Bram Ruijsink
King's College London
Cardiac Digital Twins at Scale from MRI: Open Tools and Representative Models from ~55000 UK Biobank Participants
May 27, 2025
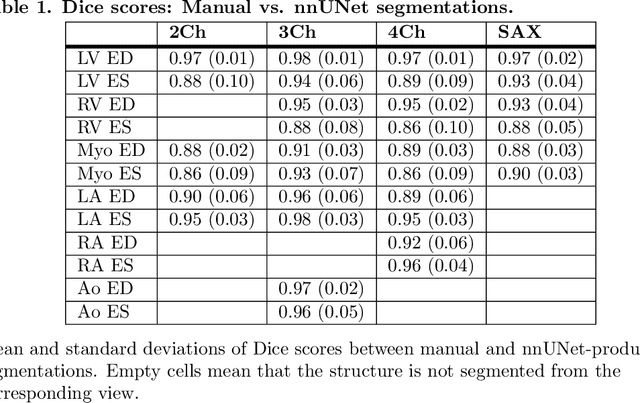
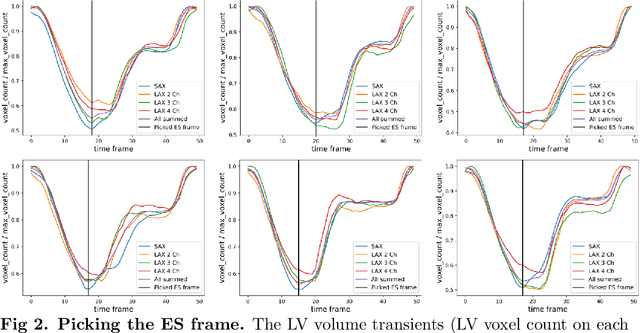
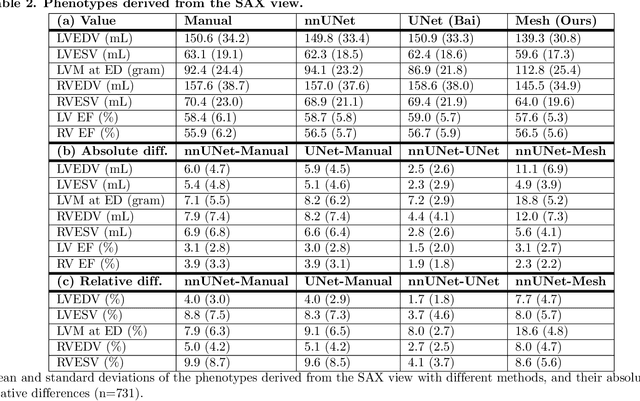
Abstract:A cardiac digital twin is a virtual replica of a patient's heart for screening, diagnosis, prognosis, risk assessment, and treatment planning of cardiovascular diseases. This requires an anatomically accurate patient-specific 3D structural representation of the heart, suitable for electro-mechanical simulations or study of disease mechanisms. However, generation of cardiac digital twins at scale is demanding and there are no public repositories of models across demographic groups. We describe an automatic open-source pipeline for creating patient-specific left and right ventricular meshes from cardiovascular magnetic resonance images, its application to a large cohort of ~55000 participants from UK Biobank, and the construction of the most comprehensive cohort of adult heart models to date, comprising 1423 representative meshes across sex (male, female), body mass index (range: 16 - 42 kg/m$^2$) and age (range: 49 - 80 years). Our code is available at https://github.com/cdttk/biv-volumetric-meshing/tree/plos2025 , and pre-trained networks, representative volumetric meshes with fibers and UVCs will be made available soon.
An investigation into the causes of race bias in AI-based cine CMR segmentation
Aug 05, 2024



Abstract:Artificial intelligence (AI) methods are being used increasingly for the automated segmentation of cine cardiac magnetic resonance (CMR) imaging. However, these methods have been shown to be subject to race bias, i.e. they exhibit different levels of performance for different races depending on the (im)balance of the data used to train the AI model. In this paper we investigate the source of this bias, seeking to understand its root cause(s) so that it can be effectively mitigated. We perform a series of classification and segmentation experiments on short-axis cine CMR images acquired from Black and White subjects from the UK Biobank and apply AI interpretability methods to understand the results. In the classification experiments, we found that race can be predicted with high accuracy from the images alone, but less accurately from ground truth segmentations, suggesting that the distributional shift between races, which is often the cause of AI bias, is mostly image-based rather than segmentation-based. The interpretability methods showed that most attention in the classification models was focused on non-heart regions, such as subcutaneous fat. Cropping the images tightly around the heart reduced classification accuracy to around chance level. Similarly, race can be predicted from the latent representations of a biased segmentation model, suggesting that race information is encoded in the model. Cropping images tightly around the heart reduced but did not eliminate segmentation bias. We also investigate the influence of possible confounders on the bias observed.
Improving Deep Learning Model Calibration for Cardiac Applications using Deterministic Uncertainty Networks and Uncertainty-aware Training
May 10, 2024



Abstract:Improving calibration performance in deep learning (DL) classification models is important when planning the use of DL in a decision-support setting. In such a scenario, a confident wrong prediction could lead to a lack of trust and/or harm in a high-risk application. We evaluate the impact on accuracy and calibration of two types of approach that aim to improve DL classification model calibration: deterministic uncertainty methods (DUM) and uncertainty-aware training. Specifically, we test the performance of three DUMs and two uncertainty-aware training approaches as well as their combinations. To evaluate their utility, we use two realistic clinical applications from the field of cardiac imaging: artefact detection from phase contrast cardiac magnetic resonance (CMR) and disease diagnosis from the public ACDC CMR dataset. Our results indicate that both DUMs and uncertainty-aware training can improve both accuracy and calibration in both of our applications, with DUMs generally offering the best improvements. We also investigate the combination of the two approaches, resulting in a novel deterministic uncertainty-aware training approach. This provides further improvements for some combinations of DUMs and uncertainty-aware training approaches.
Label Dropout: Improved Deep Learning Echocardiography Segmentation Using Multiple Datasets With Domain Shift and Partial Labelling
Mar 12, 2024Abstract:Echocardiography (echo) is the first imaging modality used when assessing cardiac function. The measurement of functional biomarkers from echo relies upon the segmentation of cardiac structures and deep learning models have been proposed to automate the segmentation process. However, in order to translate these tools to widespread clinical use it is important that the segmentation models are robust to a wide variety of images (e.g. acquired from different scanners, by operators with different levels of expertise etc.). To achieve this level of robustness it is necessary that the models are trained with multiple diverse datasets. A significant challenge faced when training with multiple diverse datasets is the variation in label presence, i.e. the combined data are often partially-labelled. Adaptations of the cross entropy loss function have been proposed to deal with partially labelled data. In this paper we show that training naively with such a loss function and multiple diverse datasets can lead to a form of shortcut learning, where the model associates label presence with domain characteristics, leading to a drop in performance. To address this problem, we propose a novel label dropout scheme to break the link between domain characteristics and the presence or absence of labels. We demonstrate that label dropout improves echo segmentation Dice score by 62% and 25% on two cardiac structures when training using multiple diverse partially labelled datasets.
Uncertainty Aware Training to Improve Deep Learning Model Calibration for Classification of Cardiac MR Images
Aug 29, 2023Abstract:Quantifying uncertainty of predictions has been identified as one way to develop more trustworthy artificial intelligence (AI) models beyond conventional reporting of performance metrics. When considering their role in a clinical decision support setting, AI classification models should ideally avoid confident wrong predictions and maximise the confidence of correct predictions. Models that do this are said to be well-calibrated with regard to confidence. However, relatively little attention has been paid to how to improve calibration when training these models, i.e., to make the training strategy uncertainty-aware. In this work we evaluate three novel uncertainty-aware training strategies comparing against two state-of-the-art approaches. We analyse performance on two different clinical applications: cardiac resynchronisation therapy (CRT) response prediction and coronary artery disease (CAD) diagnosis from cardiac magnetic resonance (CMR) images. The best-performing model in terms of both classification accuracy and the most common calibration measure, expected calibration error (ECE) was the Confidence Weight method, a novel approach that weights the loss of samples to explicitly penalise confident incorrect predictions. The method reduced the ECE by 17% for CRT response prediction and by 22% for CAD diagnosis when compared to a baseline classifier in which no uncertainty-aware strategy was included. In both applications, as well as reducing the ECE there was a slight increase in accuracy from 69% to 70% and 70% to 72% for CRT response prediction and CAD diagnosis respectively. However, our analysis showed a lack of consistency in terms of optimal models when using different calibration measures. This indicates the need for careful consideration of performance metrics when training and selecting models for complex high-risk applications in healthcare.
An investigation into the impact of deep learning model choice on sex and race bias in cardiac MR segmentation
Aug 25, 2023



Abstract:In medical imaging, artificial intelligence (AI) is increasingly being used to automate routine tasks. However, these algorithms can exhibit and exacerbate biases which lead to disparate performances between protected groups. We investigate the impact of model choice on how imbalances in subject sex and race in training datasets affect AI-based cine cardiac magnetic resonance image segmentation. We evaluate three convolutional neural network-based models and one vision transformer model. We find significant sex bias in three of the four models and racial bias in all of the models. However, the severity and nature of the bias varies between the models, highlighting the importance of model choice when attempting to train fair AI-based segmentation models for medical imaging tasks.
Automated Quality Controlled Analysis of 2D Phase Contrast Cardiovascular Magnetic Resonance Imaging
Sep 28, 2022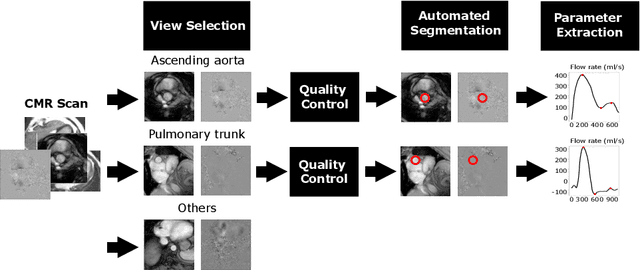
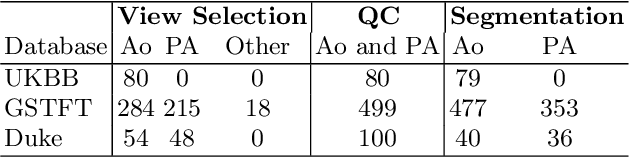
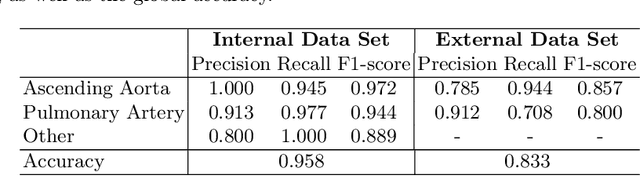
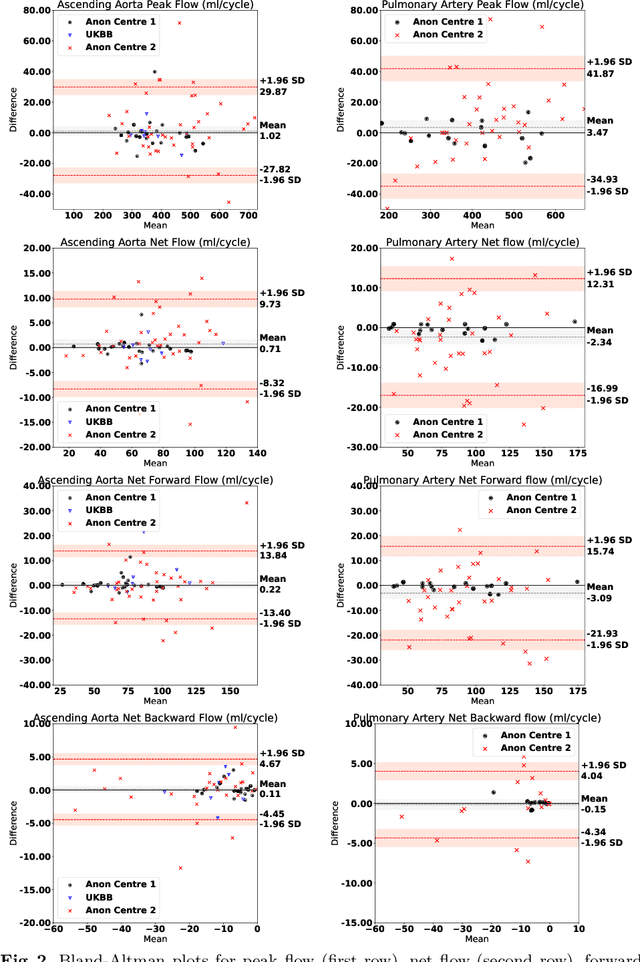
Abstract:Flow analysis carried out using phase contrast cardiac magnetic resonance imaging (PC-CMR) enables the quantification of important parameters that are used in the assessment of cardiovascular function. An essential part of this analysis is the identification of the correct CMR views and quality control (QC) to detect artefacts that could affect the flow quantification. We propose a novel deep learning based framework for the fully-automated analysis of flow from full CMR scans that first carries out these view selection and QC steps using two sequential convolutional neural networks, followed by automatic aorta and pulmonary artery segmentation to enable the quantification of key flow parameters. Accuracy values of 0.958 and 0.914 were obtained for view classification and QC, respectively. For segmentation, Dice scores were $>$0.969 and the Bland-Altman plots indicated excellent agreement between manual and automatic peak flow values. In addition, we tested our pipeline on an external validation data set, with results indicating good robustness of the pipeline. This work was carried out using multivendor clinical data consisting of 986 cases, indicating the potential for the use of this pipeline in a clinical setting.
A systematic study of race and sex bias in CNN-based cardiac MR segmentation
Sep 04, 2022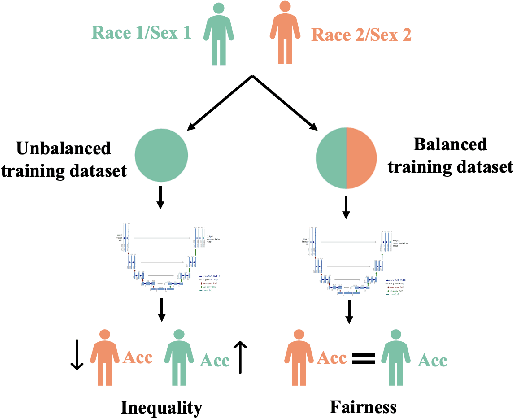
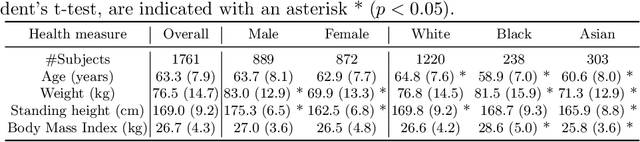
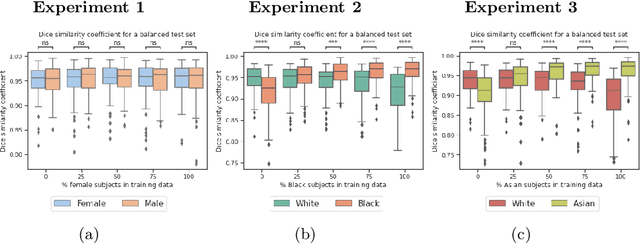
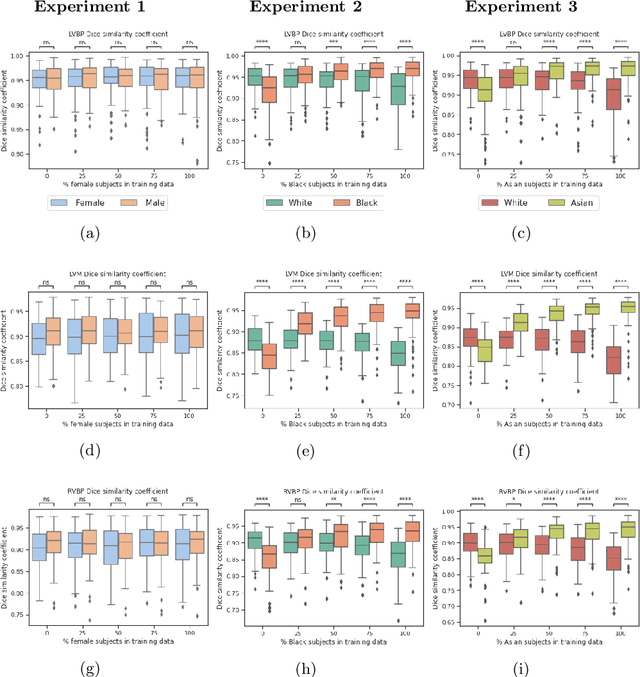
Abstract:In computer vision there has been significant research interest in assessing potential demographic bias in deep learning models. One of the main causes of such bias is imbalance in the training data. In medical imaging, where the potential impact of bias is arguably much greater, there has been less interest. In medical imaging pipelines, segmentation of structures of interest plays an important role in estimating clinical biomarkers that are subsequently used to inform patient management. Convolutional neural networks (CNNs) are starting to be used to automate this process. We present the first systematic study of the impact of training set imbalance on race and sex bias in CNN-based segmentation. We focus on segmentation of the structures of the heart from short axis cine cardiac magnetic resonance images, and train multiple CNN segmentation models with different levels of race/sex imbalance. We find no significant bias in the sex experiment but significant bias in two separate race experiments, highlighting the need to consider adequate representation of different demographic groups in health datasets.
Deep Learning-based Segmentation of Pleural Effusion From Ultrasound Using Coordinate Convolutions
Aug 05, 2022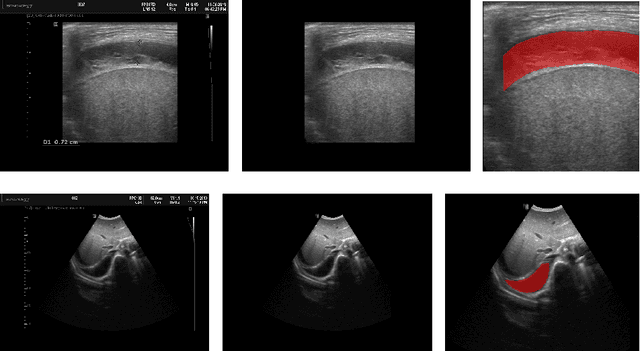

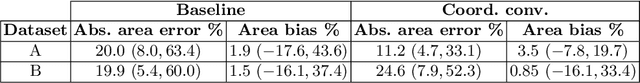
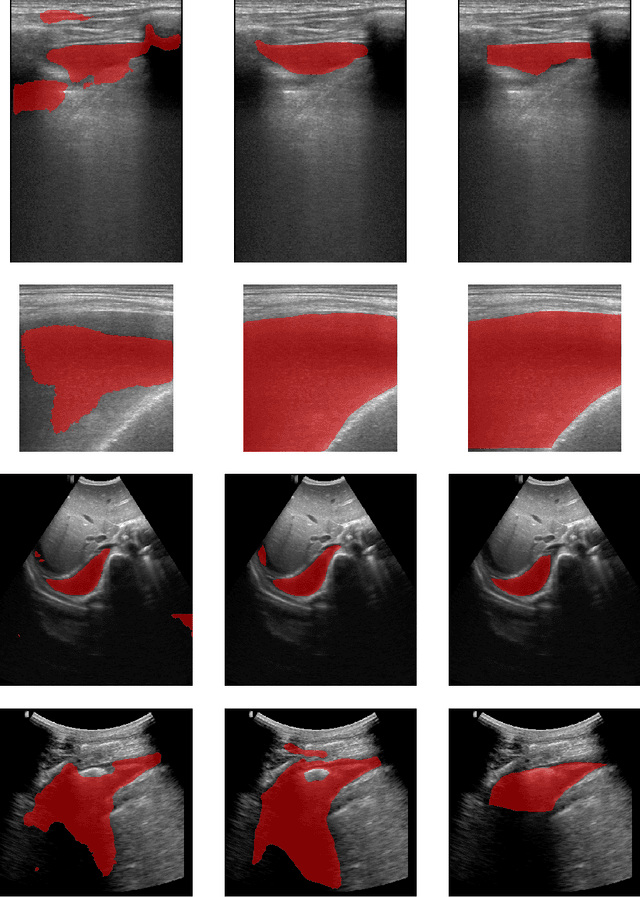
Abstract:In many low-to-middle income (LMIC) countries, ultrasound is used for assessment of pleural effusion. Typically, the extent of the effusion is manually measured by a sonographer, leading to significant intra-/inter-observer variability. In this work, we investigate the use of deep learning (DL) to automate the process of pleural effusion segmentation from ultrasound images. On two datasets acquired in a LMIC setting, we achieve median Dice Similarity Coefficients (DSCs) of 0.82 and 0.74 respectively using the nnU-net DL model. We also investigate the use of coordinate convolutions in the DL model and find that this results in a statistically significant improvement in the median DSC on the first dataset to 0.85, with no significant change on the second dataset. This work showcases, for the first time, the potential of DL in automating the process of effusion assessment from ultrasound in LMIC settings where there is often a lack of experienced radiologists to perform such tasks.
Large-scale, multi-centre, multi-disease validation of an AI clinical tool for cine CMR analysis
Jun 15, 2022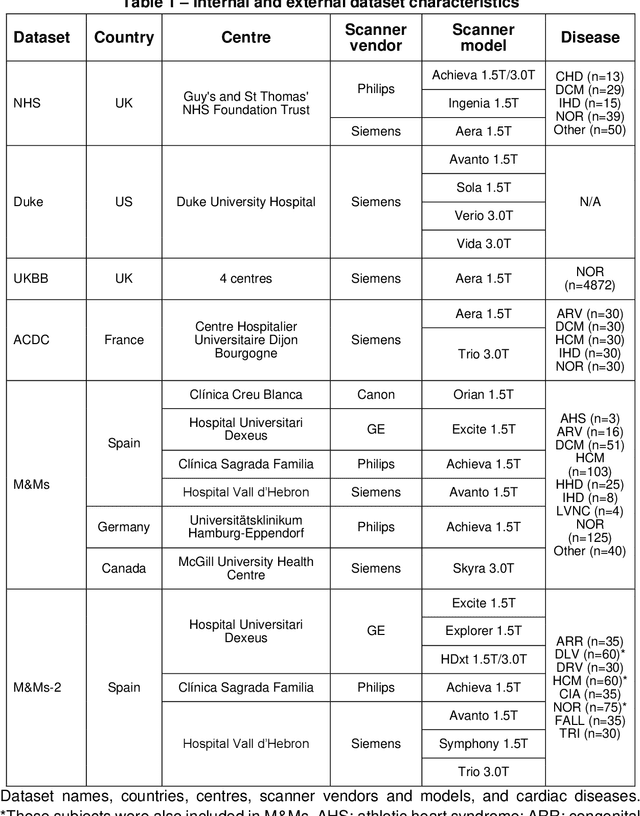
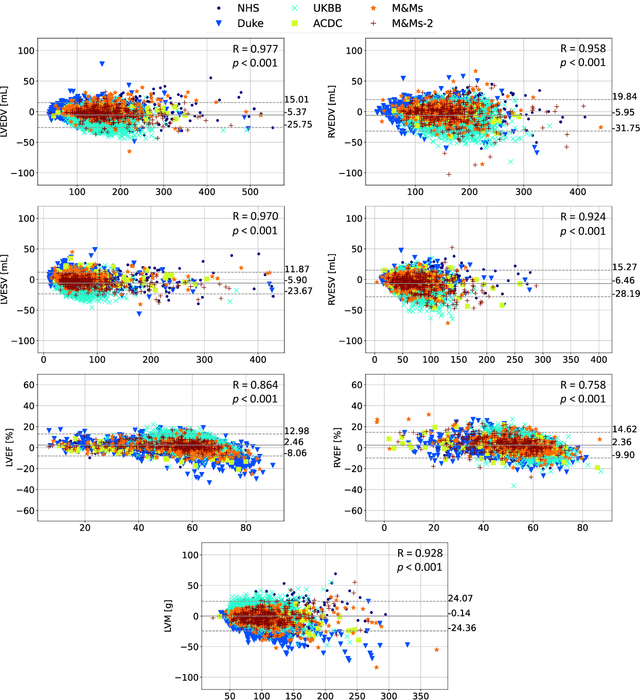
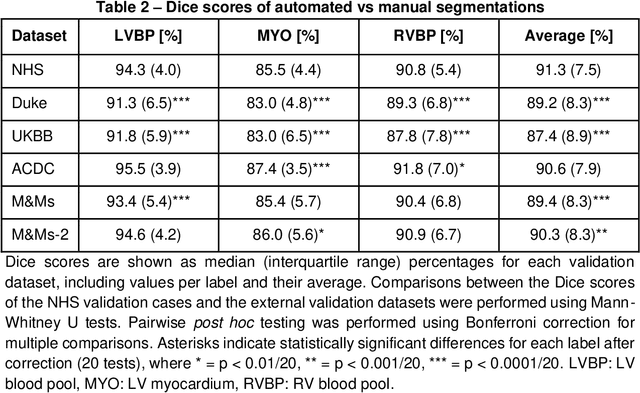
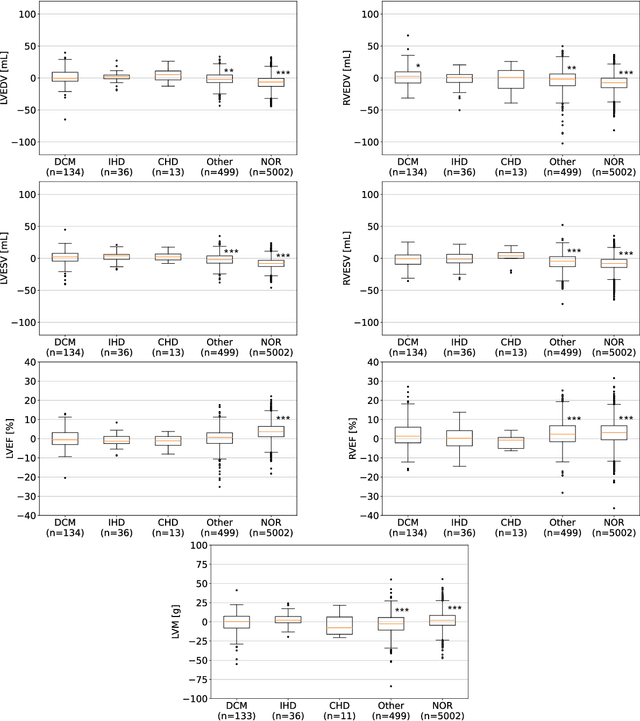
Abstract:INTRODUCTION: Artificial intelligence (AI) has the potential to facilitate the automation of CMR analysis for biomarker extraction. However, most AI algorithms are trained on a specific input domain (e.g., single scanner vendor or hospital-tailored imaging protocol) and lack the robustness to perform optimally when applied to CMR data from other input domains. METHODS: Our proposed framework consists of an AI-based algorithm for biventricular segmentation of short-axis images, followed by a post-analysis quality control to detect erroneous results. The segmentation algorithm was trained on a large dataset of clinical CMR scans from two NHS hospitals (n=2793) and validated on additional cases from this dataset (n=441) and on five external datasets (n=6808). The validation data included CMR scans of patients with a range of diseases acquired at 12 different centres using CMR scanners from all major vendors. RESULTS: Our method yielded median Dice scores over 87%, translating into median absolute errors in cardiac biomarkers within the range of inter-observer variability: <8.4mL (left ventricle), <9.2mL (right ventricle), <13.3g (left ventricular mass), and <5.9% (ejection fraction) across all datasets. Stratification of cases according to phenotypes of cardiac disease and scanner vendors showed good agreement. CONCLUSIONS: We show that our proposed tool, which combines a state-of-the-art AI algorithm trained on a large-scale multi-domain CMR dataset with a post-analysis quality control, allows us to robustly deal with routine clinical data from multiple centres, vendors, and cardiac diseases. This is a fundamental step for the clinical translation of AI algorithms. Moreover, our method yields a range of additional biomarkers of cardiac function (filling and ejection rates, regional wall motion, and strain) at no extra computational cost.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge