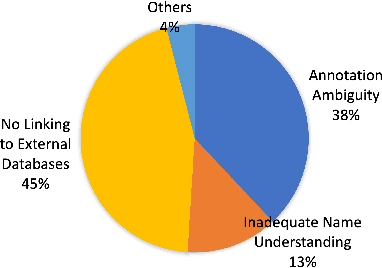
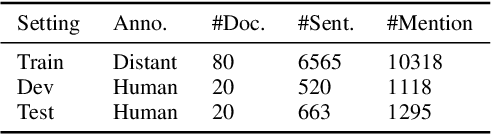
Abstract:Automated knowledge discovery from trending chemical literature is essential for more efficient biomedical research. How to extract detailed knowledge about chemical reactions from the core chemistry literature is a new emerging challenge that has not been well studied. In this paper, we study the new problem of fine-grained chemical entity typing, which poses interesting new challenges especially because of the complex name mentions frequently occurring in chemistry literature and graphic representation of entities. We introduce a new benchmark data set (CHEMET) to facilitate the study of the new task and propose a novel multi-modal representation learning framework to solve the problem of fine-grained chemical entity typing by leveraging external resources with chemical structures and using cross-modal attention to learn effective representation of text in the chemistry domain. Experiment results show that the proposed framework outperforms multiple state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge