Bertrand Thirion
MIND
Statistically Valid Variable Importance Assessment through Conditional Permutations
Sep 14, 2023Abstract:Variable importance assessment has become a crucial step in machine-learning applications when using complex learners, such as deep neural networks, on large-scale data. Removal-based importance assessment is currently the reference approach, particularly when statistical guarantees are sought to justify variable inclusion. It is often implemented with variable permutation schemes. On the flip side, these approaches risk misidentifying unimportant variables as important in the presence of correlations among covariates. Here we develop a systematic approach for studying Conditional Permutation Importance (CPI) that is model agnostic and computationally lean, as well as reusable benchmarks of state-of-the-art variable importance estimators. We show theoretically and empirically that $\textit{CPI}$ overcomes the limitations of standard permutation importance by providing accurate type-I error control. When used with a deep neural network, $\textit{CPI}$ consistently showed top accuracy across benchmarks. An empirical benchmark on real-world data analysis in a large-scale medical dataset showed that $\textit{CPI}$ provides a more parsimonious selection of statistically significant variables. Our results suggest that $\textit{CPI}$ can be readily used as drop-in replacement for permutation-based methods.
Probing Brain Context-Sensitivity with Masked-Attention Generation
May 23, 2023

Abstract:Two fundamental questions in neurolinguistics concerns the brain regions that integrate information beyond the lexical level, and the size of their window of integration. To address these questions we introduce a new approach named masked-attention generation. It uses GPT-2 transformers to generate word embeddings that capture a fixed amount of contextual information. We then tested whether these embeddings could predict fMRI brain activity in humans listening to naturalistic text. The results showed that most of the cortex within the language network is sensitive to contextual information, and that the right hemisphere is more sensitive to longer contexts than the left. Masked-attention generation supports previous analyses of context-sensitivity in the brain, and complements them by quantifying the window size of context integration per voxel.
* 2 pages, 2 figures, CCN 2023
Information-Restricted Neural Language Models Reveal Different Brain Regions' Sensitivity to Semantics, Syntax and Context
Feb 28, 2023Abstract:A fundamental question in neurolinguistics concerns the brain regions involved in syntactic and semantic processing during speech comprehension, both at the lexical (word processing) and supra-lexical levels (sentence and discourse processing). To what extent are these regions separated or intertwined? To address this question, we trained a lexical language model, Glove, and a supra-lexical language model, GPT-2, on a text corpus from which we selectively removed either syntactic or semantic information. We then assessed to what extent these information-restricted models were able to predict the time-courses of fMRI signal of humans listening to naturalistic text. We also manipulated the size of contextual information provided to GPT-2 in order to determine the windows of integration of brain regions involved in supra-lexical processing. Our analyses show that, while most brain regions involved in language are sensitive to both syntactic and semantic variables, the relative magnitudes of these effects vary a lot across these regions. Furthermore, we found an asymmetry between the left and right hemispheres, with semantic and syntactic processing being more dissociated in the left hemisphere than in the right, and the left and right hemispheres showing respectively greater sensitivity to short and long contexts. The use of information-restricted NLP models thus shed new light on the spatial organization of syntactic processing, semantic processing and compositionality.
Neural Language Models are not Born Equal to Fit Brain Data, but Training Helps
Jul 07, 2022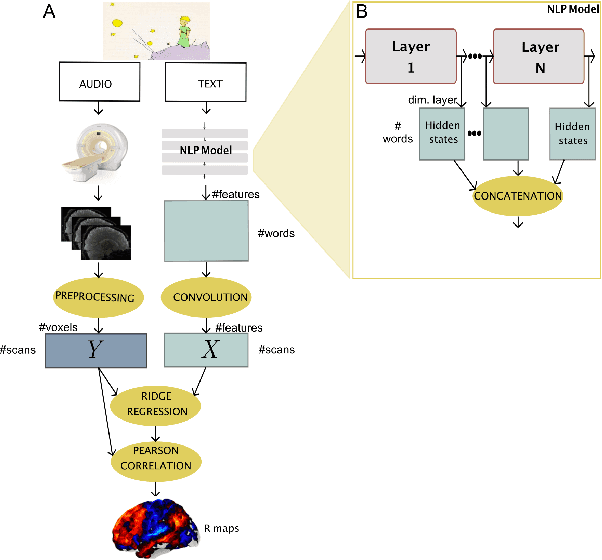
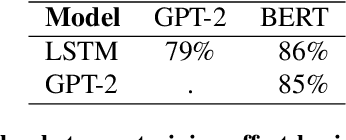
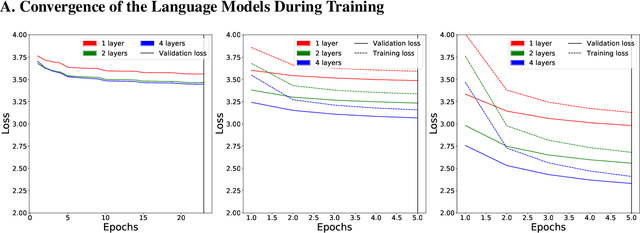
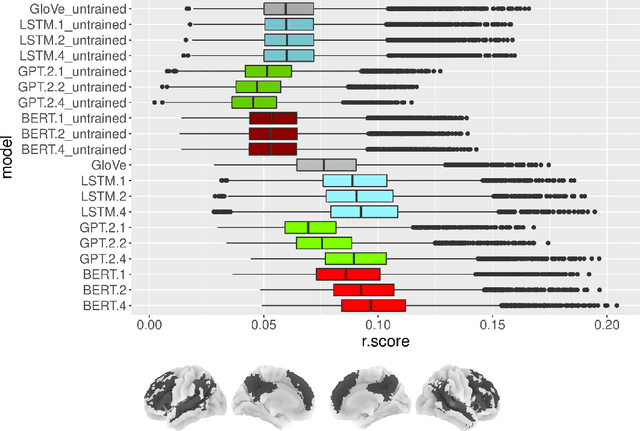
Abstract:Neural Language Models (NLMs) have made tremendous advances during the last years, achieving impressive performance on various linguistic tasks. Capitalizing on this, studies in neuroscience have started to use NLMs to study neural activity in the human brain during language processing. However, many questions remain unanswered regarding which factors determine the ability of a neural language model to capture brain activity (aka its 'brain score'). Here, we make first steps in this direction and examine the impact of test loss, training corpus and model architecture (comparing GloVe, LSTM, GPT-2 and BERT), on the prediction of functional Magnetic Resonance Imaging timecourses of participants listening to an audiobook. We find that (1) untrained versions of each model already explain significant amount of signal in the brain by capturing similarity in brain responses across identical words, with the untrained LSTM outperforming the transformerbased models, being less impacted by the effect of context; (2) that training NLP models improves brain scores in the same brain regions irrespective of the model's architecture; (3) that Perplexity (test loss) is not a good predictor of brain score; (4) that training data have a strong influence on the outcome and, notably, that off-the-shelf models may lack statistical power to detect brain activations. Overall, we outline the impact of modeltraining choices, and suggest good practices for future studies aiming at explaining the human language system using neural language models.
Aligning individual brains with Fused Unbalanced Gromov-Wasserstein
Jun 19, 2022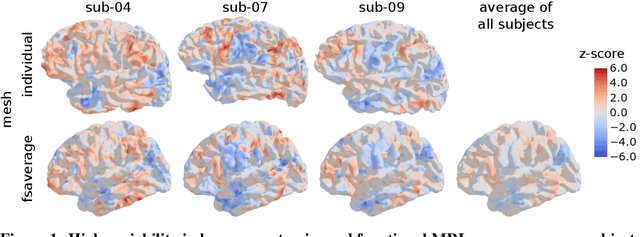
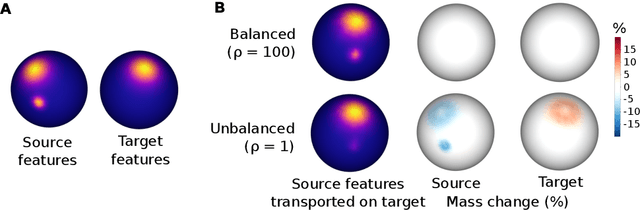
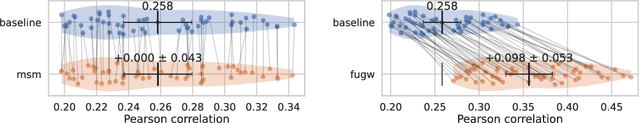
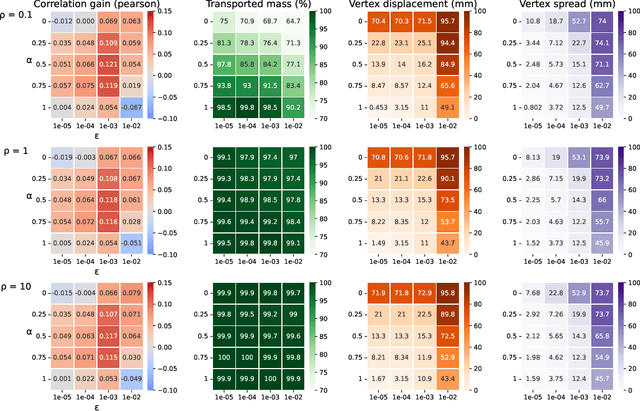
Abstract:Individual brains vary in both anatomy and functional organization, even within a given species. Inter-individual variability is a major impediment when trying to draw generalizable conclusions from neuroimaging data collected on groups of subjects. Current co-registration procedures rely on limited data, and thus lead to very coarse inter-subject alignments. In this work, we present a novel method for inter-subject alignment based on Optimal Transport, denoted as Fused Unbalanced Gromov Wasserstein (FUGW). The method aligns cortical surfaces based on the similarity of their functional signatures in response to a variety of stimulation settings, while penalizing large deformations of individual topographic organization. We demonstrate that FUGW is well-suited for whole-brain landmark-free alignment. The unbalanced feature allows to deal with the fact that functional areas vary in size across subjects. Our results show that FUGW alignment significantly increases between-subject correlation of activity for independent functional data, and leads to more precise mapping at the group level.
A Conditional Randomization Test for Sparse Logistic Regression in High-Dimension
May 29, 2022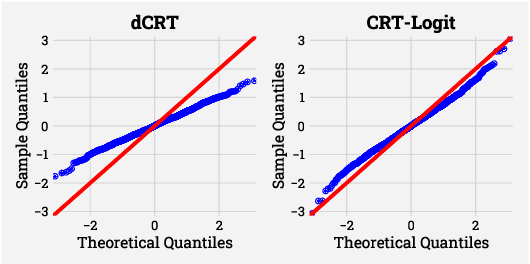

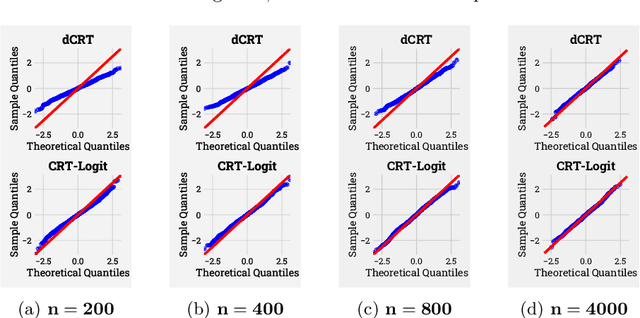
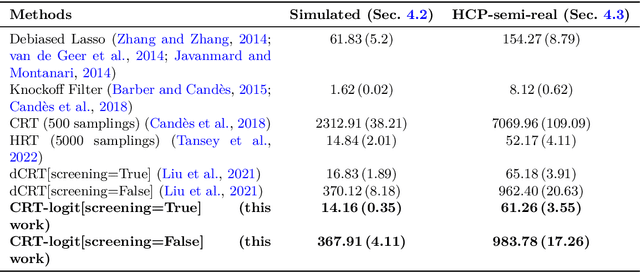
Abstract:Identifying the relevant variables for a classification model with correct confidence levels is a central but difficult task in high-dimension. Despite the core role of sparse logistic regression in statistics and machine learning, it still lacks a good solution for accurate inference in the regime where the number of features $p$ is as large as or larger than the number of samples $n$. Here, we tackle this problem by improving the Conditional Randomization Test (CRT). The original CRT algorithm shows promise as a way to output p-values while making few assumptions on the distribution of the test statistics. As it comes with a prohibitive computational cost even in mildly high-dimensional problems, faster solutions based on distillation have been proposed. Yet, they rely on unrealistic hypotheses and result in low-power solutions. To improve this, we propose \emph{CRT-logit}, an algorithm that combines a variable-distillation step and a decorrelation step that takes into account the geometry of $\ell_1$-penalized logistic regression problem. We provide a theoretical analysis of this procedure, and demonstrate its effectiveness on simulations, along with experiments on large-scale brain-imaging and genomics datasets.
Shared Independent Component Analysis for Multi-Subject Neuroimaging
Oct 26, 2021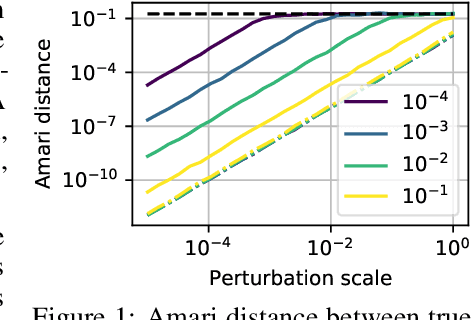
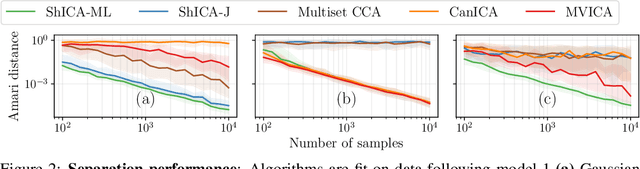
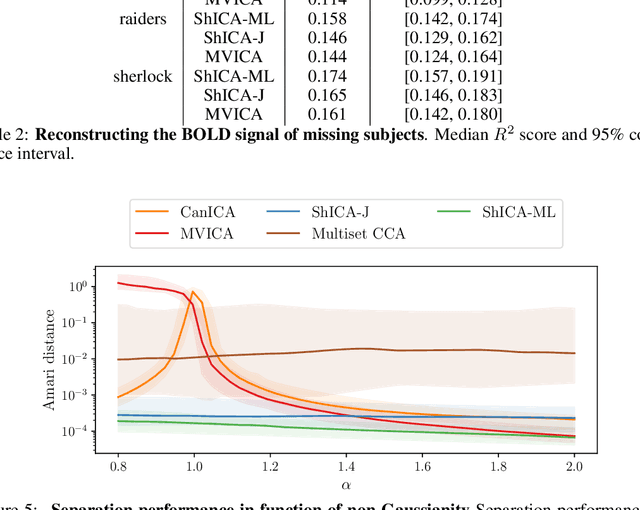
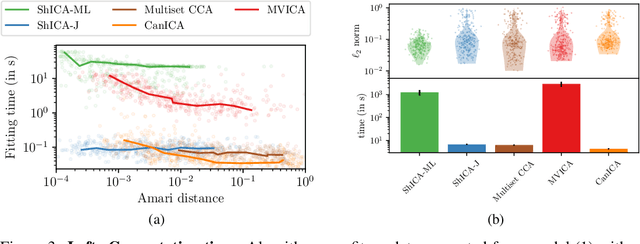
Abstract:We consider shared response modeling, a multi-view learning problem where one wants to identify common components from multiple datasets or views. We introduce Shared Independent Component Analysis (ShICA) that models each view as a linear transform of shared independent components contaminated by additive Gaussian noise. We show that this model is identifiable if the components are either non-Gaussian or have enough diversity in noise variances. We then show that in some cases multi-set canonical correlation analysis can recover the correct unmixing matrices, but that even a small amount of sampling noise makes Multiset CCA fail. To solve this problem, we propose to use joint diagonalization after Multiset CCA, leading to a new approach called ShICA-J. We show via simulations that ShICA-J leads to improved results while being very fast to fit. While ShICA-J is based on second-order statistics, we further propose to leverage non-Gaussianity of the components using a maximum-likelihood method, ShICA-ML, that is both more accurate and more costly. Further, ShICA comes with a principled method for shared components estimation. Finally, we provide empirical evidence on fMRI and MEG datasets that ShICA yields more accurate estimation of the components than alternatives.
Label scarcity in biomedicine: Data-rich latent factor discovery enhances phenotype prediction
Oct 12, 2021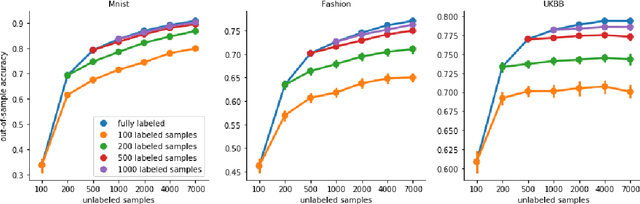
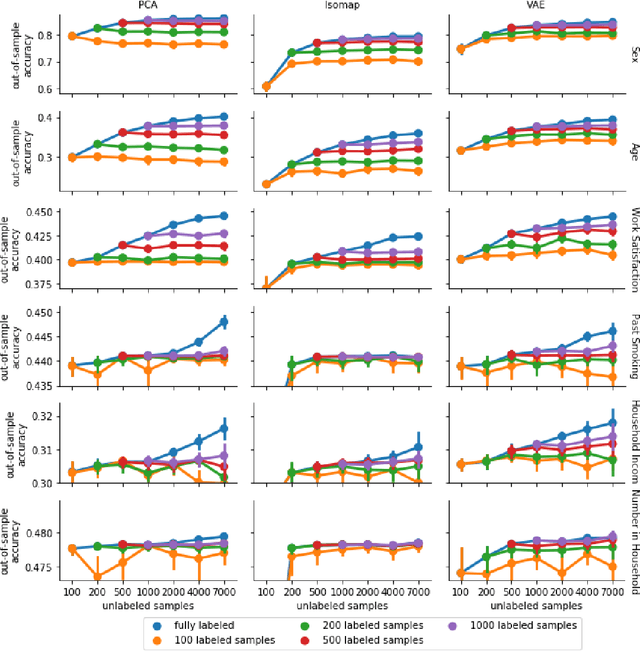
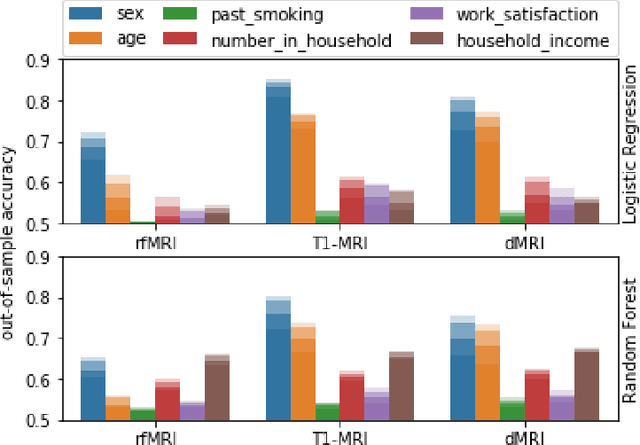
Abstract:High-quality data accumulation is now becoming ubiquitous in the health domain. There is increasing opportunity to exploit rich data from normal subjects to improve supervised estimators in specific diseases with notorious data scarcity. We demonstrate that low-dimensional embedding spaces can be derived from the UK Biobank population dataset and used to enhance data-scarce prediction of health indicators, lifestyle and demographic characteristics. Phenotype predictions facilitated by Variational Autoencoder manifolds typically scaled better with increasing unlabeled data than dimensionality reduction by PCA or Isomap. Performances gains from semisupervison approaches will probably become an important ingredient for various medical data science applications.
Functional Magnetic Resonance Imaging data augmentation through conditional ICA
Jul 14, 2021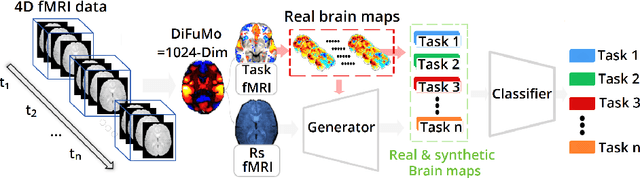
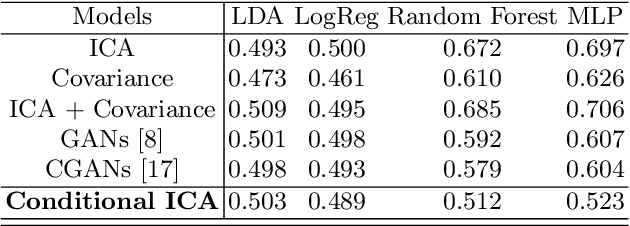
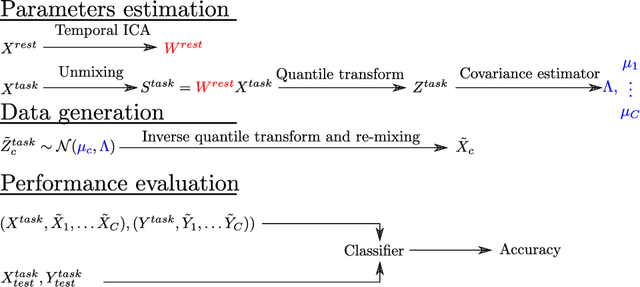
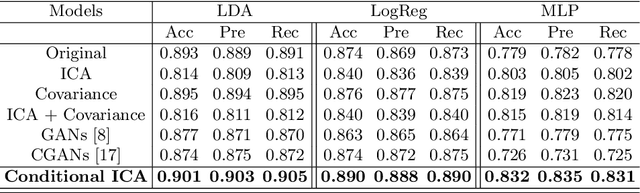
Abstract:Advances in computational cognitive neuroimaging research are related to the availability of large amounts of labeled brain imaging data, but such data are scarce and expensive to generate. While powerful data generation mechanisms, such as Generative Adversarial Networks (GANs), have been designed in the last decade for computer vision, such improvements have not yet carried over to brain imaging. A likely reason is that GANs training is ill-suited to the noisy, high-dimensional and small-sample data available in functional neuroimaging. In this paper, we introduce Conditional Independent Components Analysis (Conditional ICA): a fast functional Magnetic Resonance Imaging (fMRI) data augmentation technique, that leverages abundant resting-state data to create images by sampling from an ICA decomposition. We then propose a mechanism to condition the generator on classes observed with few samples. We first show that the generative mechanism is successful at synthesizing data indistinguishable from observations, and that it yields gains in classification accuracy in brain decoding problems. In particular it outperforms GANs while being much easier to optimize and interpret. Lastly, Conditional ICA enhances classification accuracy in eight datasets without further parameters tuning.
Spatially relaxed inference on high-dimensional linear models
Jun 04, 2021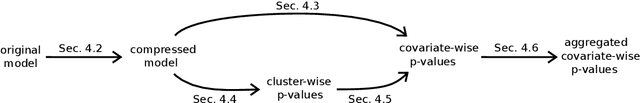
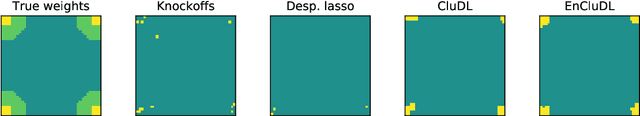
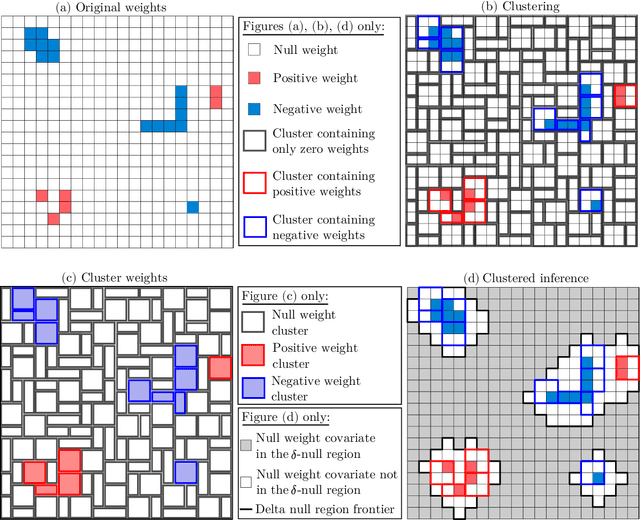

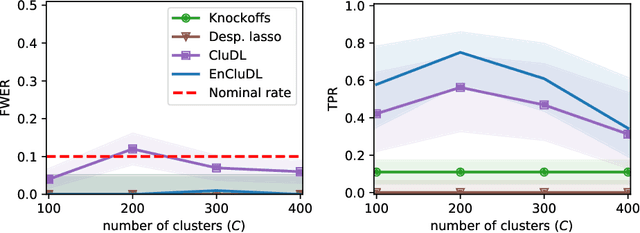
Abstract:We consider the inference problem for high-dimensional linear models, when covariates have an underlying spatial organization reflected in their correlation. A typical example of such a setting is high-resolution imaging, in which neighboring pixels are usually very similar. Accurate point and confidence intervals estimation is not possible in this context with many more covariates than samples, furthermore with high correlation between covariates. This calls for a reformulation of the statistical inference problem, that takes into account the underlying spatial structure: if covariates are locally correlated, it is acceptable to detect them up to a given spatial uncertainty. We thus propose to rely on the $\delta$-FWER, that is the probability of making a false discovery at a distance greater than $\delta$ from any true positive. With this target measure in mind, we study the properties of ensembled clustered inference algorithms which combine three techniques: spatially constrained clustering, statistical inference, and ensembling to aggregate several clustered inference solutions. We show that ensembled clustered inference algorithms control the $\delta$-FWER under standard assumptions for $\delta$ equal to the largest cluster diameter. We complement the theoretical analysis with empirical results, demonstrating accurate $\delta$-FWER control and decent power achieved by such inference algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge